1HMA
 
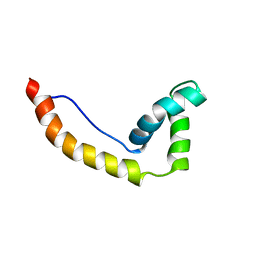 | | THE SOLUTION STRUCTURE AND DYNAMICS OF THE DNA BINDING DOMAIN OF HMG-D FROM DROSOPHILA MELANOGASTER | | Descriptor: | HMG-D | | Authors: | Jones, D.N.M, Searles, M.A, Shaw, G.L, Churchill, M.E.A, Ner, S.S, Keeler, J, Travers, A.A, Neuhaus, D. | | Deposit date: | 1994-05-12 | | Release date: | 1994-07-31 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The solution structure and dynamics of the DNA-binding domain of HMG-D from Drosophila melanogaster.
Structure, 2, 1994
|
|
4F7F
 
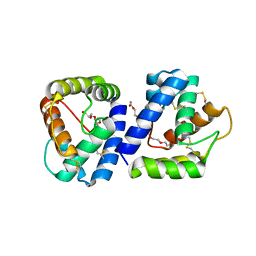 | | Structure of Anopheles gambiae odorant binding protein 20 | | Descriptor: | 1-(2-METHOXY-ETHOXY)-2-{2-[2-(2-METHOXY-ETHOXY]-ETHOXY}-ETHANE, AGAP005208-PA, NONAETHYLENE GLYCOL | | Authors: | Jones, D.N, Ziemba, B.P. | | Deposit date: | 2012-05-16 | | Release date: | 2012-10-17 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A novel mechanism of ligand binding and release in the odorant binding protein 20 from the malaria mosquito Anopheles gambiae.
Protein Sci., 22, 2013
|
|
6NBN
 
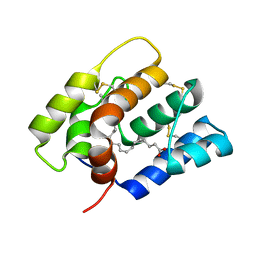 | |
3B86
 
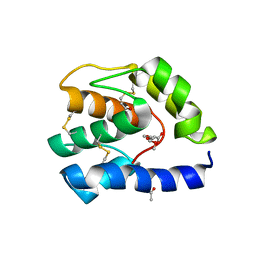 | |
3B87
 
 | | Complex of T57A Substituted Droposphila LUSH protein with Butanol | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, ACETATE ION, General odorant-binding protein lush | | Authors: | Jones, D.N.M, Thode, A.B. | | Deposit date: | 2007-10-31 | | Release date: | 2008-02-05 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The role of multiple hydrogen-bonding groups in specific alcohol binding sites in proteins: insights from structural studies of LUSH.
J.Mol.Biol., 376, 2008
|
|
3B88
 
 | | Complex of T57A Substituted Drosophila LUSH Protein with Ethanol | | Descriptor: | 2-{2-[2-2-(METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, ACETATE ION, General odorant-binding protein lush | | Authors: | Jones, D.N.M, Thode, A.B. | | Deposit date: | 2007-10-31 | | Release date: | 2008-02-05 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The role of multiple hydrogen-bonding groups in specific alcohol binding sites in proteins: insights from structural studies of LUSH.
J.Mol.Biol., 376, 2008
|
|
6OGH
 
 | | Structure of Aedes aegypti OBP22 in the complex with linoleic acid | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, AAEL005772-PA, CADMIUM ION, ... | | Authors: | Jones, D.N, Wang, J. | | Deposit date: | 2019-04-02 | | Release date: | 2019-04-24 | | Last modified: | 2020-05-06 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Aedes aegypti Odorant Binding Protein 22 selectively binds fatty acids through a conformational change in its C-terminal tail.
Sci Rep, 10, 2020
|
|
6OG0
 
 | | Structure of Aedes aegypti OBP22 | | Descriptor: | AAEL005772-PA, CADMIUM ION, CHLORIDE ION | | Authors: | Jones, D.N, Wang, J. | | Deposit date: | 2019-04-01 | | Release date: | 2019-04-17 | | Last modified: | 2020-05-06 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Aedes aegypti Odorant Binding Protein 22 selectively binds fatty acids through a conformational change in its C-terminal tail.
Sci Rep, 10, 2020
|
|
6OII
 
 | |
6OMW
 
 | |
6OPB
 
 | |
6OTL
 
 | |
6P2E
 
 | |
3B7A
 
 | |
3HXG
 
 | | Crystal structure of Schistsome eIF4E complexed with m7GpppA and 4E-BP | | Descriptor: | Eukaryotic Translation Initiation Factor 4E, Eukaryotic translation initiation factor 4E-binding protein 1, P1-7-METHYLGUANOSINE-P3-ADENOSINE-5',5'-TRIPHOSPHATE | | Authors: | Liu, W, Zhao, R, Jones, D.N.M, Davis, R.E. | | Deposit date: | 2009-06-20 | | Release date: | 2009-08-25 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural insights into parasite EIF4E binding specificity for m7G and m2,2,7G mRNA cap.
J.Biol.Chem., 284, 2009
|
|
3HXI
 
