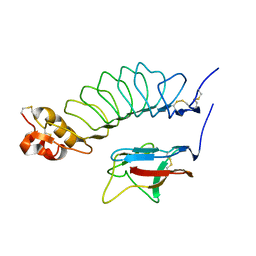
2V9T
 
 | |
1FSS
 
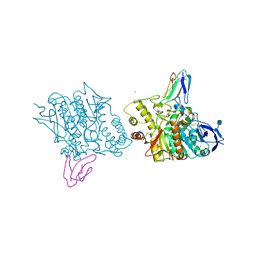 | | ACETYLCHOLINESTERASE (E.C. 3.1.1.7) COMPLEXED WITH FASCICULIN-II | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETYLCHOLINESTERASE, FASCICULIN II, ... | | Authors: | Harel, M, Kleywegt, G.J, Silman, I, Sussman, J.L. | | Deposit date: | 1995-10-25 | | Release date: | 1996-03-08 | | Last modified: | 2021-06-02 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of an acetylcholinesterase-fasciculin complex: interaction of a three-fingered toxin from snake venom with its target.
Structure, 3, 1995
|
|
4F61
 
 | | Tubulin:Stathmin-like domain complex | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Gigant, B, Mignot, I, Knossow, M. | | Deposit date: | 2012-05-14 | | Release date: | 2012-07-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (4.17 Å) | | Cite: | Design and characterization of modular scaffolds for tubulin assembly.
J.Biol.Chem., 287, 2012
|
|
4F6R
 
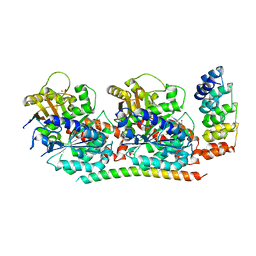 | | Tubulin:Stathmin-like domain complex | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Designed ankyrin repeat protein (DARPIN) D2, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Gigant, B, Mignot, I, Knossow, M. | | Deposit date: | 2012-05-15 | | Release date: | 2012-07-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Design and characterization of modular scaffolds for tubulin assembly.
J.Biol.Chem., 287, 2012
|
|
2BLR
 
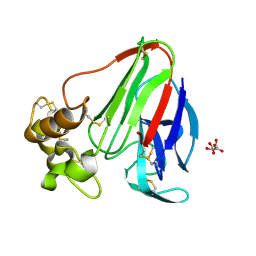 | |
3DE1
 
 | |
3DDZ
 
 | |
3D9Q
 
 | |
3DE2
 
 | |
3DE0
 
 | |
3DE6
 
 | |
3DE3
 
 | |
3DE5
 
 | |
3DE7
 
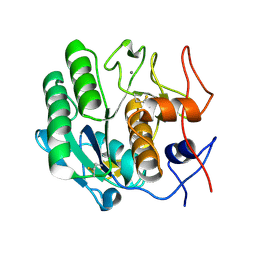 | |
3DE4
 
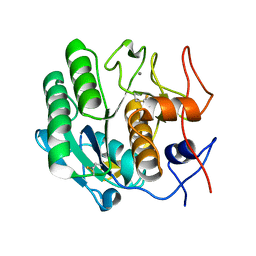 | |
2V9R
 
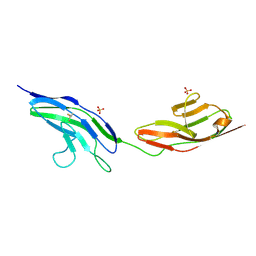 | |
2V9S
 
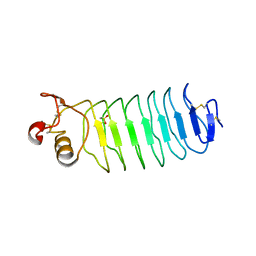 | | Second LRR domain of human Slit2 | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, SLIT HOMOLOG 2 PROTEIN N-PRODUCT | | Authors: | Morlot, C, Cusack, S, McCarthy, A.A. | | Deposit date: | 2007-08-25 | | Release date: | 2007-09-25 | | Last modified: | 2014-11-05 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Insights Into the Slit-Robo Complex.
Proc.Natl.Acad.Sci.USA, 104, 2007
|
|
4IVZ
 
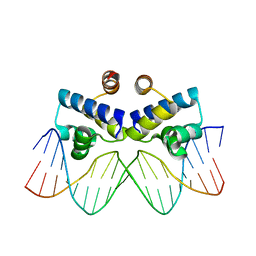 | |
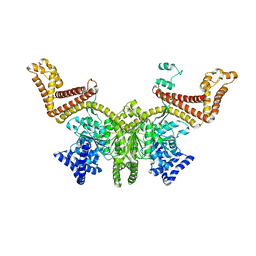
2FSI
 
 | |
2FSF
 | |
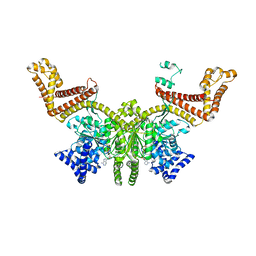
2FSH
 | |
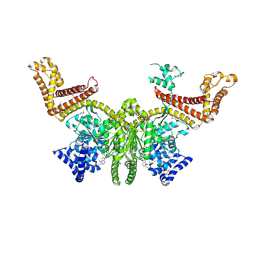
2FSG
 | |
4F8D
 
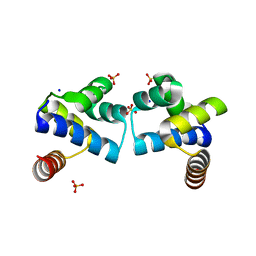 | |
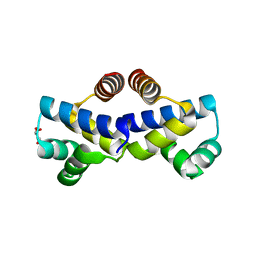
4FBI
 
 | |
5OQV
 
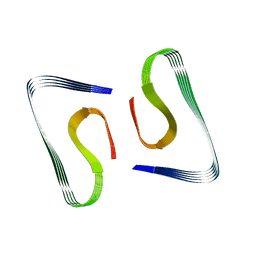 | | Near-atomic resolution fibril structure of complete amyloid-beta(1-42) by cryo-EM | | Descriptor: | Amyloid beta A4 protein | | Authors: | Gremer, L, Schoelzel, D, Schenk, C, Reinartz, E, Labahn, J, Ravelli, R, Tusche, M, Lopez-Iglesias, C, Hoyer, W, Heise, H, Willbold, D, Schroeder, G.F. | | Deposit date: | 2017-08-14 | | Release date: | 2017-09-13 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Fibril structure of amyloid-beta (1-42) by cryo-electron microscopy.
Science, 358, 2017
|
|
