3FM7
 
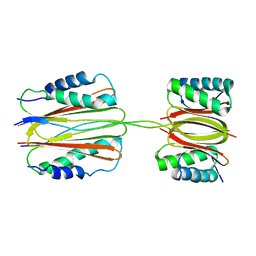 | | Quaternary Structure of Drosophila melanogaster IC/Tctex-1/LC8; Allosteric Interactions of Dynein Light Chains with Dynein Intermediate Chain | | Descriptor: | Dynein intermediate chain, cytosolic, Dynein light chain 1, ... | | Authors: | Hall, J.D, Karplus, P.A, Barbar, E.J. | | Deposit date: | 2008-12-19 | | Release date: | 2009-09-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Multivalency in the assembly of intrinsically disordered Dynein intermediate chain.
J.Biol.Chem., 284, 2009
|
|
1YF0
 
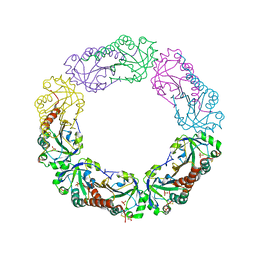 | | Structural and biochemical analysis of the link between enzymatic activity and oligomerization in AhpC, a bacterial peroxiredoxin. | | Descriptor: | Alkyl hydroperoxide reductase subunit C, SULFATE ION | | Authors: | Parsonage, D, Youngblood, D.S, Sarma, G.N, Wood, Z.A, Karplus, P.A, Poole, L.B. | | Deposit date: | 2004-12-29 | | Release date: | 2005-08-16 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Analysis of the Link between Enzymatic Activity and Oligomeric State in AhpC, a Bacterial Peroxiredoxin.
Biochemistry, 44, 2005
|
|
1YQZ
 
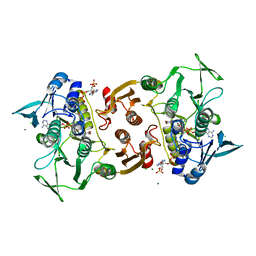 | | Structure of Coenzyme A-Disulfide Reductase from Staphylococcus aureus refined at 1.54 Angstrom resolution | | Descriptor: | CHLORIDE ION, COENZYME A, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Mallett, T.C, Wallen, J.R, Sakai, H, Luba, J, Parsonage, D, Karplus, P.A, Tsukihara, T, Claiborne, A. | | Deposit date: | 2005-02-02 | | Release date: | 2006-05-09 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Structure of coenzyme A-disulfide reductase from Staphylococcus aureus at 1.54 A resolution.
Biochemistry, 45, 2006
|
|
4L0U
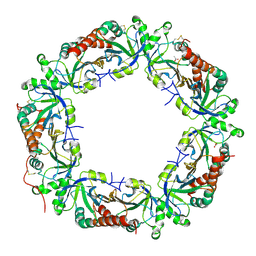 
 | | Crystal structure of Plasmodium vivax Prx1a | | Descriptor: | 2-Cys peroxiredoxin, putative, ACETATE ION | | Authors: | Gretes, M.C, Karplus, P.A. | | Deposit date: | 2013-06-01 | | Release date: | 2016-11-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Observed octameric assembly of a Plasmodium yoelii peroxiredoxin can be explained by the replacement of native "ball-and-socket" interacting residues by an affinity tag.
Protein Sci., 22, 2013
|
|
4L0W
 
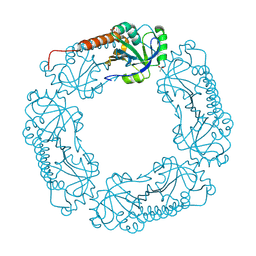 | |
4MAB
 
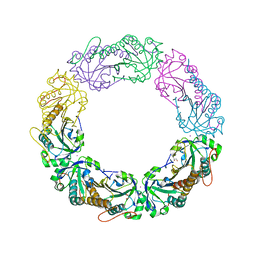 | | Resolving Cys to Ala variant of Salmonella Alkyl Hydroperoxide Reductase C in its substrate-ready conformation | | Descriptor: | Alkyl hydroperoxide reductase subunit C, CHLORIDE ION, GLYCEROL, ... | | Authors: | Perkins, A, Nelson, K.J, Williams, J.R, Poole, L.B, Karplus, P.A. | | Deposit date: | 2013-08-15 | | Release date: | 2013-11-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The sensitive balance between the fully folded and locally unfolded conformations of a model peroxiredoxin.
Biochemistry, 52, 2013
|
|
4MA9
 
