1KTG
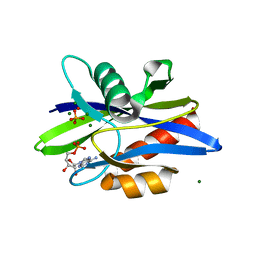 
 | | Crystal Structure of a C. elegans Ap4A Hydrolase Binary Complex | | Descriptor: | ADENOSINE MONOPHOSPHATE, Diadenosine Tetraphosphate Hydrolase, HYDROXIDE ION, ... | | Authors: | Bailey, S, Sedelnikova, S.E, Blackburn, G.M, Abdelghany, H.M, Baker, P.J, McLennan, A.G, Rafferty, J.B. | | Deposit date: | 2002-01-16 | | Release date: | 2002-05-08 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The crystal structure of diadenosine tetraphosphate hydrolase from Caenorhabditis elegans in free and binary complex forms
Structure, 10, 2002
|
|
1KT9
 
 | | Crystal Structure of C. elegans Ap4A Hydrolase | | Descriptor: | Diadenosine Tetraphosphate Hydrolase | | Authors: | Bailey, S, Sedelnikova, S.E, Blackburn, G.M, Abdelghany, H.M, Baker, P.J, McLennan, A.G, Rafferty, J.B. | | Deposit date: | 2002-01-15 | | Release date: | 2002-05-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | The crystal structure of diadenosine tetraphosphate hydrolase from Caenorhabditis elegans in free and binary complex forms
Structure, 10, 2002
|
|
1TYT
 
 | |
1NO1
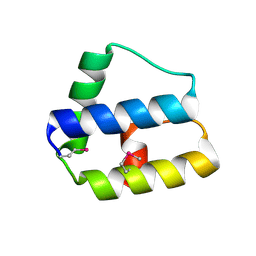 
 | | Structure of truncated variant of B.subtilis SPP1 phage G39P helicase loader/inhibitor protein | | Descriptor: | replisome organizer | | Authors: | Bailey, S, Sedelnikova, S.E, Mesa, P, Ayora, S, Waltho, J.P, Ashcroft, A.E, Baron, A.J, Alonso, J.C, Rafferty, J.B. | | Deposit date: | 2003-01-15 | | Release date: | 2003-05-06 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural analysis of Bacillus subtilis SPP1 phage helicase loader protein G39P
J.Biol.Chem., 278, 2003
|
|
2CA3
 
 | | Sulfite dehydrogenase from Starkeya Novella r55m mutant | | Descriptor: | (MOLYBDOPTERIN-S,S)-OXO-MOLYBDENUM, HEME C, SULFATE ION, ... | | Authors: | Bailey, S, Kappler, U, Feng, C, Honeychurch, M.J, Bernhardt, P.V, Tollin, G, Enemark, J.H. | | Deposit date: | 2005-12-16 | | Release date: | 2007-02-20 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Molecular basis for enzymatic sulfite oxidation: how three conserved active site residues shape enzyme activity.
J.Biol.Chem., 284, 2009
|
|
2CA4
 
 | | Sulfite dehydrogenase from Starkeya Novella mutant | | Descriptor: | (MOLYBDOPTERIN-S,S)-OXO-MOLYBDENUM, HEME C, SULFITE:CYTOCHROME C OXIDOREDUCTASE SUBUNIT A, ... | | Authors: | Bailey, S, Kappler, U. | | Deposit date: | 2005-12-16 | | Release date: | 2007-02-20 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Molecular Basis for Enzymatic Sulfite Oxidation: How Three Conserved Active Site Residues Shape Enzyme Activity.
J.Biol.Chem., 284, 2009
|
|
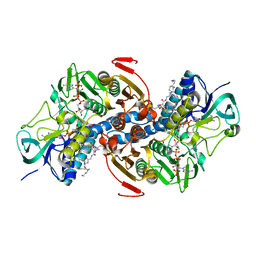
1TYP
 | |
7D6B
 
 | | Crystal structure of Oryza sativa Os4BGlu18 monolignol beta-glucosidase with delta-gluconolactone | | Descriptor: | Beta-glucosidase 18, D-glucono-1,5-lactone, GLYCEROL, ... | | Authors: | Baiya, S, Pengthaisong, S, Ketudat Cairns, J.R. | | Deposit date: | 2020-09-29 | | Release date: | 2021-01-13 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural analysis of rice Os4BGlu18 monolignol beta-glucosidase.
Plos One, 16, 2021
|
|
7D6A
 
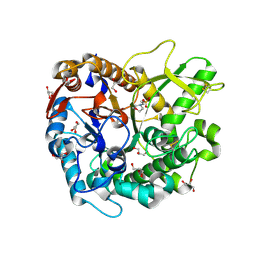 | | Crystal structure of Oryza sativa Os4BGlu18 monolignol beta-glucosidase | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Beta-glucosidase 18, GLYCEROL, ... | | Authors: | Baiya, S, Pengthaisong, S, Ketudat Cairns, J.R. | | Deposit date: | 2020-09-29 | | Release date: | 2021-01-13 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural analysis of rice Os4BGlu18 monolignol beta-glucosidase.
Plos One, 16, 2021
|
|
2HPI
 
