6F2J
 
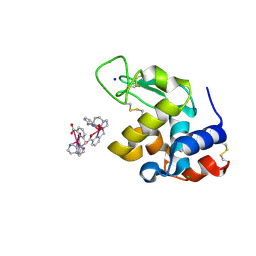 | | Crystal structure of Hen Egg-White Lysozyme co-crystallized in presence of 100 mM Tb-Xo4 and 100 mM sodium sulfate | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Engilberge, S, Riobe, F, Di Pietro, S, Girard, E, Dumont, E, Maury, O. | | Deposit date: | 2017-11-24 | | Release date: | 2018-10-03 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Unveiling the Binding Modes of the Crystallophore, a Terbium-based Nucleating and Phasing Molecular Agent for Protein Crystallography.
Chemistry, 24, 2018
|
|
6F2H
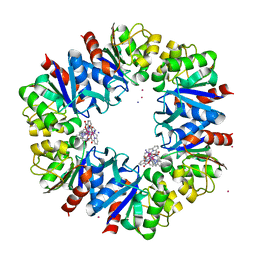 
 | | Structure of Protease 1 from Pyrococcus horikoshii co-crystallized in presence of 10 mM Tb-Xo4 and potassium iodide. | | Descriptor: | Deglycase PH1704, IODIDE ION, TERBIUM(III) ION, ... | | Authors: | Engilberge, S, Riobe, F, Di Pietro, S, Franzetti, B, Girard, E, Dumont, E, Maury, O. | | Deposit date: | 2017-11-24 | | Release date: | 2018-10-03 | | Last modified: | 2019-05-29 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Unveiling the Binding Modes of the Crystallophore, a Terbium-based Nucleating and Phasing Molecular Agent for Protein Crystallography.
Chemistry, 24, 2018
|
|
6FRO
 
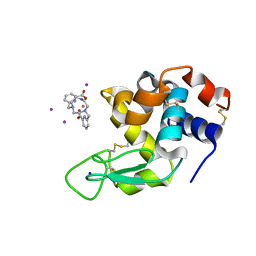 | | Crystal structure of Hen Egg-White Lysozyme co-crystallized in presence of 100 mM Tb-Xo4 and 100 mM potassium iodide. | | Descriptor: | IODIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Engilberge, S, Riobe, F, Di Pietro, S, Girard, E, Dumont, E, Maury, O. | | Deposit date: | 2018-02-16 | | Release date: | 2018-10-03 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Unveiling the Binding Modes of the Crystallophore, a Terbium-based Nucleating and Phasing Molecular Agent for Protein Crystallography.
Chemistry, 24, 2018
|
|
5G0R
 
 | |
2OHH
 
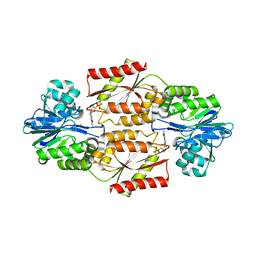 | | Crystal Structure of coenzyme F420H2 oxidase (FprA), a diiron flavoprotein, active oxidized state | | Descriptor: | FE (III) ION, FLAVIN MONONUCLEOTIDE, SULFATE ION, ... | | Authors: | Seedorf, H, Warkentin, E, Ermler, U. | | Deposit date: | 2007-01-10 | | Release date: | 2007-05-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of coenzyme F420H2 oxidase (FprA), a di-iron flavoprotein from methanogenic Archaea catalyzing the reduction of O2 to H2O.
Febs J., 274, 2007
|
|
2OHI
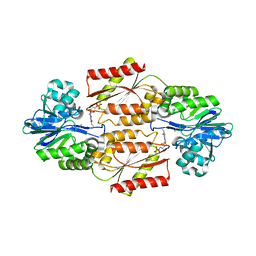 
 | | Crystal Structure of coenzyme F420H2 oxidase (FprA), a diiron flavoprotein, reduced state | | Descriptor: | CHLORIDE ION, FE (III) ION, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Seedorf, H, Warkentin, E, Ermler, U. | | Deposit date: | 2007-01-10 | | Release date: | 2007-05-22 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of coenzyme F420H2 oxidase (FprA), a di-iron flavoprotein from methanogenic Archaea catalyzing the reduction of O2 to H2O.
Febs J., 274, 2007
|
|
2OHJ
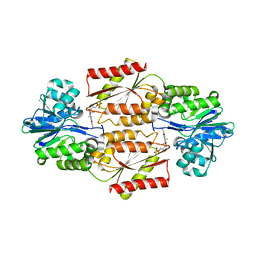 
 | | Crystal Structure of coenzyme F420H2 oxidase (FprA), a diiron flavoprotein, inactive oxidized state | | Descriptor: | CHLORIDE ION, FE (III) ION, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Seedorf, H, Warkentin, E, Ermler, U. | | Deposit date: | 2007-01-10 | | Release date: | 2007-05-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Structure of coenzyme F420H2 oxidase (FprA), a di-iron flavoprotein from methanogenic Archaea catalyzing the reduction of O2 to H2O.
Febs J., 274, 2007
|
|
1HBO
 
 | | METHYL-COENZYME M REDUCTASE MCR-RED1-SILENT | | Descriptor: | 1-THIOETHANESULFONIC ACID, CHLORIDE ION, Coenzyme B, ... | | Authors: | Grabarse, W. | | Deposit date: | 2001-04-20 | | Release date: | 2001-08-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | On the Mechanism of Biological Methane Formation: Structural Evidence for Conformational Changes in Methyl-Coenzyme M Reductase Upon Substrate Binding
J.Mol.Biol., 309, 2001
|
|
1HBN
 
 | | METHYL-COENZYME M REDUCTASE | | Descriptor: | 1-THIOETHANESULFONIC ACID, CHLORIDE ION, Coenzyme B, ... | | Authors: | Ermler, U, Grabarse, W. | | Deposit date: | 2001-04-20 | | Release date: | 2001-08-16 | | Last modified: | 2019-07-24 | | Method: | X-RAY DIFFRACTION (1.16 Å) | | Cite: | On the Mechanism of Biological Methane Formation: Structural Evidence for Conformational Changes in Methyl-Coenzyme M Reductase Upon Substrate Binding
J.Mol.Biol., 309, 2001
|
|
1HBM
 
 | | METHYL-COENZYME M REDUCTASE ENZYME PRODUCT COMPLEX | | Descriptor: | CHLORIDE ION, FACTOR 430, GLYCEROL, ... | | Authors: | Ermler, U, Grabarse, W. | | Deposit date: | 2001-04-20 | | Release date: | 2001-08-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | On the Mechanism of Biological Methane Formation: Structural Evidence for Conformational Changes in Methyl-Coenzyme M Reductase Upon Substrate Binding
J.Mol.Biol., 309, 2001
|
|
5A8K
 
 | |
5A0Y
 
 | |
5A8W
 
 | |
5A8R
 
 | |
