2K6Y
 
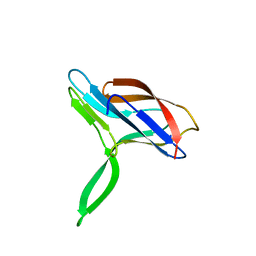 | | Solution structures of apo form PCuA (cis conformation of the peptide bond involving the nitrogen of P14) | | Descriptor: | Putative uncharacterized protein TTHA1943 | | Authors: | Abriata, L.A, Banci, L, Bertini, I, Ciofi-Baffoni, S, Gkazonis, P, Spyroulias, G.A, Vila, A.J, Wang, S. | | Deposit date: | 2008-07-28 | | Release date: | 2008-09-09 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Mechanism of Cu(A) assembly.
Nat.Chem.Biol., 4, 2008
|
|
2L4D
 
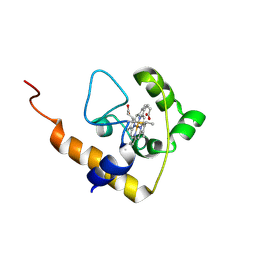 | | cytochrome c domain of pp3183 protein from Pseudomonas putida | | Descriptor: | HEME C, SCO1/SenC family protein/cytochrome c | | Authors: | Banci, L, Bertini, I, Ciofi-Baffoni, S, Kozyreva, T, Mori, M, Wang, S. | | Deposit date: | 2010-10-04 | | Release date: | 2011-01-26 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Sco proteins are involved in electron transfer processes
J.Biol.Inorg.Chem., 16, 2011
|
|
2GCF
 
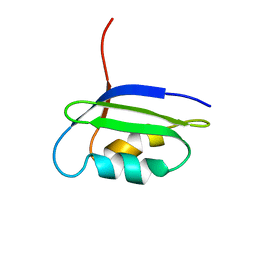 | | Solution structure of the N-terminal domain of the coppper(I) ATPase PacS in its apo form | | Descriptor: | Cation-transporting ATPase pacS | | Authors: | Banci, L, Bertini, I, Ciofi-Baffoni, S, Kandias, N.G, Spyroulias, G.A, Robinson, N.J, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2006-03-14 | | Release date: | 2006-05-30 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | The delivery of copper for thylakoid import observed by NMR.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
2LQL
 
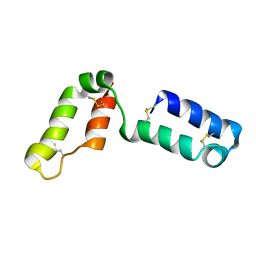 | | Solution structure of CHCH5 | | Descriptor: | Coiled-coil-helix-coiled-coil-helix domain-containing protein 5 | | Authors: | Peruzzini, R, Ciofi-Baffoni, S, Banci, L, Bertini, I. | | Deposit date: | 2012-03-09 | | Release date: | 2012-10-24 | | Last modified: | 2023-12-06 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of CHCHD5 and CHCHD7: Two atypical human twin CX(9)C proteins.
J.Struct.Biol., 180, 2012
|
|
2L0Y
 
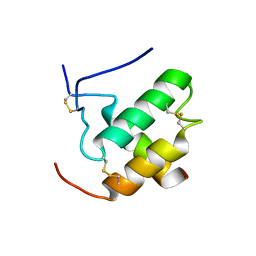 | | Complex hMia40-hCox17 | | Descriptor: | COX17 cytochrome c oxidase assembly homolog (S. cerevisiae) pseudogene (COX17), Mitochondrial intermembrane space import and assembly protein 40 | | Authors: | Bertini, I, Ciofi-Baffoni, S, Gallo, A. | | Deposit date: | 2010-07-20 | | Release date: | 2010-11-24 | | Last modified: | 2020-02-05 | | Method: | SOLUTION NMR | | Cite: | Molecular chaperone function of Mia40 triggers consecutive induced folding steps of the substrate in mitochondrial protein import.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2LQT
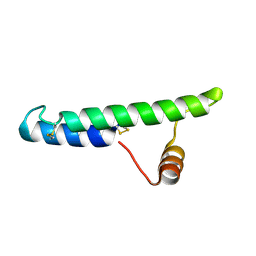 
 | |
2JSC
 
