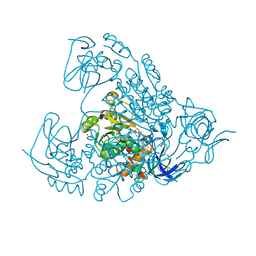
2D8A
 
 | |
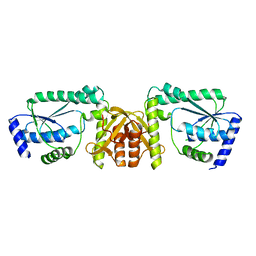
2DPL
 
 | |
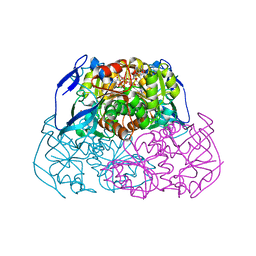
2EER
 
 | |
2EKC
 
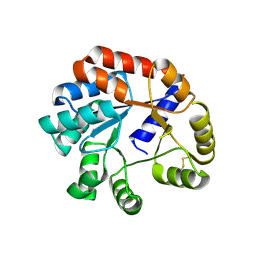 | |
2CU0
 | |
2EKD
 
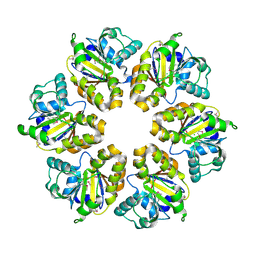 | |
2EP5
 
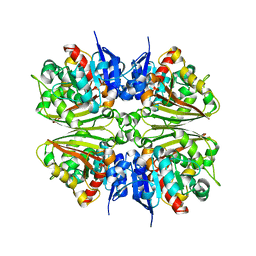 | |
2E8S
 
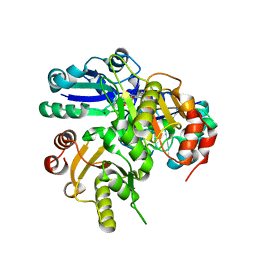 | | Structural study of Project ID PH0725 from Pyrococcus horikoshii OT3 | | Descriptor: | Probable diphthine synthase, S-ADENOSYL-L-HOMOCYSTEINE, SODIUM ION | | Authors: | Asada, Y, Shimada, H, Taketa, M, Nakamoto, T, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-01-23 | | Release date: | 2007-07-24 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural study of Project ID PH0725 from Pyrococcus horikoshii OT3
To be Published
|
|
2E8Q
 
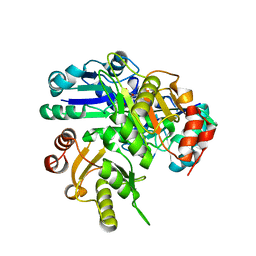 | | Structural study of Project ID PH0725 from Pyrococcus horikoshii OT3 (K19M) | | Descriptor: | Probable diphthine synthase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Asada, Y, Shimada, H, Taketa, M, Nakamoto, T, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-01-23 | | Release date: | 2007-07-24 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural study of Project ID PH0725 from Pyrococcus horikoshii OT3 (K19M)
To be Published
|
|
2EP7
 
 | |
2DPN
 
 | |
1J33
 
 | |
1UAY
 
 | | Crystal Structure of Type II 3-Hydroxyacyl-CoA Dehydrogenase from Thermus thermophilus HB8 | | Descriptor: | ADENOSINE, Type II 3-hydroxyacyl-CoA dehydrogenase | | Authors: | Kunishima, N, Asada, Y, Yokoyama, S, Kuramitsu, S, Miyano, M, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-03-25 | | Release date: | 2003-04-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Crystal Structure of Type II 3-Hydroxyacyl-CoA Dehydrogenase from Thermus thermophilus HB8
To be Published
|
|
2HD9
 
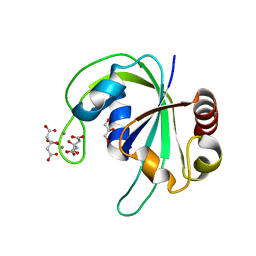 | | Crystal structure of PH1033 from Pyrococcus horikoshii OT3 | | Descriptor: | CALCIUM ION, CITRIC ACID, GLYCEROL, ... | | Authors: | Sugahara, M, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-06-20 | | Release date: | 2006-12-20 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Nucleant-mediated protein crystallization with the application of microporous synthetic zeolites.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
5B3T
 | |
5B3U
 
