2OKW
 | |
2OKY
 | |
3EOJ
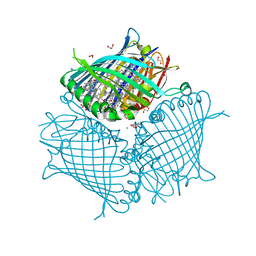 
 | | Fmo protein from Prosthecochloris Aestuarii 2K AT 1.3A Resolution | | Descriptor: | 1,2-ETHANEDIOL, AMMONIUM ION, BACTERIOCHLOROPHYLL A, ... | | Authors: | Tronrud, D.E, Wen, J, Gay, L, Blankenship, R.E. | | Deposit date: | 2008-09-27 | | Release date: | 2009-05-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | The structural basis for the difference in absorbance spectra for the FMO antenna protein from various green sulfur bacteria.
Photosynth.Res., 100, 2009
|
|
192L
 
 | |
190L
 
 | |
1B9C
 
 | |
191L
 
 | |
1GYM
 
 | | PHOSPHATIDYLINOSITOL-SPECIFIC PHOSPHOLIPASE C IN COMPLEX WITH GLUCOSAMINE-(ALPHA-1-6)-MYO-INOSITOL | | Descriptor: | (1R,2R,3R,4R,5R,6S)-2,3,4,5,6-pentahydroxycyclohexyl 2-amino-2-deoxy-alpha-D-glucopyranoside, PHOSPHATIDYLINOSITOL-SPECIFIC PHOSPHOLIPASE C | | Authors: | Heinz, D.W, Ryan, M, Smith, M.P, Weaver, L.H, Keana, J.F.W, Griffith, O.H. | | Deposit date: | 1996-05-02 | | Release date: | 1996-11-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of phosphatidylinositol-specific phospholipase C from Bacillus cereus in complex with glucosaminyl(alpha 1-->6)-D-myo-inositol, an essential fragment of GPI anchors.
Biochemistry, 35, 1996
|
|
