3IVS
 
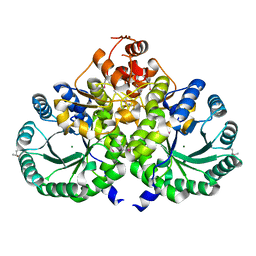 | | Homocitrate Synthase Lys4 | | Descriptor: | COBALT (II) ION, Homocitrate synthase, mitochondrial, ... | | Authors: | Bulfer, S.L, Scott, E.M, Couture, J.-F, Pillus, L, Trievel, R.C. | | Deposit date: | 2009-09-01 | | Release date: | 2009-09-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Crystal structure and functional analysis of homocitrate synthase, an essential enzyme in lysine biosynthesis.
J.Biol.Chem., 284, 2009
|
|
3F9Z
 
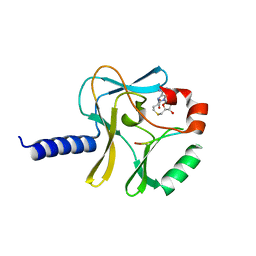 | | Structural Insights into Lysine Multiple Methylation by SET Domain Methyltransferases, SET8-Y245F / H4-Lys20 / AdoHcy | | Descriptor: | Histone H4, Histone-lysine N-methyltransferase SETD8, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Couture, J.-F, Dirk, L.M.A, Brunzelle, J.S, Houtz, R.L, Trievel, R.C. | | Deposit date: | 2008-11-14 | | Release date: | 2008-11-25 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural origins for the product specificity of SET domain protein methyltransferases.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
3IVU
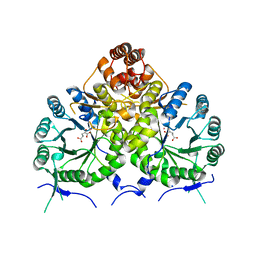 
 | | Homocitrate Synthase Lys4 bound to 2-OG | | Descriptor: | 2-OXOGLUTARIC ACID, COBALT (II) ION, Homocitrate synthase, ... | | Authors: | Bulfer, S.L, Scott, E.M, Couture, J.-F, Pillus, L, Trievel, R.C. | | Deposit date: | 2009-09-01 | | Release date: | 2009-09-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.72 Å) | | Cite: | Crystal structure and functional analysis of homocitrate synthase, an essential enzyme in lysine biosynthesis.
J.Biol.Chem., 284, 2009
|
|
3IVT
 
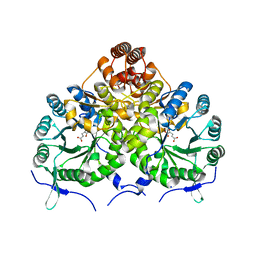 | | Homocitrate Synthase Lys4 bound to 2-OG | | Descriptor: | 2-OXOGLUTARIC ACID, Homocitrate synthase, mitochondrial, ... | | Authors: | Bulfer, S.L, Scott, E.M, Couture, J.-F, Pillus, L, Trievel, R.C. | | Deposit date: | 2009-09-01 | | Release date: | 2009-09-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.67 Å) | | Cite: | Crystal structure and functional analysis of homocitrate synthase, an essential enzyme in lysine biosynthesis.
J.Biol.Chem., 284, 2009
|
|
6BT2
 
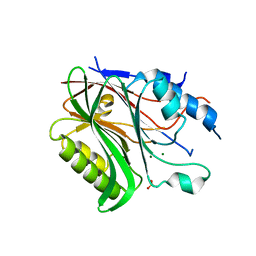 | | Structure of the human Nocturnin catalytic domain with bound sulfate anion | | Descriptor: | MAGNESIUM ION, Nocturnin, SULFATE ION | | Authors: | Abshire, E.T, Chasseur, J, Del Rizzo, P, Trievel, R. | | Deposit date: | 2017-12-04 | | Release date: | 2018-05-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.411 Å) | | Cite: | The structure of human Nocturnin reveals a conserved ribonuclease domain that represses target transcript translation and abundance in cells.
Nucleic Acids Res., 46, 2018
|
|
6BT1
 