 | | Crystal structure of Schistosome eIF4E complexed with m7GpppG and 4E-BP | | Descriptor: | 7-METHYL-GUANOSINE-5'-TRIPHOSPHATE-5'-GUANOSINE, Eukaryotic Translation Initiation 4E, Eukaryotic translation initiation factor 4E-binding protein 1 | | Authors: | Liu, W, Zhao, R, Jones, D.N.M, Davis, R.E. | | Deposit date: | 2009-06-20 | | Release date: | 2009-08-25 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural insights into parasite EIF4E binding specificity for m7G and m2,2,7G mRNA cap.
J.Biol.Chem., 284, 2009
|
|
3V2L
 
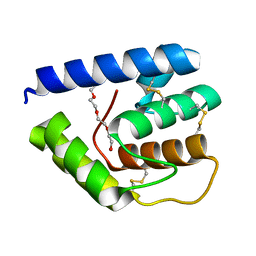 | |
3VB1
 
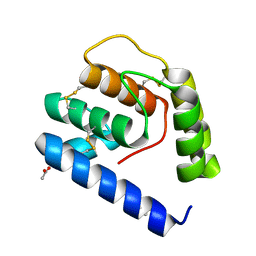 | |
2GTE
 | | Drosophila OBP LUSH bound to attractant pheromone 11-cis-vaccenyl acetate | | Descriptor: | (Z)-OCTADEC-11-ENYL ACETATE, General odorant-binding protein lush, PHOSPHATE ION | | Authors: | Laughlin, J.D, Ha, T, Smith, D.P, Jones, D.N.M. | | Deposit date: | 2006-04-27 | | Release date: | 2007-06-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Activation of pheromone-sensitive neurons is mediated by conformational activation of pheromone-binding protein
Cell(Cambridge,Mass.), 133, 2008
|
|
3B6X
 
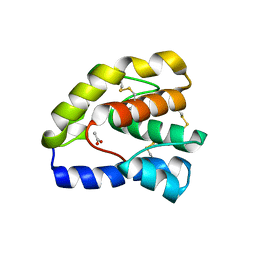 | |
2QDI
 
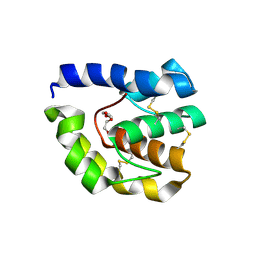 | | Drosophila OBP LUSH D118A mutation | | Descriptor: | General odorant-binding protein lush, TRIETHYLENE GLYCOL | | Authors: | Laughlin, J.D, Jones, D.N.M. | | Deposit date: | 2007-06-20 | | Release date: | 2008-06-03 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Activation of pheromone-sensitive neurons is mediated by conformational activation of pheromone-binding protein.
Cell(Cambridge,Mass.), 133, 2008
|
|
1QYP
 
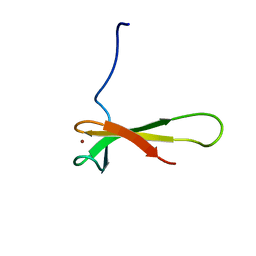 | | THERMOCOCCUS CELER RPB9, NMR, 25 STRUCTURES | | Descriptor: | RNA POLYMERASE II, ZINC ION | | Authors: | Wang, B, Jones, D.N.M, Kaine, B.P, Weiss, M.A. | | Deposit date: | 1997-08-19 | | Release date: | 1997-12-24 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | High-resolution structure of an archaeal zinc ribbon defines a general architectural motif in eukaryotic RNA polymerases.
Structure, 6, 1998
|
|
1T14
 
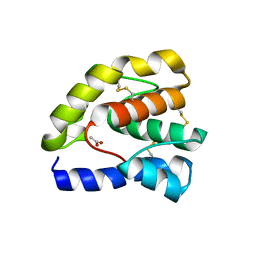 | |
3Q8I
 
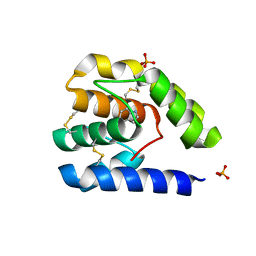 | |
1OOI
 
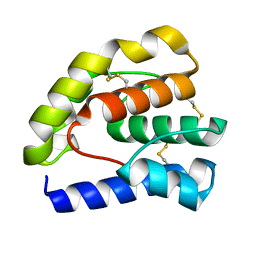 | |
