6UZL
 
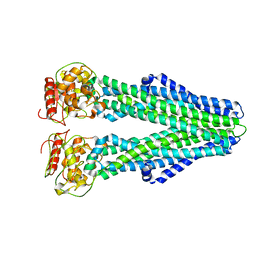 | | Cryo-EM structure of nucleotide-free MsbA reconstituted into peptidiscs, conformation 2 | | Descriptor: | Lipid A export ATP-binding/permease protein MsbA | | Authors: | Angiulli, G, Walz, T, Dhupar, H.S, Suzuki, H, Wason, I.S, Duong Van Hoa, F. | | Deposit date: | 2019-11-15 | | Release date: | 2020-03-04 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | New approach for membrane protein reconstitution into peptidiscs and basis for their adaptability to different proteins.
Elife, 9, 2020
|
|
6UZH
 
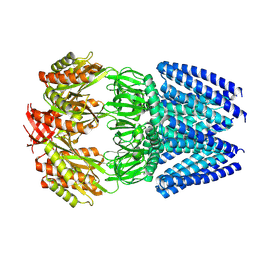 | | Cryo-EM structure of mechanosensitive channel MscS reconstituted into peptidiscs | | Descriptor: | Small-conductance mechanosensitive channel | | Authors: | Angiulli, G, Walz, T, Dhupar, H.S, Suzuki, H, Wason, I.S, Duong Van Hoa, F. | | Deposit date: | 2019-11-15 | | Release date: | 2020-03-04 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | New approach for membrane protein reconstitution into peptidiscs and basis for their adaptability to different proteins.
Elife, 9, 2020
|
|
6UZ2
 
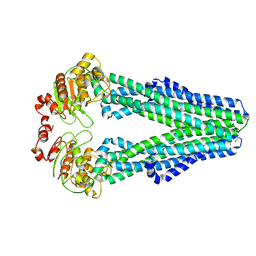 | | Cryo-EM structure of nucleotide-free MsbA reconstituted into peptidiscs, conformation 1 | | Descriptor: | Lipid A export ATP-binding/permease protein MsbA | | Authors: | Angiulli, G, Walz, T, Dhupar, H.S, Suzuki, H, Wason, I.S, Duong Van Hoa, F. | | Deposit date: | 2019-11-14 | | Release date: | 2020-03-04 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | New approach for membrane protein reconstitution into peptidiscs and basis for their adaptability to different proteins.
Elife, 9, 2020
|
|
5EBK
 
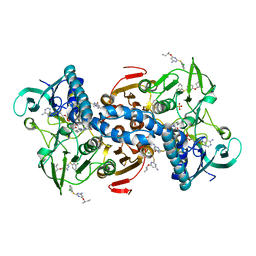 | |
4K1F
 
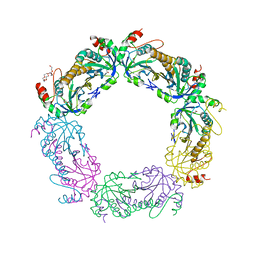 | | Crystal structure of reduced tryparedoxin peroxidase from leishmania major at 2.34 A resolution | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Ilari, A, Fiorillo, A, Di Chiaro, F. | | Deposit date: | 2013-04-05 | | Release date: | 2014-04-09 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Structure-based discovery of the first non-covalent inhibitors of Leishmania major tryparedoxin peroxidase by high throughput docking
Sci Rep, 5, 2015
|
|
