4RMN
 
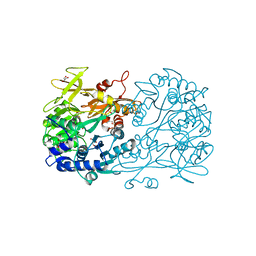 | | Crystal structure of a benzoate coenzyme A ligase with 2-Thiophene Carboxylic acid | | Descriptor: | Benzoate-coenzyme A ligase, GLYCEROL, THIOPHENE-2-CARBOXYLIC ACID | | Authors: | Strom, S, Nosrati, M, Thornburg, C, Walker, K.D, Geiger, J.H. | | Deposit date: | 2014-10-21 | | Release date: | 2015-09-30 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Kinetically and Crystallographically Guided Mutations of a Benzoate CoA Ligase (BadA) Elucidate Mechanism and Expand Substrate Permissivity.
Biochemistry, 54, 2015
|
|
4RUU
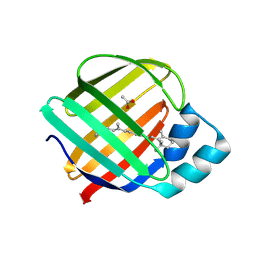 
 | |
4RM2
 
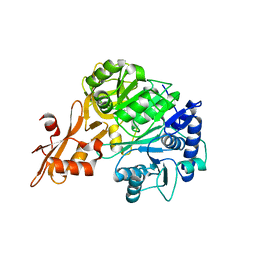 | | Crystal structure of a benzoate coenzyme A ligase with 2-Fluoro benzoic acid | | Descriptor: | 2-fluorobenzoic acid, Benzoate-coenzyme A ligase | | Authors: | Strom, S, Nosrati, M, Thornburg, C, Walker, K.D, Geiger, J.H. | | Deposit date: | 2014-10-18 | | Release date: | 2015-09-30 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Kinetically and Crystallographically Guided Mutations of a Benzoate CoA Ligase (BadA) Elucidate Mechanism and Expand Substrate Permissivity.
Biochemistry, 54, 2015
|
|
4YGH
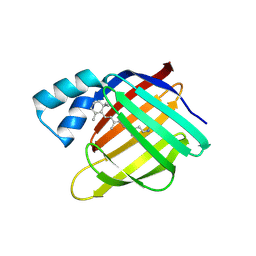 
 | |
4YH0
 | |
4YDA
 | |
4YDB
 | |
4YCE
 | |
4YFQ
 | |
4YFR
 
 | |
4YKM
 
 | |
4YBP
 | |
4YGZ
 | |
4YFP
 | |
4YGG
 | |
4YBU
 | |
4YCH
 | |
4ZJZ
 
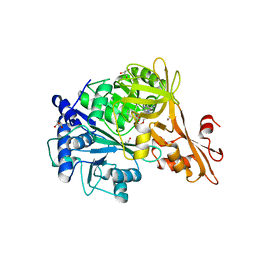 | | Crystal structure of a benzoate coenzyme A ligase with Benzoyl-AMP | | Descriptor: | 5'-O-[(R)-(benzoyloxy)(hydroxy)phosphoryl]adenosine, BENZOIC ACID, Benzoate-coenzyme A ligase, ... | | Authors: | Strom, S, Nosrati, M, Thornburg, C, Walker, K.D, Geiger, J.H. | | Deposit date: | 2015-04-29 | | Release date: | 2015-09-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Kinetically and Crystallographically Guided Mutations of a Benzoate CoA Ligase (BadA) Elucidate Mechanism and Expand Substrate Permissivity.
Biochemistry, 54, 2015
|
|
6ON7
 
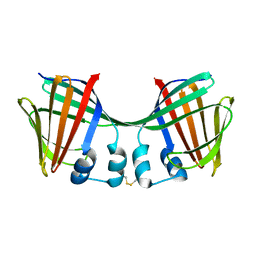 | |
3UNV
 
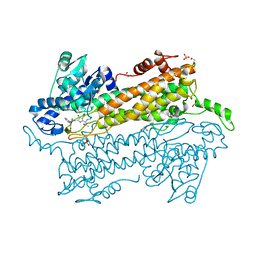 | | Pantoea agglomerans Phenylalanine Aminomutase | | Descriptor: | (3S)-3-amino-3-phenylpropanoic acid, 1,2-ETHANEDIOL, AdmH, ... | | Authors: | Geiger, J, Strom, S. | | Deposit date: | 2011-11-16 | | Release date: | 2012-02-22 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Insights into the mechanistic pathway of the Pantoea agglomerans phenylalanine aminomutase.
Angew.Chem.Int.Ed.Engl., 51, 2012
|
|
2QZS
 
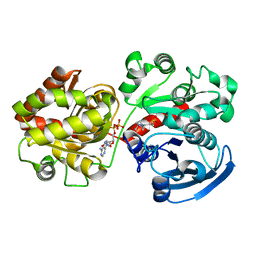 | | Crystal Structure of Wild-type E.coli GS in complex with ADP and Glucose(wtGSb) | | Descriptor: | (2R)-2-hydroxy-3-[4-(2-hydroxyethyl)piperazin-1-yl]propane-1-sulfonic acid, ADENOSINE-5'-DIPHOSPHATE, alpha-D-glucopyranose, ... | | Authors: | Sheng, F, Geiger, J. | | Deposit date: | 2007-08-17 | | Release date: | 2008-09-09 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Crystal Structures of the Open and Catalytically Competent Closed Conformation of Escherichia coli Glycogen Synthase.
J.Biol.Chem., 284, 2009
|
|
2R4U
 
 | | Crystal Structure of Wild-type E.coli GS in complex with ADP and Glucose(wtGSd) | | Descriptor: | (2R)-2-hydroxy-3-[4-(2-hydroxyethyl)piperazin-1-yl]propane-1-sulfonic acid, 3,6,9,12,15,18,21,24,27,30,33,36,39-TRIDECAOXAHENTETRACONTANE-1,41-DIOL, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Sheng, F, Geiger, J. | | Deposit date: | 2007-09-01 | | Release date: | 2008-09-09 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.367 Å) | | Cite: | The crystal structures of the open and catalytically competent closed conformation of Escherichia coli glycogen synthase.
J.Biol.Chem., 284, 2009
|
|
2R4T
 
 | | Crystal Structure of Wild-type E.coli GS in Complex with ADP and Glucose(wtGSc) | | Descriptor: | (2R)-2-hydroxy-3-[4-(2-hydroxyethyl)piperazin-1-yl]propane-1-sulfonic acid, 3,6,9,12,15,18,21,24,27,30,33,36,39-TRIDECAOXAHENTETRACONTANE-1,41-DIOL, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Sheng, F, Geiger, J. | | Deposit date: | 2007-09-01 | | Release date: | 2008-09-09 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.258 Å) | | Cite: | The crystal structures of the open and catalytically competent closed conformation of Escherichia coli glycogen synthase.
J.Biol.Chem., 284, 2009
|
|
6WP0
 
 | |
5DG4
 | | Crystal structure of monomer human cellular retinol binding protein II-Y60L | | Descriptor: | ACETATE ION, Retinol-binding protein 2 | | Authors: | Assar, Z, Nossoni, Z, Wang, W, Gieger, J.H, Borhan, B. | | Deposit date: | 2015-08-27 | | Release date: | 2016-09-21 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Domain-Swapped Dimers of Intracellular Lipid-Binding Proteins: Evidence for Ordered Folding Intermediates.
Structure, 24, 2016
|
|
