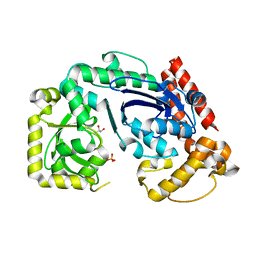
1FXX
 
 | |
3HRU
 
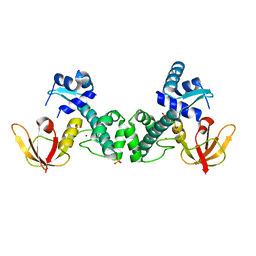 | | Crystal Structure of ScaR with bound Zn2+ | | Descriptor: | Metalloregulator ScaR, SULFATE ION, ZINC ION | | Authors: | Stoll, K.E, Draper, W.E, Kliegman, J.I, Golynskiy, M.V, Brew-Appiah, R.A.T, Brown, H.K, Breyer, W.A, Jakubovics, N.S, Jenkinson, H.F, Brennan, R.B, Cohen, S.M, Glasfeld, A. | | Deposit date: | 2009-06-09 | | Release date: | 2009-06-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Characterization and structure of the manganese-responsive transcriptional regulator ScaR.
Biochemistry, 48, 2009
|
|
3HRT
 
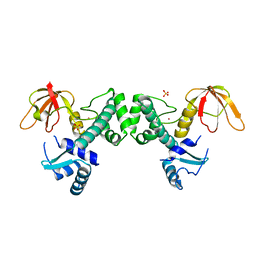 | | Crystal Structure of ScaR with bound Cd2+ | | Descriptor: | CADMIUM ION, Metalloregulator ScaR, SULFATE ION | | Authors: | Stoll, K.E, Draper, W.E, Kliegman, J.I, Golynskiy, M.V, Brew-Appiah, R.A.T, Brown, H.K, Breyer, W.A, Jakubovics, N.S, Jenkinson, H.F, Brennan, R.B, Cohen, S.M, Glasfeld, A. | | Deposit date: | 2009-06-09 | | Release date: | 2009-06-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Characterization and structure of the manganese-responsive transcriptional regulator ScaR.
Biochemistry, 48, 2009
|
|
3HRS
 
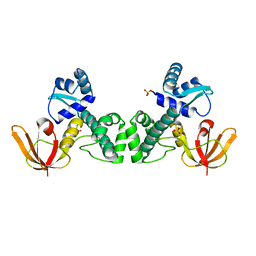 | | Crystal Structure of the Manganese-activated Repressor ScaR: apo form | | Descriptor: | Metalloregulator ScaR, SULFATE ION | | Authors: | Stoll, K.E, Draper, W.E, Kliegman, J.I, Golynskiy, M.V, Brew-Appiah, R.A.T, Brown, H.K, Breyer, W.A, Jakubovics, N.S, Jenkinson, H.F, Brennan, R.B, Cohen, S.M, Glasfeld, A. | | Deposit date: | 2009-06-09 | | Release date: | 2009-06-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Characterization and structure of the manganese-responsive transcriptional regulator ScaR.
Biochemistry, 48, 2009
|
|
1L5T
 
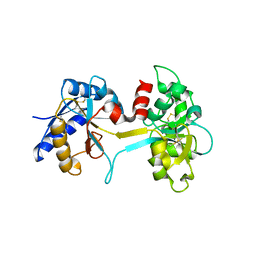 | | Crystal Structure of a Domain-Opened Mutant (R121D) of the Human Lactoferrin N-lobe Refined From a Merohedrally-Twinned Crystal Form. | | Descriptor: | lactoferrin | | Authors: | Jameson, G.B, Anderson, B.F, Breyer, W.A, Tweedie, J.W, Baker, E.N. | | Deposit date: | 2002-03-07 | | Release date: | 2002-03-27 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of a domain-opened mutant (R121D) of the human lactoferrin N-lobe refined from a merohedrally twinned crystal form.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
5CVI
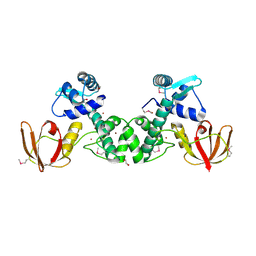 
 | | Structure of the manganese regulator SloR | | Descriptor: | 1,2-ETHANEDIOL, SloR, ZINC ION | | Authors: | Nye, D, Glasfeld, A. | | Deposit date: | 2015-07-27 | | Release date: | 2015-08-12 | | Last modified: | 2020-02-26 | | Method: | X-RAY DIFFRACTION (2.804 Å) | | Cite: | SloR structure and DNA binding studies inform the SloR-SRE interaction in Streptococcus mutans
to be published
|
|
1B6I
 
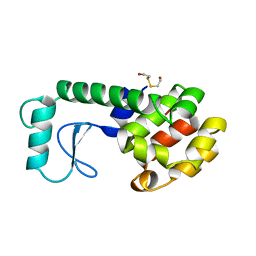 | | T4 LYSOZYME MUTANT WITH CYS 54 REPLACED BY THR, CYS 97 REPLACED BY ALA, THR 21 REPLACED BY CYS AND LYS 124 REPLACED BY CYS (C54T,C97A,T21C,K124C) | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, PROTEIN (LYSOZYME) | | Authors: | Vetter, I.R, Baase, W.A, Snow, S, Matthews, B.W. | | Deposit date: | 1999-01-14 | | Release date: | 2000-01-12 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Solid-state synthesis and mechanical unfolding of polymers of T4 lysozyme.
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
1C6E
 | |
1C61
 
 | |
1C60
 
 | |
1C62
 | |
1C65
 | |
1C67
 | |
1C64
 | |
1C69
 | |
1C6B
 | |
1C63
 | |
1C68
 | |
1C6C
 | |
1C6D
 | |
1C6F
 | |
1C6H
 | |
1C6G
 | |
1C6I
 | |
1C6J
 | |
