2Y2Y
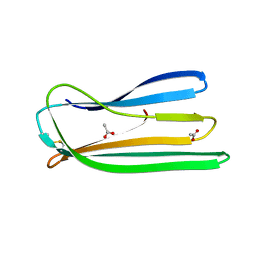 
 | | Oxidised form of E. coli CsgC | | Descriptor: | ACETATE ION, CURLI PRODUCTION PROTEIN CSGC | | Authors: | Taylor, J.D, Salgado, P.S, Constable, S.C, Cota, E, Mathews, S.J. | | Deposit date: | 2010-12-16 | | Release date: | 2011-09-21 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Atomic Resolution Insights Into Curli Fiber Biogenesis.
Structure, 19, 2011
|
|
1WAC
 
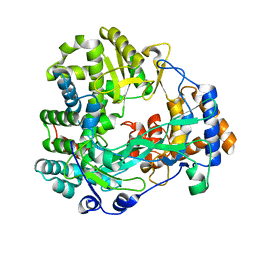 | | Back-priming mode of Phi6 RNA-dependent RNA polymerase | | Descriptor: | P2 PROTEIN | | Authors: | Laurila, M.R.L, Salgado, P.S, Stuart, D.I, Grimes, J.M, Bamford, D.H. | | Deposit date: | 2004-10-26 | | Release date: | 2005-01-27 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Back-Priming Mode of Phi6 RNA-Dependent RNA Polymerase
J.Gen.Virol., 86, 2005
|
|
6T8S
 
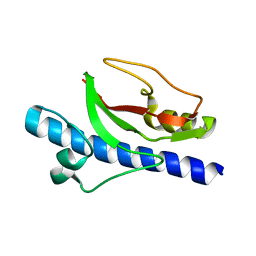 | |
4ZXR
 
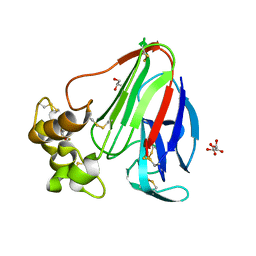 | | Structure of Thaumatin wrapped in graphene within vacuum | | Descriptor: | GLYCEROL, L(+)-TARTARIC ACID, Thaumatin-1 | | Authors: | Aller, P, Warren, A.J, Trincao, J, Evans, G. | | Deposit date: | 2015-05-20 | | Release date: | 2015-10-14 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | In vacuo X-ray data collection from graphene-wrapped protein crystals.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
7QGQ
 
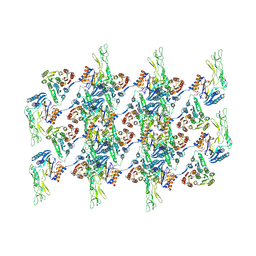 | |
4LEB
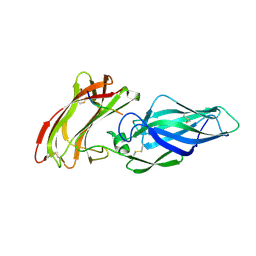 
 | |
4LEE
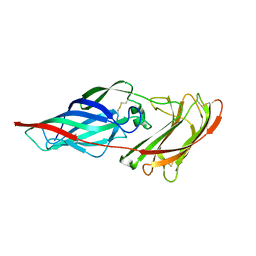 
 | | Structure of the Als3 adhesin from Candida albicans, residues 1-313 (mature sequence), triple mutant in the binding cavity: K59M, A116V, Y301F | | Descriptor: | Agglutinin-like protein 3 | | Authors: | Lin, J, Garnett, J.A, Cota, E. | | Deposit date: | 2013-06-25 | | Release date: | 2014-05-14 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The Peptide-binding Cavity Is Essential for Als3-mediated Adhesion of Candida albicans to Human Cells.
J.Biol.Chem., 289, 2014
|
|
