1DY4
 
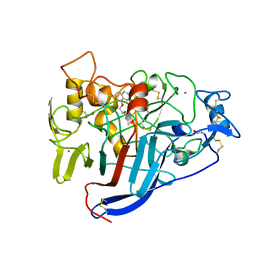 | | CBH1 IN COMPLEX WITH S-PROPRANOLOL | | Descriptor: | 1-(ISOPROPYLAMINO)-3-(1-NAPHTHYLOXY)-2-PROPANOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, COBALT (II) ION, ... | | Authors: | Stahlberg, J, Henriksson, H, Divne, C, Isaksson, R, Pettersson, G, Johansson, G, Jones, T.A. | | Deposit date: | 2000-01-26 | | Release date: | 2000-12-18 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Basis for Enantiomer Binding and Separation of a Common Beta-Blocker: Crystal Structure of Cellobiohydrolase Cel7A with Bound (S)-Propranolol at 1.9 A Resolution
J.Mol.Biol., 305, 2001
|
|
2WGS
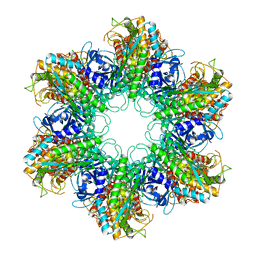 
 | | Crystal structure of Mycobacterium Tuberculosis Glutamine Synthetase in complex with a purine analogue inhibitor. | | Descriptor: | 1-(3,4-dichlorobenzyl)-3,7-dimethyl-8-morpholin-4-yl-3,7-dihydro-1H-purine-2,6-dione, CHLORIDE ION, GLUTAMINE SYNTHETASE 1 | | Authors: | Nilsson, M.T, Krajewski, W.W, Jones, T.A, Mowbray, S.L. | | Deposit date: | 2009-04-27 | | Release date: | 2009-09-01 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structural Basis for the Inhibition of Mycobacterium Tuberculosis Glutamine Synthetase by Novel ATP-Competitive Inhibitors.
J.Mol.Biol., 393, 2009
|
|
2WHI
 
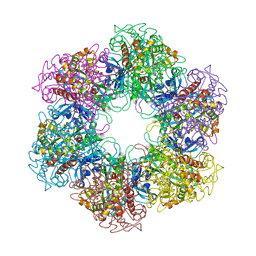 | | Crystal structure of Mycobacterium Tuberculosis Glutamine Synthetase in complex with a purine analogue inhibitor and L-methionine-S- sulfoximine phosphate. | | Descriptor: | 1-(3,4-dichlorobenzyl)-3,7-dimethyl-8-morpholin-4-yl-3,7-dihydro-1H-purine-2,6-dione, CHLORIDE ION, GLUTAMINE SYNTHETASE 1, ... | | Authors: | Nilsson, M.T, Krajewski, W.W, Jones, T.A, Mowbray, S.L. | | Deposit date: | 2009-05-05 | | Release date: | 2009-09-01 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural Basis for the Inhibition of Mycobacterium Tuberculosis Glutamine Synthetase by Novel ATP-Competitive Inhibitors.
J.Mol.Biol., 393, 2009
|
|
1EG1
 
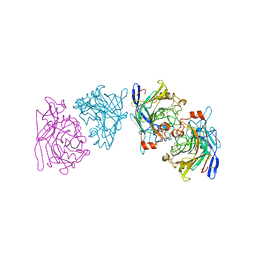 | | ENDOGLUCANASE I FROM TRICHODERMA REESEI | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ENDOGLUCANASE I | | Authors: | Kleywegt, G.J, Zou, J.-Y, Jones, T.A. | | Deposit date: | 1996-11-26 | | Release date: | 1997-08-20 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | The crystal structure of the catalytic core domain of endoglucanase I from Trichoderma reesei at 3.6 A resolution, and a comparison with related enzymes.
J.Mol.Biol., 272, 1997
|
|
1CB2
 
