8IQ5
 
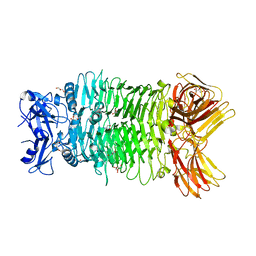 | | Crystal structure of trimeric K2-2 TSP | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, BICINE, GLYCEROL, ... | | Authors: | Ye, T.J, Huang, K.F, Ko, T.P. | | Deposit date: | 2023-03-15 | | Release date: | 2024-02-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Klebsiella pneumoniae K2 capsular polysaccharide degradation by a bacteriophage depolymerase does not require trimer formation.
Mbio, 15, 2024
|
|
6M5N
 
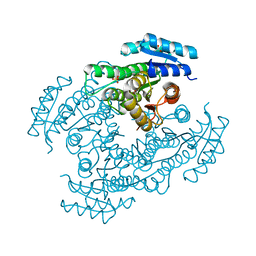 | | Apo-Form Structure of Borneol Dehydrogenase | | Descriptor: | ACETATE ION, Borneol dehydrogenase, SULFATE ION | | Authors: | Khine, A.A, Huang, K.F, Ko, T.P, Chen, H.P. | | Deposit date: | 2020-03-11 | | Release date: | 2020-07-15 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Structural characterization of borneol dehydrogenase from Pseudomonas sp. TCU-HL1
Acta Crystallogr.,Sect.F, 76, 2020
|
|
4J4M
 
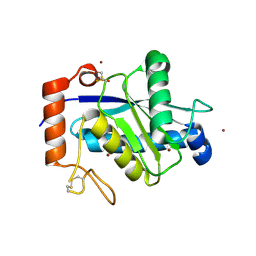 | | Crystal structure of TM-1, a Trimeresurus mucrosquamatus venom metalloproteinase | | Descriptor: | ZINC ION, zinc-dependent metalloproteinase | | Authors: | Chou, T.L, Wu, C.H, Huang, K.F, Wang, A.H. | | Deposit date: | 2013-02-07 | | Release date: | 2013-07-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a Trimeresurus mucrosquamatus venom metalloproteinase providing new insights into the inhibition by endogenous tripeptide inhibitors.
Toxicon, 71C, 2013
|
|
8K7I
 
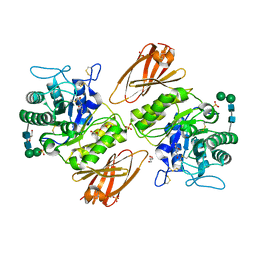 | | Crystal structure of human lysosomal alpha-galactosidase A in complex with (2R,3S,4R,5R)-2-(hydroxymethyl)-5-((methylamino)methyl)pyrrolidine-3,4-diol | | Descriptor: | (2~{R},3~{S},4~{R},5~{R})-2-(hydroxymethyl)-5-(methylaminomethyl)pyrrolidine-3,4-diol, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, H.Y, Huang, K.F, Ko, T.P, Cheng, W.C. | | Deposit date: | 2023-07-26 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Mechanistic Insights into Dibasic Iminosugars as pH-Selective Pharmacological Chaperones to Stabilize Human alpha-Galactosidase.
Jacs Au, 4, 2024
|
|
8K7J
 
 | | Crystal structure of human lysosomal alpha-galactosidase A in complex with (2R,3R,4S,5R)-2-((dimethylamino)methyl)-5-(hydroxymethyl)pyrrolidine-3,4-diol | | Descriptor: | (2~{R},3~{R},4~{S},5~{R})-2-[(dimethylamino)methyl]-5-(hydroxymethyl)pyrrolidine-3,4-diol, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, H.Y, Huang, K.F, Ko, T.P, Cheng, W.C. | | Deposit date: | 2023-07-26 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Mechanistic Insights into Dibasic Iminosugars as pH-Selective Pharmacological Chaperones to Stabilize Human alpha-Galactosidase.
Jacs Au, 4, 2024
|
|
8K7H
 