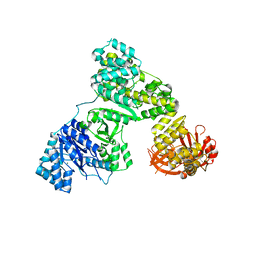 | | Eubacterial and Eukaryotic Replicative DNA Polymerases are not Homologous: X-ray Structure of DNA Polymerase III | | Descriptor: | CHLORIDE ION, DNA polymerase III alpha subunit, MAGNESIUM ION, ... | | Authors: | Bailey, S, Wing, R.A, Steitz, T.A. | | Deposit date: | 2006-07-17 | | Release date: | 2006-09-19 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The Structure of T. aquaticus DNA Polymerase III Is Distinct from Eukaryotic Replicative DNA Polymerases.
Cell(Cambridge,Mass.), 126, 2006
|
|
2HPM
 
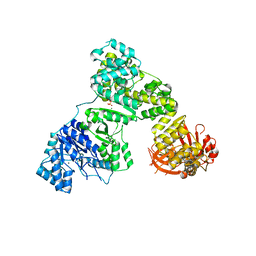 | | Eubacterial and Eukaryotic Replicative DNA Polymerases are not Homologous: X-ray Structure of DNA Polymerase III | | Descriptor: | CHLORIDE ION, DNA Polymerase III alpha subunit, MAGNESIUM ION, ... | | Authors: | Bailey, S, Wing, R.A, Steitz, T.A. | | Deposit date: | 2006-07-17 | | Release date: | 2006-09-19 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | The Structure of T. aquaticus DNA Polymerase III Is Distinct from Eukaryotic Replicative DNA Polymerases.
Cell(Cambridge,Mass.), 126, 2006
|
|
2R6C
 
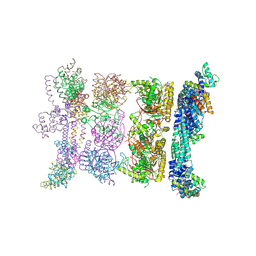 | | Crystal Form BH2 | | Descriptor: | DnaG Primase, Helicase Binding Domain, Replicative helicase | | Authors: | Bailey, S, Eliason, W.K, Steitz, T.A. | | Deposit date: | 2007-09-05 | | Release date: | 2008-06-10 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (4 Å) | | Cite: | Structure of hexameric DnaB helicase and its complex with a domain of DnaG primase
Science, 318, 2007
|
|
2R6A
 
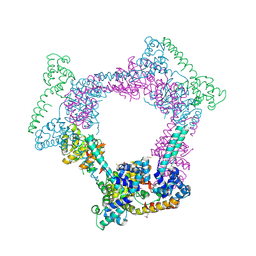 | | Crystal Form BH1 | | Descriptor: | DnaG Primase, Helicase Binding Domain, Replicative helicase, ... | | Authors: | Bailey, S, Eliason, W.K, Steitz, T.A. | | Deposit date: | 2007-09-05 | | Release date: | 2007-11-06 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structure of hexameric DnaB helicase and its complex with a domain of DnaG primase
Science, 318, 2007
|
|
2R6E
 
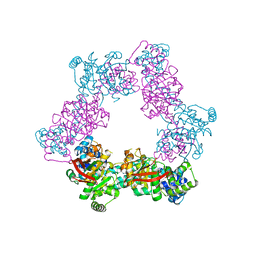 | | Crystal Form B2 | | Descriptor: | Replicative helicase, SULFATE ION | | Authors: | Bailey, S, Eliason, W.K, Steitz, T.A. | | Deposit date: | 2007-09-05 | | Release date: | 2007-11-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (5.019 Å) | | Cite: | Structure of hexameric DnaB helicase and its complex with a domain of DnaG primase
Science, 318, 2007
|
|
2R6D
 
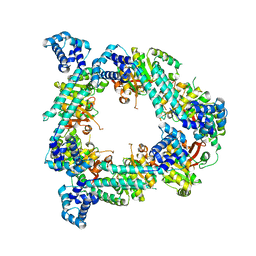 | | Crystal Form B1 | | Descriptor: | Replicative helicase | | Authors: | Bailey, S, Eliason, W.K, Steitz, T.A. | | Deposit date: | 2007-09-05 | | Release date: | 2008-06-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | Structure of hexameric DnaB helicase and its complex with a domain of DnaG primase
Science, 318, 2007
|
|
2CC3
 