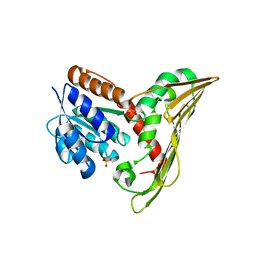 | |
5B3V
 
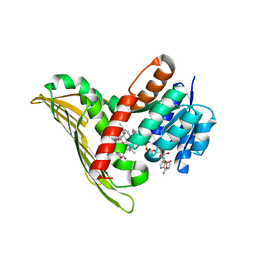 | |
2POQ
 
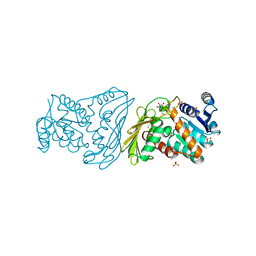 | | Dimeric Dihydrodiol Dehydrogenase complexed with inhibitor, Isoascorbic acid | | Descriptor: | BETA-MERCAPTOETHANOL, Dimeric dihydrodiol dehydrogenase, ISOASCORBIC ACID, ... | | Authors: | Carbone, V, El-Kabbani, O. | | Deposit date: | 2007-04-27 | | Release date: | 2007-07-31 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Structure of monkey dimeric dihydrodiol dehydrogenase in complex with isoascorbic acid.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
1J1Y
 
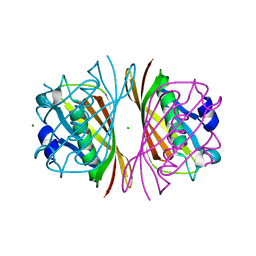 | | Crystal Structure of PaaI from Thermus thermophilus HB8 | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, PaaI protein | | Authors: | Kunishima, N, Sugahara, M, Kuramitsu, S, Yokoyama, S, Miyano, M, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2002-12-24 | | Release date: | 2004-02-17 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A Novel Induced-fit Reaction Mechanism of Asymmetric Hot Dog Thioesterase PaaI
J.Mol.Biol., 352, 2005
|
|
1WMM
 
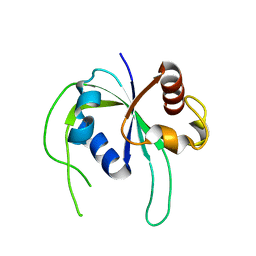 | |
2ZBN
 
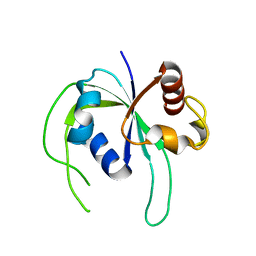 | |
1WLV
 
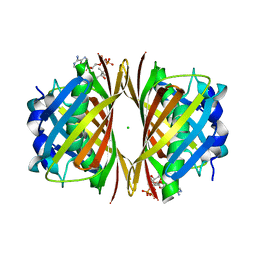 | | Crystal structure of TT0310 protein from Thermus thermophilus HB8 | | Descriptor: | ACETATE ION, CHLORIDE ION, COENZYME A, ... | | Authors: | Kunishima, N, Sugahara, M, Miyano, M, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-06-29 | | Release date: | 2005-07-05 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A Novel Induced-fit Reaction Mechanism of Asymmetric Hot Dog Thioesterase PaaI
J.Mol.Biol., 352, 2005
|
|
1WM6
 
 | |
1WLU
 
 | | Crystal structure of TT0310 protein from Thermus thermophilus HB8 | | Descriptor: | CHLORIDE ION, GLYCEROL, phenylacetic acid degradation protein PaaI | | Authors: | Kunishima, N, Sugahara, M, Miyano, M, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-06-29 | | Release date: | 2005-07-05 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | A Novel Induced-fit Reaction Mechanism of Asymmetric Hot Dog Thioesterase PaaI
J.Mol.Biol., 352, 2005
|
|
1WN3
 
 | | Crystal structure of TT0310 protein from Thermus thermophilus HB8 | | Descriptor: | ACETATE ION, CHLORIDE ION, HEXANOYL-COENZYME A, ... | | Authors: | Kunishima, N, Sugahara, M, Miyano, M, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-07-27 | | Release date: | 2005-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A Novel Induced-fit Reaction Mechanism of Asymmetric Hot Dog Thioesterase PaaI
J.Mol.Biol., 352, 2005
|
|
