2KVQ
 
 | |
2LCL
 
 | |
5OWJ
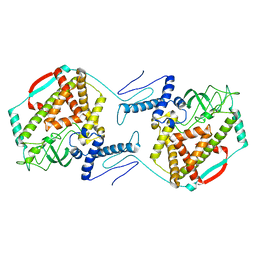 
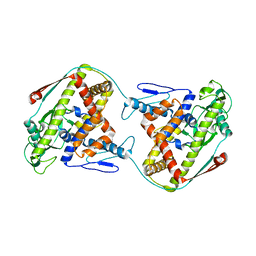
 | | The dynamic dimer structure of the chaperone Trigger Factor (conformer 2) | | Descriptor: | Trigger factor | | Authors: | Morgado, L, Burmann, B.M, Sharpe, T, Mazur, A, Hiller, S. | | Deposit date: | 2017-09-01 | | Release date: | 2017-11-29 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The dynamic dimer structure of the chaperone Trigger Factor.
Nat Commun, 8, 2017
|
|
5OWI
 
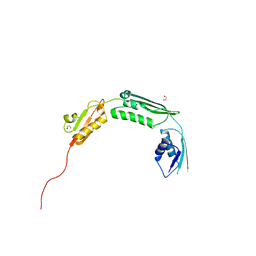
 | | The dynamic dimer structure of the chaperone Trigger Factor (conformer 1) | | Descriptor: | Trigger factor | | Authors: | Morgado, L, Burmann, B.M, Sharpe, T, Mazur, A, Hiller, S. | | Deposit date: | 2017-09-01 | | Release date: | 2017-11-29 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The dynamic dimer structure of the chaperone Trigger Factor.
Nat Commun, 8, 2017
|
|
4BZA
 
 | | Crystal structure of TamA POTRA domains 1-3 from E. coli | | Descriptor: | 1,2-ETHANEDIOL, TRANSLOCATION AND ASSEMBLY MODULE TAMA | | Authors: | Jakob, R.P, Gruss, F, Zaehringer, F, Burmann, B.M, Hiller, S, Maier, T. | | Deposit date: | 2013-07-24 | | Release date: | 2013-09-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.839 Å) | | Cite: | The Structural Basis of Autotransporter Translocation by Tama
Nat.Struct.Mol.Biol., 20, 2013
|
|
4C00
 
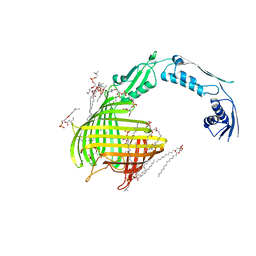 | | Crystal structure of TamA from E. coli | | Descriptor: | 1,2-DIMYRISTOYL-RAC-GLYCERO-3-PHOSPHOCHOLINE, ACETATE ION, CHLORIDE ION, ... | | Authors: | Gruss, F, Zaehringer, F, Jakob, R.P, Burmann, B.M, Hiller, S, Maier, T. | | Deposit date: | 2013-07-30 | | Release date: | 2013-09-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | The Structural Basis of Autotransporter Translocation by Tama
Nat.Struct.Mol.Biol., 20, 2013
|
|
3IMQ
 
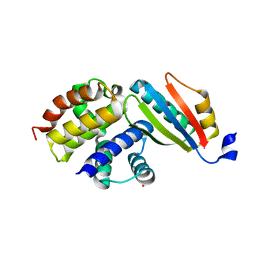 | | Crystal structure of the NusB101-S10(delta loop) complex | | Descriptor: | 30S ribosomal protein S10, N utilization substance protein B, POTASSIUM ION | | Authors: | Luo, X, Wahl, M.C. | | Deposit date: | 2009-08-11 | | Release date: | 2009-11-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Fine tuning of the E. coli NusB:NusE complex affinity to BoxA RNA is required for processive antitermination.
Nucleic Acids Res., 38, 2010
|
|
2LQ8
 
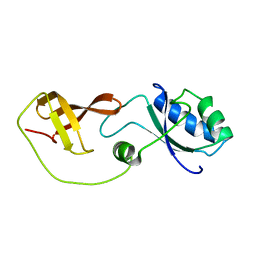 | | Domain interaction in Thermotoga maritima NusG | | Descriptor: | Transcription antitermination protein nusG | | Authors: | Droegemueller, J, Stegmann, C, Burmann, B, Roesch, P, Wahl, M.C, Schweimer, K. | | Deposit date: | 2012-02-27 | | Release date: | 2013-01-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | An Autoinhibited State in the Structure of Thermotoga maritima NusG.
Structure, 21, 2013
|
|
6YHZ
 
