6Q09
 
 | |
6PTR
 
 | |
6Q10
 
 | |
6PTV
 
 | |
6PTH
 
 | |
4WJM
 
 | |
4WNY
 
 | |
4WSO
 
 | |
4XIJ
 
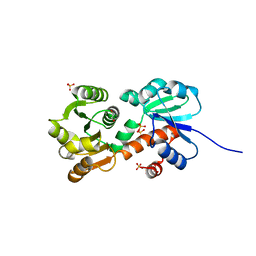 | |
4XK1
 
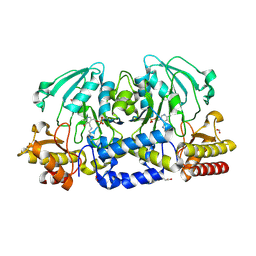 | |
4XDQ
 
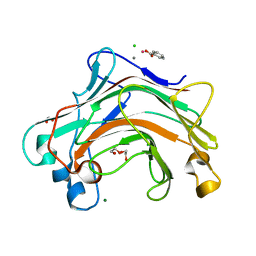 | |
4XUZ
 
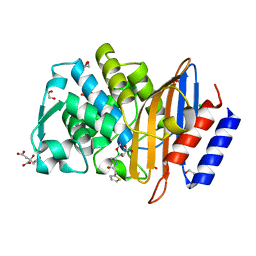 | | Structure of CTX-M-15 bound to RPX-7009 at 1.5 A | | Descriptor: | 1,2-ETHANEDIOL, Beta-lactamase, CHLORIDE ION, ... | | Authors: | Clifton, M.C, Gardberg, A. | | Deposit date: | 2015-01-26 | | Release date: | 2015-04-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Discovery of a Cyclic Boronic Acid beta-Lactamase Inhibitor (RPX7009) with Utility vs Class A Serine Carbapenemases.
J.Med.Chem., 58, 2015
|
|
4XXV
 
 | |
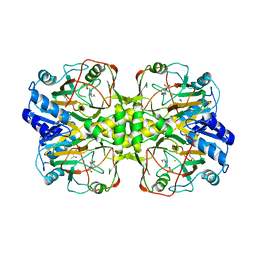
4Y0E
 | |
6WCI
 
 | |
6WNG
 
 | |
6WCT
 
 | |
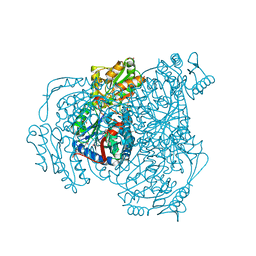
6WSA
 | |
6WSB
 
 | |
6DA0
 
 | |
6XDH
 
 | |
6XRS
 
 | |
3U0A
 
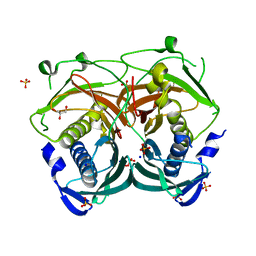 | | Crystal structure of an Acyl-CoA thioesterase II TesB2 from Mycobacterium marinum | | Descriptor: | Acyl-CoA thioesterase II TesB2, GLYCEROL, SODIUM ION, ... | | Authors: | Arakaki, T.L, Staker, B.L, Clifton, M.C, Sankaran, B, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2011-09-28 | | Release date: | 2011-10-05 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Increasing the structural coverage of tuberculosis drug targets.
Tuberculosis (Edinb), 95, 2015
|
|
3TL3
 
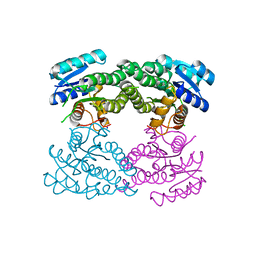 | |
3TX2
 
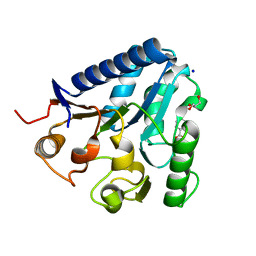 | |