 | | Crystal structure of human lysosomal alpha-galactosidase A in complex with (2R,3S,4R)-2-(hydroxymethyl)-1-methylpyrrolidine-3,4-diol | | Descriptor: | (2~{R},3~{S},4~{R})-2-(hydroxymethyl)-1-methyl-pyrrolidine-3,4-diol, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, H.Y, Huang, K.F, Ko, T.P, Cheng, W.C. | | Deposit date: | 2023-07-26 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Mechanistic Insights into Dibasic Iminosugars as pH-Selective Pharmacological Chaperones to Stabilize Human alpha-Galactosidase.
Jacs Au, 4, 2024
|
|
8K7L
 
 | | Crystal structure of human lysosomal alpha-galactosidase A in complex with (2R,3R,4S,5R)-2-(aminomethyl)-5-(hydroxymethyl)-1-methylpyrrolidine-3,4-diol | | Descriptor: | (2~{R},3~{R},4~{S},5~{R})-2-(aminomethyl)-5-(hydroxymethyl)-1-methyl-pyrrolidine-3,4-diol, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, H.Y, Huang, K.F, Ko, T.P, Cheng, W.C. | | Deposit date: | 2023-07-26 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Mechanistic Insights into Dibasic Iminosugars as pH-Selective Pharmacological Chaperones to Stabilize Human alpha-Galactosidase.
Jacs Au, 4, 2024
|
|
8K7D
 
 | | Crystal structure of human lysosomal alpha-galactosidase A in complex with (2R,3R,4S,5R)-2-(aminomethyl)-5-(hydroxymethyl)pyrrolidine-3,4-diol | | Descriptor: | (2~{R},3~{R},4~{S},5~{R})-2-(aminomethyl)-5-(hydroxymethyl)pyrrolidine-3,4-diol, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, H.Y, Huang, K.F, Ko, T.P, Cheng, W.C. | | Deposit date: | 2023-07-26 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Mechanistic Insights into Dibasic Iminosugars as pH-Selective Pharmacological Chaperones to Stabilize Human alpha-Galactosidase.
Jacs Au, 4, 2024
|
|
8K7F
 
 | | Crystal structure of human lysosomal alpha-galactosidase A in complex with (2R,3S,4R,5R)-2,5-bis(hydroxymethyl)pyrrolidine-3,4-diol | | Descriptor: | (2~{R},3~{R},4~{S},5~{R})-2,5-bis(hydroxymethyl)pyrrolidine-3,4-diol, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, H.Y, Huang, K.F, Ko, T.P, Cheng, W.C. | | Deposit date: | 2023-07-26 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Mechanistic Insights into Dibasic Iminosugars as pH-Selective Pharmacological Chaperones to Stabilize Human alpha-Galactosidase.
Jacs Au, 4, 2024
|
|
8K7G
 
 | | Crystal structure of human lysosomal alpha-galactosidase A in complex with (2R,3S,4R,5R)-2,5-bis(hydroxymethyl)pyrrolidine-3,4-diol | | Descriptor: | (2~{R},3~{R},4~{S},5~{R})-2,5-bis(hydroxymethyl)pyrrolidine-3,4-diol, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, H.Y, Huang, K.F, Ko, T.P, Cheng, W.C. | | Deposit date: | 2023-07-26 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Mechanistic Insights into Dibasic Iminosugars as pH-Selective Pharmacological Chaperones to Stabilize Human alpha-Galactosidase.
Jacs Au, 4, 2024
|
|
8K7E
 | | Crystal structure of human lysosomal alpha-galactosidase A in complex with (2R,3R,4S,5R)-2-(aminomethyl)-5-(hydroxymethyl)pyrrolidine-3,4-diol | | Descriptor: | (2~{R},3~{R},4~{S},5~{R})-2-(aminomethyl)-5-(hydroxymethyl)pyrrolidine-3,4-diol, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, H.Y, Huang, K.F, Ko, T.P, Cheng, W.C. | | Deposit date: | 2023-07-26 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mechanistic Insights into Dibasic Iminosugars as pH-Selective Pharmacological Chaperones to Stabilize Human alpha-Galactosidase.
Jacs Au, 4, 2024
|
|
8K7K
 
