4YVB
 
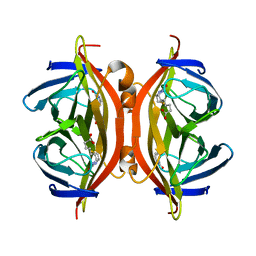 | | Structure of D128N streptavidin | | Descriptor: | BIOTIN, Streptavidin | | Authors: | Baugh, L, Le Trong, I, Stayton, P.S, Stenkamp, R.E, Lybrand, T.P. | | Deposit date: | 2015-03-19 | | Release date: | 2016-03-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.351 Å) | | Cite: | A Streptavidin Binding Site Mutation Yields an Unexpected Result
To Be Published
|
|
3T6F
 
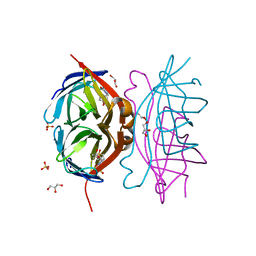 | | Biotin complex of Y54F core streptavidin | | Descriptor: | BIOTIN, BIOTIN-D-SULFOXIDE, GLYCEROL, ... | | Authors: | Baugh, L, Le Trong, I, Cerutti, D.S, Mehta, N, Gulich, S, Stayton, P.S, Stenkamp, R.E, Lybrand, T.P. | | Deposit date: | 2011-07-28 | | Release date: | 2011-12-21 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.22 Å) | | Cite: | Second-Contact Shell Mutation Diminishes Streptavidin-Biotin Binding Affinity through Transmitted Effects on Equilibrium Dynamics.
Biochemistry, 51, 2012
|
|
3T6L
 
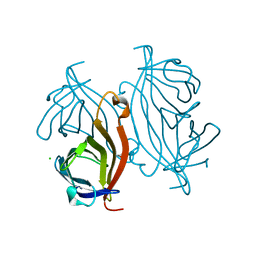 | | Y54F mutant of core streptavidin | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Streptavidin | | Authors: | Baugh, L, Le Trong, I, Stayton, P.S, Stenkamp, R.E, Lybrand, T.P. | | Deposit date: | 2011-07-28 | | Release date: | 2011-12-21 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Second-Contact Shell Mutation Diminishes Streptavidin-Biotin Binding Affinity through Transmitted Effects on Equilibrium Dynamics.
Biochemistry, 51, 2012
|
|
3MG5
 
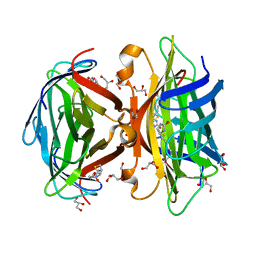 | | Core-streptavidin mutant F130L in complex with biotin | | Descriptor: | BIOTIN, GLYCEROL, Streptavidin | | Authors: | Le Trong, I, Baugh, L, Stayton, P.S, Lybrand, T.P, Stenkamp, R.E. | | Deposit date: | 2010-04-05 | | Release date: | 2010-05-26 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | A distal point mutation in the streptavidin-biotin complex preserves structure but diminishes binding affinity: experimental evidence of electronic polarization effects?
Biochemistry, 49, 2010
|
|
3FTP
 
 | |
4EFI
 
 | | Crystal Structure of 3-oxoacyl-(Acyl-carrier protein) Synthase from Burkholderia Xenovorans LB400 | | Descriptor: | 3-oxoacyl-(Acyl-carrier protein) synthase, CHLORIDE ION, FORMIC ACID, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease, Craig, T.K, Abendroth, J, Staker, B, Stewart, L, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2012-03-29 | | Release date: | 2012-04-11 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Combining functional and structural genomics to sample the essential Burkholderia structome.
Plos One, 8, 2013
|
|
4EG0
 
 | |
4EK2
 
 | |
4EGJ
 
 | |
3SZ8
 
 | |
4EXQ
 
 | |
3TML
 
 | |
4O2D
 
 | |
4EGF
 
 | |
4EYE
 
 | |
4ED4
 
 | |
4EO9
 
 | |
3TDE
 
 | |
3SWX
 
 | |
3TCR
 
 | |
4EMD
 
 | |
3TAV
 
 | |
4EGE
 
 | |
4KAM
 
 | |
4LGV
 | |
