6OTE
 
 | |
6OR9
 
 | |
4TVI
 
 | |
4TWR
 
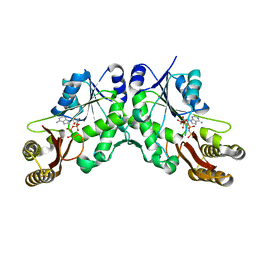
 | | Structure of UDP-glucose 4-epimerase from Brucella abortus | | Descriptor: | NAD binding site:NAD-dependent epimerase/dehydratase:UDP-glucose 4-epimerase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ZINC ION | | Authors: | Horanyi, P.S, Abendroth, J, Lorimer, D, Edwards, T, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2014-07-01 | | Release date: | 2014-10-08 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of UDP-glucose 4-epimerase from Brucella melitensis
To Be Published
|
|
6PHE
 | |
6PI4
 
 | |
6P61
 
 | |
6Q09
 
 | |
6Q1Y
 
 | |
6PTG
 
 | |
6PZJ
 | |
5JAY
 
 | |
5JI5
 | |
5K0T
 
 | |
5JTK
 
 | |
5JRY
 
 | |
5K0S
 
 | |
5KEU
 
 | |
5K9F
 
 | |
5JSC
 
 | |
5KF0
 
 | |
5KIA
 | |
5KHA
 
 | |
5KOI
 
 | |
5KWV
 
 | |