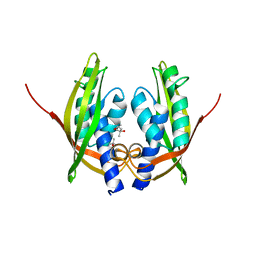 | | Structure of Agrobacterium tumefaciens VirB8 protein | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, PROTEIN VIRB8 | | Authors: | Bailey, S, Ward, D, Middleton, R, Grossmann, G, Zambryski, P.C. | | Deposit date: | 2006-01-11 | | Release date: | 2006-01-30 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Agrobacterium Tumefaciens Virb8 Structure Reveals Potential Protein-Protein Interactions Sites.
Proc.Natl.Acad.Sci.USA, 103, 2006
|
|
2Q6T
 
 | |
3SKD
 
 | |
4F3E
 
 | |
2BLF
 
 | | Sulfite dehydrogenase from Starkeya Novella | | Descriptor: | (MOLYBDOPTERIN-S,S)-OXO-MOLYBDENUM, HEME C, SULFATE ION, ... | | Authors: | Bailey, S, Kappler, U. | | Deposit date: | 2005-03-04 | | Release date: | 2005-04-28 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Molecular Basis of Intramolecular Electron Transfer in Sulfite-Oxidizing Enzymes is Revealed by High Resolution Structure of a Heterodimeric Complex of the Catalytic Molybdopterin Subunit and a C-Type Cytochrome Subunit.
J.Biol.Chem., 280, 2005
|
|
2BPB
 
 | | Sulfite dehydrogenase from Starkeya Novella | | Descriptor: | (MOLYBDOPTERIN-S,S)-OXO-MOLYBDENUM, HEME C, SULFITE:CYTOCHROME C OXIDOREDUCTASE SUBUNIT A, ... | | Authors: | Bailey, S, Kappler, U. | | Deposit date: | 2005-04-19 | | Release date: | 2005-04-28 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular Basis of Intramolecular Electron Transfer in Sulfite-Oxidizing Enzymes is Revealed by High Resolution Structure of a Heterodimeric Complex of the Catalytic Molybdopterin Subunit and a C-Type Cytochrome Subunit.
J.Biol.Chem., 280, 2005
|
|
2C9X
 
 | | Sulfite dehydrogenase from Starkeya Novella Y236F mutant | | Descriptor: | (MOLYBDOPTERIN-S,S)-OXO-MOLYBDENUM, HEME C, SULFATE ION, ... | | Authors: | Bailey, S, Kappler, U, Feng, C, Honeychurch, M.J, Bernhardt, P, Tollin, G, Enemark, J. | | Deposit date: | 2005-12-15 | | Release date: | 2006-08-16 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Kinetic and Structural Evidence for the Importance of Tyr236 for the Integrity of the Mo Active Site in a Bacterial Sulfite Dehydrogenase.
Biochemistry, 45, 2006
|
|
3SK9
 
 | | Crystal structure of the Thermus thermophilus cas3 HD domain | | Descriptor: | Putative uncharacterized protein TTHB187 | | Authors: | Bailey, S, Mulepati, S. | | Deposit date: | 2011-06-22 | | Release date: | 2011-07-20 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and Biochemical Analysis of Nuclease Domain of Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR)-associated Protein 3 (Cas3).
J.Biol.Chem., 286, 2011
|
|
1H5N
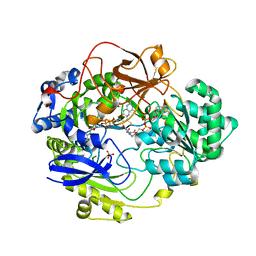 
 | | DMSO REDUCTASE MODIFIED BY THE PRESENCE OF DMS AND AIR | | Descriptor: | 2-AMINO-5,6-DIMERCAPTO-7-METHYL-3,7,8A,9-TETRAHYDRO-8-OXA-1,3,9,10-TETRAAZA-ANTHRACEN-4-ONE GUANOSINE DINUCLEOTIDE, Dimethyl sulfoxide/trimethylamine N-oxide reductase, MOLYBDENUM(VI) ION, ... | | Authors: | Bailey, S, Adams, B, Smith, A.T, Richards, R.L, Lowe, D.J, Bray, R.C. | | Deposit date: | 2001-05-22 | | Release date: | 2002-05-17 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Reactions of Dimethyl Sulphoxide Reductase in the Presence of Dimethylsulphide and the Structure of the Dimethylsulphide-Modified Enzyme
Biochemistry, 40, 2001
|
|
1E5V
 
 | | OXIDIZED DMSO REDUCTASE EXPOSED TO HEPES BUFFER | | Descriptor: | 2-AMINO-5,6-DIMERCAPTO-7-METHYL-3,7,8A,9-TETRAHYDRO-8-OXA-1,3,9,10-TETRAAZA-ANTHRACEN-4-ONE GUANOSINE DINUCLEOTIDE, Dimethyl sulfoxide/trimethylamine N-oxide reductase, MOLYBDENUM (IV)OXIDE, ... | | Authors: | Bailey, S, Bennett, B, Adams, B, Smith, A.T, Bray, R.C. | | Deposit date: | 2000-08-03 | | Release date: | 2000-08-25 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Reversible Dissociation of Thiolate Ligands from Molybdenum in an Enzyme of the Dimethyl Sulfoxide Reductase Family
Biochemistry, 39, 2000
|
|