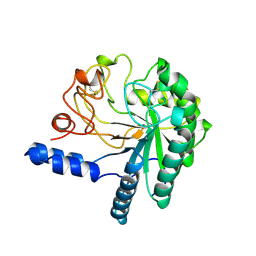 | | CELLOBIOHYDROLASE II, CATALYTIC DOMAIN, MUTANT Y169F | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CELLOBIOHYDROLASE II, alpha-D-mannopyranose | | Authors: | Kleywegt, G.J, Szardenings, M, Jones, T.A. | | Deposit date: | 1995-11-25 | | Release date: | 1996-10-14 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The active site of Trichoderma reesei cellobiohydrolase II: the role of tyrosine 169.
Protein Eng., 9, 1996
|
|
1CA2
 
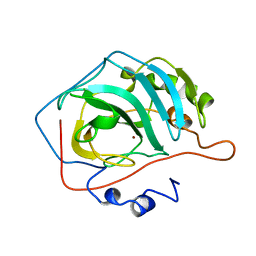 | |
1HGW
 
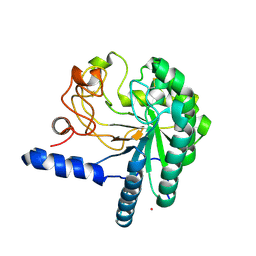 | | CEL6A D175A mutant | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CELLOBIOHYDROLASE CEL6A (FORMERLY CALLED CBH II), COBALT (II) ION, ... | | Authors: | Zou, J.-Y, Jones, T.A. | | Deposit date: | 2000-12-15 | | Release date: | 2002-01-15 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Active Site of Cellobiohydrolase Cel6A from Trichoderma Reesei: The Roles of Aspartic Acids D221 and D175
J.Am.Chem.Soc., 124, 2002
|
|
1GSD
 
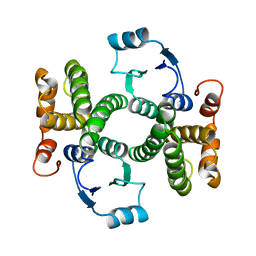 | |
1CEL
 
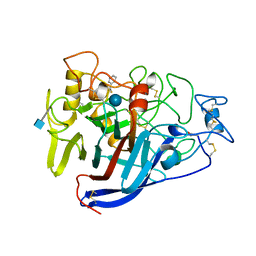 | |
1GSE
 
 | |
1HGY
 
 | | CEL6A D221A mutant | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CELLOBIOHYDROLASE CEL6A (FORMERLY CALLED CBH II), alpha-D-glucopyranose, ... | | Authors: | Zou, J.-Y, Kleywegt, G.J, Jones, T.A. | | Deposit date: | 2000-12-15 | | Release date: | 2002-01-15 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Active Site of Cellobiohydrolase Cel6A from Trichoderma Reesei: The Roles of Aspartic Acids D221 and D175
J.Am.Chem.Soc., 124, 2002
|
|
5ADH
 
 | |
2UU7
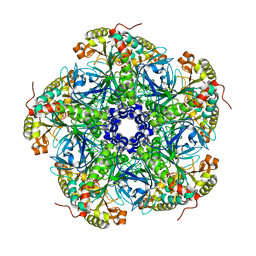 
 | |
2WA1
 
 | | Structure of the methyltransferase domain from Modoc Virus, a Flavivirus with No Known Vector (NKV) | | Descriptor: | NON-STRUCTURAL PROTEIN 5, SULFATE ION | | Authors: | Jansson, A.M, Johansson, P, Jones, T.A. | | Deposit date: | 2009-02-02 | | Release date: | 2009-08-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the Methyltransferase Domain from the Modoc Virus, a Flavivirus with No Known Vector.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
2WA2
 
 | | Structure of the methyltransferase domain from Modoc Virus, a Flavivirus with No Known Vector (NKV) | | Descriptor: | NON-STRUCTURAL PROTEIN 5, S-ADENOSYLMETHIONINE, SULFATE ION | | Authors: | Jansson, A.M, Johansson, P, Jones, T.A. | | Deposit date: | 2009-02-02 | | Release date: | 2009-08-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of the Methyltransferase Domain from the Modoc Virus, a Flavivirus with No Known Vector.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
1QK0
 