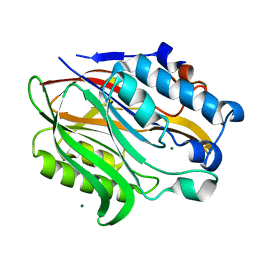 | | Structure of the human Nocturnin catalytic domain | | Descriptor: | MAGNESIUM ION, Nocturnin | | Authors: | Abshire, E.T, Chasseur, J, Del Rizzo, P, Trievel, R. | | Deposit date: | 2017-12-04 | | Release date: | 2018-05-16 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | The structure of human Nocturnin reveals a conserved ribonuclease domain that represses target transcript translation and abundance in cells.
Nucleic Acids Res., 46, 2018
|
|
5WO1
 
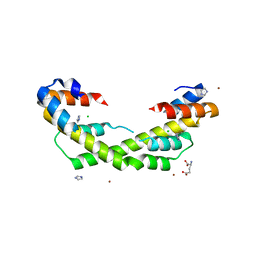 | | Chaperone Spy H96L bound to Im7 L18A L19A L37A (Im7 un-modeled) | | Descriptor: | CHLORIDE ION, GLUTAMIC ACID, IMIDAZOLE, ... | | Authors: | Horowitz, S, Koldewey, P, Martin, R, Bardwell, J.C.A. | | Deposit date: | 2017-08-01 | | Release date: | 2017-08-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Visualizing chaperone-assisted protein folding.
Nat. Struct. Mol. Biol., 23, 2016
|
|
5WO2
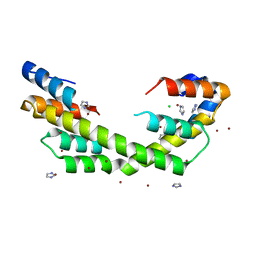 
 | | Chaperone Spy bound to Casein Fragment (Casein un-modeled) | | Descriptor: | CHLORIDE ION, IMIDAZOLE, Periplasmic chaperone Spy, ... | | Authors: | Horowitz, S, Koldewey, P, Martin, R, Bardwell, J.C.A. | | Deposit date: | 2017-08-01 | | Release date: | 2017-08-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.769 Å) | | Cite: | Visualizing chaperone-assisted protein folding.
Nat. Struct. Mol. Biol., 23, 2016
|
|
5WO3
 
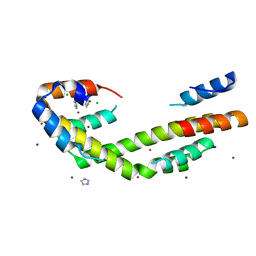 | | Chaperone Spy bound to Im7 (Im7 un-modeled) | | Descriptor: | CHLORIDE ION, IMIDAZOLE, Periplasmic chaperone Spy, ... | | Authors: | Horowitz, S, Koldewey, P, Martin, R, Bardwell, J.C.A. | | Deposit date: | 2017-08-01 | | Release date: | 2017-08-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Visualizing chaperone-assisted protein folding.
Nat. Struct. Mol. Biol., 23, 2016
|
|
3KK6
 
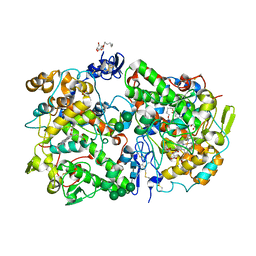 | | Crystal Structure of Cyclooxygenase-1 in complex with celecoxib | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 4-[5-(4-METHYLPHENYL)-3-(TRIFLUOROMETHYL)-1H-PYRAZOL-1-YL]BENZENESULFONAMIDE, CITRATE ANION, ... | | Authors: | Sidhu, R.S. | | Deposit date: | 2009-11-04 | | Release date: | 2009-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Coxibs interfere with the action of aspirin by binding tightly to one monomer of cyclooxygenase-1.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