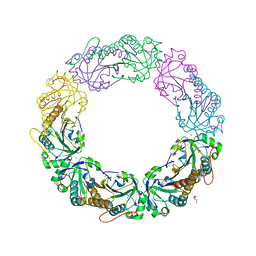 | |
4ND7
 
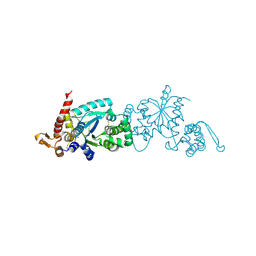 | | Crystal structure of apo 3-nitro-tyrosine tRNA synthetase (5B) in the closed form | | Descriptor: | BETA-MERCAPTOETHANOL, SODIUM ION, Tyrosine--tRNA ligase | | Authors: | Cooley, R.B, Driggers, C.M, Karplus, P.A, Mehl, R.A. | | Deposit date: | 2013-10-25 | | Release date: | 2014-03-19 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis of Improved Second-Generation 3-Nitro-tyrosine tRNA Synthetases.
Biochemistry, 53, 2014
|
|
4NDA
 
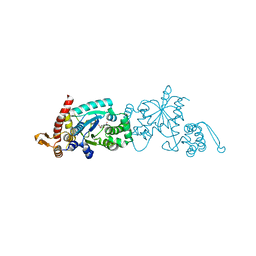 | | Crystal structure of 3-nitro-tyrosine tRNA synthetase (5B) bound to 3-nitro-tyrosine | | Descriptor: | GLYCEROL, META-NITRO-TYROSINE, SODIUM ION, ... | | Authors: | Cooley, R.B, Driggers, C.M, Karplus, P.A, Mehl, R.A. | | Deposit date: | 2013-10-25 | | Release date: | 2014-03-19 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis of Improved Second-Generation 3-Nitro-tyrosine tRNA Synthetases.
Biochemistry, 53, 2014
|
|
4ND6
 
 | | Crystal structure of apo 3-nitro-tyrosine tRNA synthetase (5B) in the open form | | Descriptor: | GLYCEROL, Tyrosine-tRNA ligase | | Authors: | Cooley, R.B, Driggers, C.M, Karplus, P.A, Mehl, R.A. | | Deposit date: | 2013-10-25 | | Release date: | 2014-03-19 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis of Improved Second-Generation 3-Nitro-tyrosine tRNA Synthetases.
Biochemistry, 53, 2014
|
|
4P53
 
 | | ValA (2-epi-5-epi-valiolone synthase) from Streptomyces hygroscopicus subsp. jinggangensis 5008 with NAD+ and Zn2+ bound | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, Cyclase, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Kean, K.M, Codding, S.J, Karplus, P.A. | | Deposit date: | 2014-03-13 | | Release date: | 2014-07-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of a sedoheptulose 7-phosphate cyclase: ValA from Streptomyces hygroscopicus.
Biochemistry, 53, 2014
|
|
4PBT
 
 | | Crystal structure of the M. jannaschii G2 tRNA synthetase variant bound to 4-trans-cyclooctene-amidopheylalanine (Tco-amF) | | Descriptor: | 1-ETHOXY-2-(2-ETHOXYETHOXY)ETHANE, 4-{[(1R,4E)-cyclooct-4-en-1-ylcarbonyl]amino}-L-phenylalanine, GLYCEROL, ... | | Authors: | Cooley, R.B, Karplus, P.A, Mehl, R.A. | | Deposit date: | 2014-04-13 | | Release date: | 2014-07-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Gleaning Unexpected Fruits from Hard-Won Synthetases: Probing Principles of Permissivity in Non-canonical Amino Acid-tRNA Synthetases.
Chembiochem, 15, 2014
|
|
4PBS
 
 | |
4PJY
 
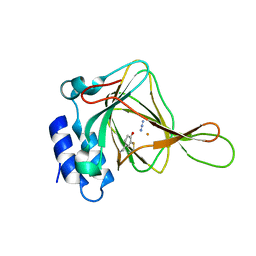 | | Azide bound Cysteine Dioxygenase at pH 6.2 | | Descriptor: | AZIDE ION, Cysteine dioxygenase type 1, FE (III) ION | | Authors: | Driggers, C.M, Karplus, P.A. | | Deposit date: | 2014-05-12 | | Release date: | 2016-02-17 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.501 Å) | | Cite: | Structure-Based Insights into the Role of the Cys-Tyr Crosslink and Inhibitor Recognition by Mammalian Cysteine Dioxygenase.
J. Mol. Biol., 428, 2016
|
|
4PTZ
 