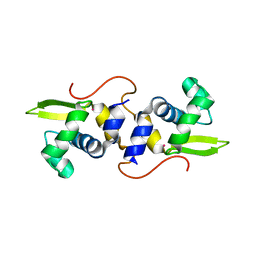 | | NMR structure of the cadmium metal-sensor CMTR from Mycobacterium tuberculosis | | Descriptor: | CADMIUM ION, Transcriptional regulator Rv1994c/MT2050 | | Authors: | Banci, L, Bertini, I, Cantini, F, Ciofi-Baffoni, S, Cavet, J.S, Dennison, C, Graham, A.I, Harvie, D.R, Robinson, N.J, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2007-07-02 | | Release date: | 2007-07-31 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | NMR Structural Analysis of Cadmium Sensing by Winged Helix Repressor CmtR.
J.Biol.Chem., 282, 2007
|
|
2LD4
 
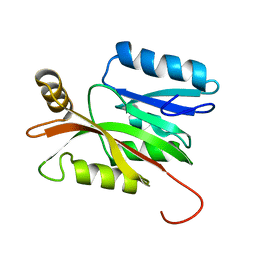 | | Solution structure of the N-terminal domain of human anamorsin | | Descriptor: | Anamorsin | | Authors: | Banci, L, Bertini, I, Ciofi-Baffoni, S, Boscaro, F, Chatzi, A, Mikolajczyk, M, Tokatlidis, K, Winkelmann, J. | | Deposit date: | 2011-05-13 | | Release date: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Anamorsin Is a [2Fe-2S] Cluster-Containing Substrate of the Mia40-Dependent Mitochondrial Protein Trapping Machinery.
Chem.Biol., 18, 2011
|
|
2LGQ
 
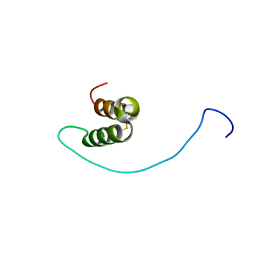 | | Human C30S/C59S-Cox17 mutant | | Descriptor: | Cytochrome c oxidase copper chaperone | | Authors: | Bertini, I, Ciofi-Baffoni, S, Gallo, A. | | Deposit date: | 2011-08-01 | | Release date: | 2011-08-17 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Functional role of two interhelical disulfide bonds in human cox17 protein from a structural perspective.
J.Biol.Chem., 286, 2011
|
|
5LCI
 