 | | Crystal structure of human lysosomal alpha-galactosidase A in complex with (2R,3S,4R,5R)-2,5-bis(hydroxymethyl)-1-methylpyrrolidine-3,4-diol | | Descriptor: | (2~{R},3~{S},4~{R},5~{R})-2,5-bis(hydroxymethyl)-1-methyl-pyrrolidine-3,4-diol, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, H.Y, Huang, K.F, Ko, T.P, Cheng, W.C. | | Deposit date: | 2023-07-26 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Mechanistic Insights into Dibasic Iminosugars as pH-Selective Pharmacological Chaperones to Stabilize Human alpha-Galactosidase.
Jacs Au, 4, 2024
|
|
3VA4
 
 | | Crystal structure of the mammalian MDC1 FHA domain complexed with CHK2 pThr68 peptide | | Descriptor: | Mediator of DNA damage checkpoint protein 1, Serine/threonine-protein kinase Chk2 | | Authors: | Wu, H.H, Wu, P.Y, Huang, K.F, Kao, Y.Y, Tsai, M.D. | | Deposit date: | 2011-12-28 | | Release date: | 2012-02-01 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Structural Delineation of MDC1-FHA Domain Binding with CHK2-pThr68.
Biochemistry, 2012
|
|
5XFT
 
 | |
5ZYO
 
 | |
6JC5
 
 | |
6JC6
 
 | |
7EI3
 
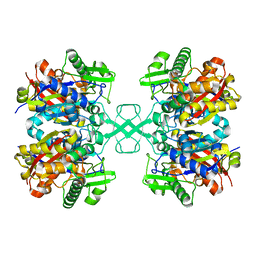 | |
7EI4
 
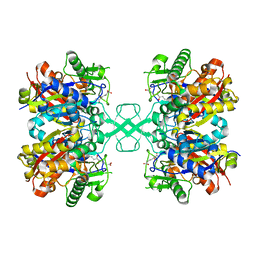 | | Crystal structure of MasL in complex with a novel covalent inhibitor, collimonin C | | Descriptor: | (6S,7R,9E)-6,7-bis(oxidanyl)hexadeca-9,15-dien-11,13-diynoic acid, Acetyl-CoA C-acyltransferase | | Authors: | Lin, C.C, Huang, K.F, Yang, Y.L. | | Deposit date: | 2021-03-30 | | Release date: | 2022-04-06 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Integrated omics approach to unveil antifungal bacterial polyynes as acetyl-CoA acetyltransferase inhibitors.
Commun Biol, 5, 2022
|
|
7EV4
 
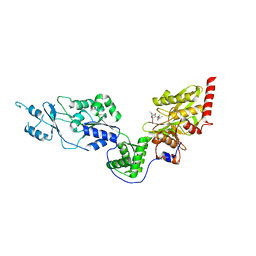 | | Crystal structure of the Lon-like protease MtaLonC with S582A mutation in complex with F-b20-Q | | Descriptor: | Endopeptidase La, F-b20-Q peptide {ortho-aminobenzoic acid (Abz)- QLRSLNGEWRFAWFPAPEAV[Tyr(3-NO2)]A}, PHOSPHATE ION | | Authors: | Hsieh, K.Y, Kuo, C.I, Su, S.C, Huang, K.F, Chang, C.I. | | Deposit date: | 2021-05-20 | | Release date: | 2021-11-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Processive cleavage of substrate at individual proteolytic active sites of the Lon protease complex.
Sci Adv, 7, 2021
|
|
7EV6
 
