1RP4
 
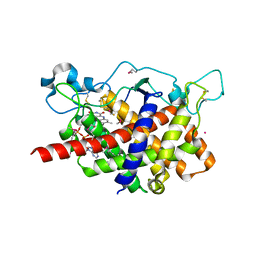 | | Structure of Ero1p, Source of Disulfide Bonds for Oxidative Protein Folding in the Cell | | Descriptor: | 1-ETHYL-PYRROLIDINE-2,5-DIONE, CADMIUM ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Gross, E, Kastner, D.B, Kaiser, C.A, Fass, D. | | Deposit date: | 2003-12-03 | | Release date: | 2004-06-08 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of ero1p, source of disulfide bonds for oxidative protein folding in the cell.
Cell(Cambridge,Mass.), 117, 2004
|
|
1JR8
 
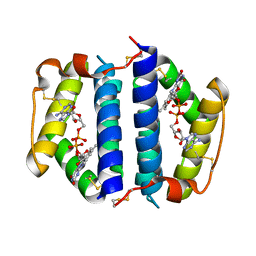 | | Crystal Structure of Erv2p | | Descriptor: | Erv2 PROTEIN, mitochondrial, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Gross, E, Sevier, C.S, Vala, A, Kaiser, C.A, Fass, D. | | Deposit date: | 2001-08-13 | | Release date: | 2001-12-28 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | A new FAD-binding fold and intersubunit disulfide shuttle in the thiol oxidase Erv2p.
Nat.Struct.Biol., 9, 2002
|
|
1JRA
 
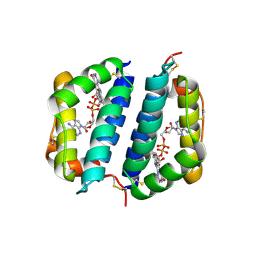 | | Crystal Structure of Erv2p | | Descriptor: | ERV2 PROTEIN, MITOCHONDRIAL, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Gross, E, Sevier, C.S, Vala, A, Kaiser, C.A, Fass, D. | | Deposit date: | 2001-08-13 | | Release date: | 2001-12-28 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A new FAD-binding fold and intersubunit disulfide shuttle in the thiol oxidase Erv2p.
Nat.Struct.Biol., 9, 2002
|
|
1RQ1
 
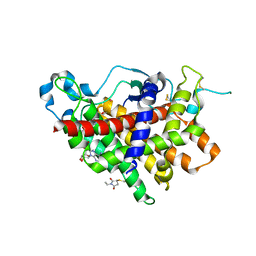 | | Structure of Ero1p, Source of Disulfide Bonds for Oxidative Protein Folding in the Cell | | Descriptor: | 1-ETHYL-PYRROLIDINE-2,5-DIONE, CADMIUM ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Gross, E, Kastner, D.B, Kaiser, C.A, Fass, D. | | Deposit date: | 2003-12-04 | | Release date: | 2004-06-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of ero1p, source of disulfide bonds for oxidative protein folding in the cell.
Cell(Cambridge,Mass.), 117, 2004
|
|
3ED3
 
 | | Crystal Structure of the Yeast Dithiol/Disulfide Oxidoreductase Mpd1p | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Protein disulfide-isomerase MPD1 | | Authors: | Vitu, E, Greenblatt, H.M, Fass, D. | | Deposit date: | 2008-09-02 | | Release date: | 2008-11-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Yeast Mpd1p reveals the structural diversity of the protein disulfide isomerase family
J.Mol.Biol., 384, 2008
|
|
3ZPH
 | |
4D4F
 | | Mutant P250A of bacterial chalcone isomerase from Eubacterium ramulus | | Descriptor: | CHALCONE ISOMERASE, CHLORIDE ION, GLYCEROL | | Authors: | Thomsen, M, Kratzat, H, Hinrichs, W. | | Deposit date: | 2014-10-28 | | Release date: | 2016-01-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Structural Basis for (2 R ,3 R )-Taxifolin Binding and Reaction Products to the Bacterial Chalcone Isomerase of Eubacterium ramulus.
Molecules, 27, 2022
|
|
8SMU
 
 | | Integral fusion of the HtaA CR2 domain from Corynebacterium diphtheriae within EGFP | | Descriptor: | CHLORIDE ION, GLYCEROL, HtaACR2 integral fusion within enhanced green fluorescent protein, ... | | Authors: | Mahoney, B.J, Cascio, D, Clubb, R.T. | | Deposit date: | 2023-04-26 | | Release date: | 2023-09-27 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Development and atomic structure of a new fluorescence-based sensor to probe heme transfer in bacterial pathogens.
J.Inorg.Biochem., 249, 2023
|
|
7R0U
 
 | | Structure of a cytosolic sulfotransferase of Anopheles gambiae (AGAP001425) in complex with 3'-phosphoadenosine 5-phosphate and vanillin. | | Descriptor: | 4-hydroxy-3-methoxybenzaldehyde, ADENOSINE-3'-5'-DIPHOSPHATE, AGAP001425-PA, ... | | Authors: | Esposito Verza, A, Miggiano, R, Rizzi, R, Rossi, F. | | Deposit date: | 2022-02-02 | | Release date: | 2022-08-10 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Biochemical and structural analysis of a cytosolic sulfotransferase of the malaria vector Anopheles gambiae overexpressed in the reproductive tissues.
Curr Res Struct Biol, 4, 2022
|
|
7R0O
 
 | | Structure of a cytosolic sulfotransferase of Anopheles gambiae (AGAP001425). | | Descriptor: | AGAP001425-PA, CHLORIDE ION, GLYCEROL | | Authors: | Esposito Verza, A, Miggiano, R, Rizzi, R, Rossi, F. | | Deposit date: | 2022-02-02 | | Release date: | 2022-08-10 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Biochemical and structural analysis of a cytosolic sulfotransferase of the malaria vector Anopheles gambiae overexpressed in the reproductive tissues.
Curr Res Struct Biol, 4, 2022
|
|
7R0S
 
 | | Structure of a cytosolic sulfotransferase of Anopheles gambiae (AGAP001425) in complex with vanillin | | Descriptor: | 4-hydroxy-3-methoxybenzaldehyde, AGAP001425-PA, CHLORIDE ION, ... | | Authors: | Esposito Verza, A, Miggiano, R, Rizzi, R, Rossi, F. | | Deposit date: | 2022-02-02 | | Release date: | 2022-08-10 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Biochemical and structural analysis of a cytosolic sulfotransferase of the malaria vector Anopheles gambiae overexpressed in the reproductive tissues.
Curr Res Struct Biol, 4, 2022
|
|