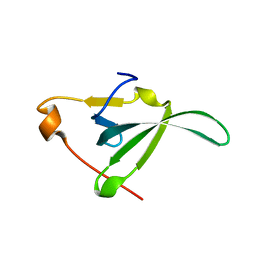 | |
6YI2
 
 | |
2XHC
 
 | |
2XHA
 
 | |
4WO7
 
 | |
5CMT
 
 | |
5CGL
 
 | |
5CKL
 
 | |
9HG6
 
 | | BAM-SurA complex in the swing-in state | | Descriptor: | Chaperone SurA, Outer membrane protein assembly factor BamA, Outer membrane protein assembly factor BamB, ... | | Authors: | Lehner, P.A, Jakob, R.P, Hiller, S. | | Deposit date: | 2024-11-19 | | Release date: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (2.73 Å) | | Cite: | Architecture and conformational dynamics of the BAM-SurA holo insertase complex.
Sci Adv, 11, 2025
|
|
9HG7
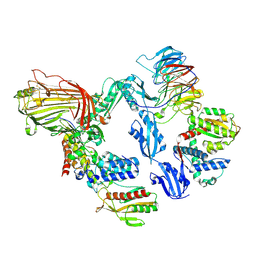 
 | | BAM-SurA complex in the swing-out state | | Descriptor: | Chaperone SurA, Outer membrane protein assembly factor BamA, Outer membrane protein assembly factor BamB, ... | | Authors: | Lehner, P.A, Jakob, R.P, Hiller, S. | | Deposit date: | 2024-11-19 | | Release date: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (2.83 Å) | | Cite: | Architecture and conformational dynamics of the BAM-SurA holo insertase complex.
Sci Adv, 11, 2025
|
|
9HG8
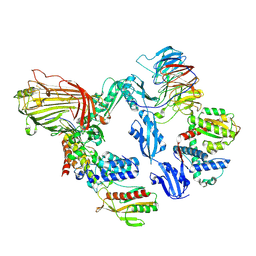 
 | | BAM-SurA complex in the swing-out state with BamC resolved | | Descriptor: | Chaperone SurA, Outer membrane protein assembly factor BamA, Outer membrane protein assembly factor BamB, ... | | Authors: | Lehner, P.A, Jakob, R.P, Hiller, S. | | Deposit date: | 2024-11-19 | | Release date: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Architecture and conformational dynamics of the BAM-SurA holo insertase complex.
Sci Adv, 11, 2025
|
|
9HGA
 
 | | BAM-SurA-darobactin complex in the swing-out state | | Descriptor: | Chaperone SurA, Darobactin A, Outer membrane protein assembly factor BamA, ... | | Authors: | Lehner, P.A, Jakob, R.P, Hiller, S. | | Deposit date: | 2024-11-19 | | Release date: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (3.62 Å) | | Cite: | Architecture and conformational dynamics of the BAM-SurA holo insertase complex.
Sci Adv, 11, 2025
|
|
9HG5
 
 | | BAM-SurA complex in the swing-in state with full length BamC resolved | | Descriptor: | Chaperone SurA, Outer membrane protein assembly factor BamA, Outer membrane protein assembly factor BamB, ... | | Authors: | Lehner, P.A, Jakob, R.P, Hiller, S. | | Deposit date: | 2024-11-19 | | Release date: | 2025-04-16 | | Last modified: | 2025-07-09 | | Method: | ELECTRON MICROSCOPY (2.82 Å) | | Cite: | Architecture and conformational dynamics of the BAM-SurA holo insertase complex.
Sci Adv, 11, 2025
|
|
9HG9
 
 | | BAM-SurA-darobactin complex in the swing-in state | | Descriptor: | Chaperone SurA, Darobactin A, MAGNESIUM ION, ... | | Authors: | Lehner, P.A, Jakob, R.P, Hiller, S. | | Deposit date: | 2024-11-19 | | Release date: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (2.88 Å) | | Cite: | Architecture and conformational dynamics of the BAM-SurA holo insertase complex.
Sci Adv, 11, 2025
|
|