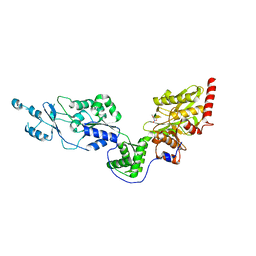 | | Crystal structure of the Lon-like protease MtaLonC with D581A mutation in complex with F-b20-Q | | Descriptor: | Endopeptidase La, F-b20-Q peptide {ortho-aminobenzoic acid (Abz)- QLRSLNGEWRFAWFPAPEAV[Tyr(3-NO2)]A}, PHOSPHATE ION | | Authors: | Hsieh, K.Y, Kuo, C.I, Su, S.C, Huang, K.F, Chang, C.I. | | Deposit date: | 2021-05-20 | | Release date: | 2021-11-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Processive cleavage of substrate at individual proteolytic active sites of the Lon protease complex.
Sci Adv, 7, 2021
|
|
7EUY
 
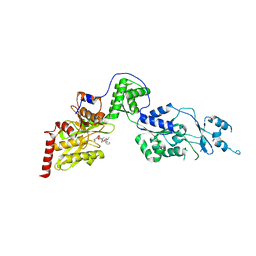 | | Crystal structure of the Lon-like protease MtaLonC with D582A mutation in complex with substrate polypeptide | | Descriptor: | ALA-PRO-GLU-ALA-VAL, Endopeptidase La, PHOSPHATE ION | | Authors: | Hsieh, K.Y, Kuo, C.I, Su, S.C, Huang, K.F, Chang, C.I. | | Deposit date: | 2021-05-19 | | Release date: | 2021-11-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Processive cleavage of substrate at individual proteolytic active sites of the Lon protease complex.
Sci Adv, 7, 2021
|
|
7EUX
 
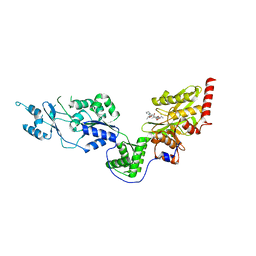 | | Crystal structure of the Lon-like protease MtaLonC with D581A mutation in complex with substrate polypeptide | | Descriptor: | ALA-PRO-GLU-ALA-VAL, Endopeptidase La, PHOSPHATE ION | | Authors: | Hsieh, K.Y, Kuo, C.I, Su, S.C, Huang, K.F, Chang, C.I. | | Deposit date: | 2021-05-19 | | Release date: | 2021-11-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Processive cleavage of substrate at individual proteolytic active sites of the Lon protease complex.
Sci Adv, 7, 2021
|
|
7FEA
 
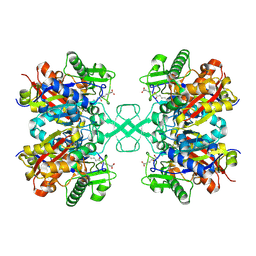 | | PY14 in complex with Col-D | | Descriptor: | (6~{R},7~{R},9~{E})-6,7-bis(oxidanyl)hexadeca-9,15-dien-11,13-diynoic acid, Acetyl-CoA C-acyltransferase | | Authors: | Lin, C.C, Ko, T.P, Huang, K.F, Yang, Y.L. | | Deposit date: | 2021-07-19 | | Release date: | 2022-07-27 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Integrated omics approach to unveil antifungal bacterial polyynes as acetyl-CoA acetyltransferase inhibitors.
Commun Biol, 5, 2022
|
|
7VT7
 
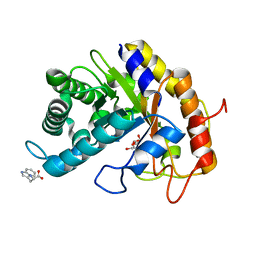 | | Crystal structure of CBM deleted MtGlu5 in complex with CBI | | Descriptor: | Endoglucanase H, GLYCEROL, TRYPTOPHAN, ... | | Authors: | Ye, T.J, Ko, P.T, Huang, K.F, Wu, S.H. | | Deposit date: | 2021-10-28 | | Release date: | 2022-09-07 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Synergic action of an inserted carbohydrate-binding module in a glycoside hydrolase family 5 endoglucanase.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