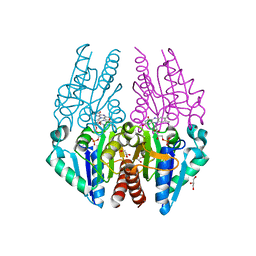 | | Crystal structure of the Escherichia coli alkanesulfonate FMN reductase SsuE in FMN-bound form | | Descriptor: | FLAVIN MONONUCLEOTIDE, FMN reductase SsuE, GLYCEROL, ... | | Authors: | Driggers, C.M, Ellis, H.R, Karplus, P.A. | | Deposit date: | 2014-03-11 | | Release date: | 2014-06-18 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9007 Å) | | Cite: | Crystal Structure of Escherichia coli SsuE: Defining a General Catalytic Cycle for FMN Reductases of the Flavodoxin-like Superfamily.
Biochemistry, 53, 2014
|
|
4PBR
 
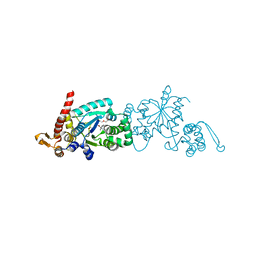 | | Crystal structure of the M. jannaschii G2 tRNA synthetase variant bound to 4-(2-bromoisobutyramido)-phenylalanine (BibaF) | | Descriptor: | 4-[(2-bromo-2-methylpropanoyl)amino]-L-phenylalanine, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Cooley, R.B, Karplus, P.A, Mehl, R.A. | | Deposit date: | 2014-04-13 | | Release date: | 2014-07-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Gleaning Unexpected Fruits from Hard-Won Synthetases: Probing Principles of Permissivity in Non-canonical Amino Acid-tRNA Synthetases.
Chembiochem, 15, 2014
|
|
4PU0
 
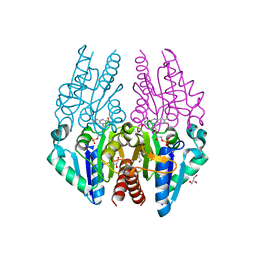 | | Crystal structure of the Escherichia coli alkanesulfonate FMN reductase SsuE in FMNH2-bound form | | Descriptor: | FLAVIN MONONUCLEOTIDE, FMN reductase SsuE, GLYCEROL, ... | | Authors: | Driggers, C.M, Ellis, H.R, Karplus, P.A. | | Deposit date: | 2014-03-11 | | Release date: | 2014-06-18 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.3018 Å) | | Cite: | Crystal Structure of Escherichia coli SsuE: Defining a General Catalytic Cycle for FMN Reductases of the Flavodoxin-like Superfamily.
Biochemistry, 53, 2014
|
|
4PIY
 
 | |
4PIX
 
 | | Unliganded Cysteine Dioxygenase C93A variant at pH 6.2 | | Descriptor: | CHLORIDE ION, Cysteine dioxygenase type 1, FE (III) ION | | Authors: | Driggers, C.M, Karplus, P.A. | | Deposit date: | 2014-05-09 | | Release date: | 2016-02-17 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structure-Based Insights into the Role of the Cys-Tyr Crosslink and Inhibitor Recognition by Mammalian Cysteine Dioxygenase.
J. Mol. Biol., 428, 2016
|
|
4PIZ
 
 | | Homocysteine-bound Cysteine Dioxygenase at pH 6.2 | | Descriptor: | 2-AMINO-4-MERCAPTO-BUTYRIC ACID, Cysteine dioxygenase type 1, FE (III) ION | | Authors: | Driggers, C.M, Karplus, P.A. | | Deposit date: | 2014-05-09 | | Release date: | 2016-02-17 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structure-Based Insights into the Role of the Cys-Tyr Crosslink and Inhibitor Recognition by Mammalian Cysteine Dioxygenase.
J. Mol. Biol., 428, 2016
|
|
4PTY
 
 | |
4QM8
 
 | |
1FRN
 
 | |
1FWJ
 
 | | KLEBSIELLA AEROGENES UREASE, NATIVE | | Descriptor: | NICKEL (II) ION, UREASE | | Authors: | Pearson, M.A, Karplus, P.A. | | Deposit date: | 1997-04-23 | | Release date: | 1997-10-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structures of Cys319 variants and acetohydroxamate-inhibited Klebsiella aerogenes urease.
Biochemistry, 36, 1997
|
|
1FWF
 
 | |