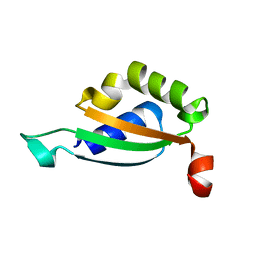 | |
1FVS
 
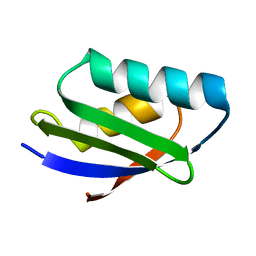 | | SOLUTION STRUCTURE OF THE YEAST COPPER TRANSPORTER DOMAIN CCC2A IN THE APO AND CU(I) LOAD STATES | | Descriptor: | COPPER (II) ION, COPPER-TRANSPORTING ATPASE | | Authors: | Banci, L, Bertini, I, Ciofi Baffoni, S, Huffman, D.L, O'Halloran, T.V. | | Deposit date: | 2000-09-20 | | Release date: | 2001-03-14 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the yeast copper transporter domain Ccc2a in the apo and Cu(I)-loaded states.
J.Biol.Chem., 276, 2001
|
|
1FVQ
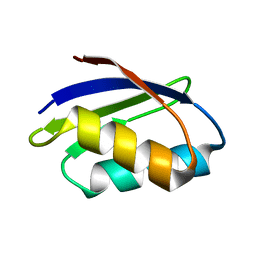 
 | | SOLUTION STRUCTURE OF THE YEAST COPPER TRANSPORTER DOMAIN CCC2A IN THE APO AND CU(I) LOADED STATES | | Descriptor: | COPPER-TRANSPORTING ATPASE | | Authors: | Banci, L, Bertini, I, Ciofi Baffoni, S, Huffman, D.L, O'Halloran, T.V. | | Deposit date: | 2000-09-20 | | Release date: | 2001-03-14 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the yeast copper transporter domain Ccc2a in the apo and Cu(I)-loaded states.
J.Biol.Chem., 276, 2001
|
|
8OKB
 
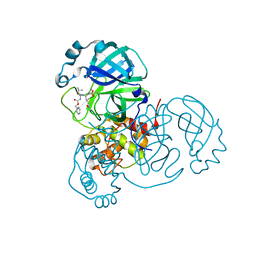 | | SARS-CoV2 NSP5 in complex with a peptidomimetic ligand | | Descriptor: | 3C-like proteinase nsp5, methyl (4~{S})-4-[[(2~{S})-4-methyl-2-(phenylmethoxycarbonylamino)pentanoyl]amino]-5-[(3~{S})-2-oxidanylidenepyrrolidin-3-yl]pentanoate | | Authors: | Calderone, V. | | Deposit date: | 2023-03-28 | | Release date: | 2024-01-17 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Development of a GC-376 Based Peptidomimetic PROTAC as a Degrader of 3-Chymotrypsin-like Protease of SARS-CoV-2.
Acs Med.Chem.Lett., 15, 2024
|
|
8OKC
 
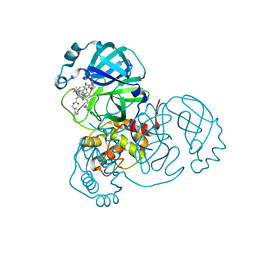 | | SARS-CoV2 NSP5 in complex with a GC-376 based peptidomimetic PROTAC | | Descriptor: | (phenylmethyl) ~{N}-[(2~{R})-1-[[(~{Z},2~{S})-5-[4-[[1-[2-[(3~{R})-2,6-bis(oxidanylidene)piperidin-3-yl]-6-fluoranyl-1,3-bis(oxidanylidene)isoindol-5-yl]piperidin-4-yl]methyl]piperazin-1-yl]-5-oxidanylidene-1-[(3~{R})-2-oxidanylidenepyrrolidin-3-yl]pent-3-en-2-yl]amino]-4-methyl-1-oxidanylidene-pentan-2-yl]carbamate, 3C-like proteinase nsp5 | | Authors: | Calderone, V. | | Deposit date: | 2023-03-28 | | Release date: | 2024-01-17 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Development of a GC-376 Based Peptidomimetic PROTAC as a Degrader of 3-Chymotrypsin-like Protease of SARS-CoV-2.
Acs Med.Chem.Lett., 15, 2024
|
|
2K3J
 
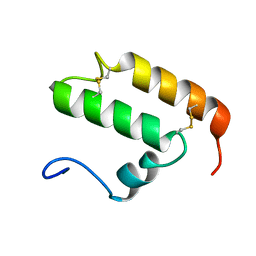 | | The solution structure of human Mia40 | | Descriptor: | Mitochondrial intermembrane space import and assembly protein 40 | | Authors: | Ciofi Baffoni, S, Bertini, I, Gallo, A. | | Deposit date: | 2008-05-08 | | Release date: | 2009-02-10 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | MIA40 is an oxidoreductase that catalyzes oxidative protein folding in mitochondria.
Nat.Struct.Mol.Biol., 16, 2009
|
|