 | | CEL6A WITH A NON-HYDROLYSABLE CELLOTETRAOSE | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3-IODO-BENZYL ALCOHOL, CELLOBIOHYDROLASE CEL6A (FORMERLY CALLED CBH II), ... | | Authors: | Zou, J.-Y, Jones, T.A. | | Deposit date: | 1999-07-08 | | Release date: | 1999-09-18 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystallographic Evidence for Substrate Ring Distortion and Protein Conformational Changes During Catalysis in Cellobiohydrolase Cel6A from Trichoderma Reesei
Structure, 7, 1999
|
|
1QK2
 
 | | WILD TYPE CEL6A WITH A NON-HYDROLYSABLE CELLOTETRAOSE | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CELLOBIOHYDROLASE CEL6A (FORMERLY CALLED CBH II), alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-4-thio-beta-D-glucopyranose-(1-4)-methyl beta-D-glucopyranoside, ... | | Authors: | Zou, J.-Y, Jones, T.A. | | Deposit date: | 1999-07-09 | | Release date: | 1999-09-18 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystallographic Evidence for Substrate Ring Distortion and Protein Conformational Changes During Catalysis in Cellobiohydrolase Cel6A from Trichoderma Reesei
Structure, 7, 1999
|
|
1KX9
 
 | | ANTENNAL CHEMOSENSORY PROTEIN A6 FROM THE MOTH MAMESTRA BRASSICAE | | Descriptor: | ACETATE ION, CHEMOSENSORY PROTEIN A6 | | Authors: | Lartigue, A, Campanacci, V, Roussel, A, Larsson, A.M, Jones, T.A, Tegoni, M, Cambillau, C. | | Deposit date: | 2002-01-31 | | Release date: | 2002-12-04 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | X-ray structure and ligand binding study of a moth chemosensory protein
J.Biol.Chem., 277, 2002
|
|
1KX8
 
 | | Antennal Chemosensory Protein A6 from Mamestra brassicae, tetragonal form | | Descriptor: | CHEMOSENSORY PROTEIN A6 | | Authors: | Lartigue, A, Campanacci, V, Roussel, A, Larsson, A.M, Jones, T.A, Tegoni, M, Cambillau, C. | | Deposit date: | 2002-01-31 | | Release date: | 2002-12-04 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | X-Ray Structure and Ligand Binding Study of a Chemosensory Protein
J.Biol.Chem., 277, 2002
|
|
1BH5
 
 | | HUMAN GLYOXALASE I Q33E, E172Q DOUBLE MUTANT | | Descriptor: | LACTOYLGLUTATHIONE LYASE, S-HEXYLGLUTATHIONE, ZINC ION | | Authors: | Cameron, A.D, Jones, T.A. | | Deposit date: | 1998-06-13 | | Release date: | 1998-11-04 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Involvement of an active-site Zn2+ ligand in the catalytic mechanism of human glyoxalase I.
J.Biol.Chem., 273, 1998
|
|
1FRO
 
 | |
1FCC
 
 | | CRYSTAL STRUCTURE OF THE C2 FRAGMENT OF STREPTOCOCCAL PROTEIN G IN COMPLEX WITH THE FC DOMAIN OF HUMAN IGG | | Descriptor: | IGG1 MO61 FC, STREPTOCOCCAL PROTEIN G (C2 FRAGMENT) | | Authors: | Sauer-Eriksson, A.E, Kleywegt, G.J, Uhlen, M, Jones, T.A. | | Deposit date: | 1995-01-17 | | Release date: | 1995-04-20 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal structure of the C2 fragment of streptococcal protein G in complex with the Fc domain of human IgG.
Structure, 3, 1995
|
|
1HB6
 
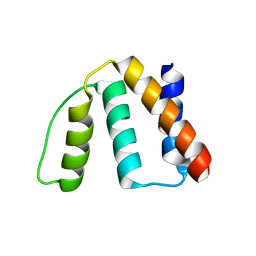 | | Structure of bovine Acyl-CoA binding protein in orthorhombic crystal form | | Descriptor: | ACYL-COA BINDING PROTEIN, CADMIUM ION | | Authors: | Zou, J.Y, Kleywegt, G.J, Bergfors, T, Knudsen, J, Jones, T.A. | | Deposit date: | 2001-04-12 | | Release date: | 2002-03-11 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Binding Site Differences Revealed by Crystal Structures of Plasmodium Falciparum and Bovine Acyl-Coa Binding Protein
J.Mol.Biol., 309, 2001
|
|
1H8V
 
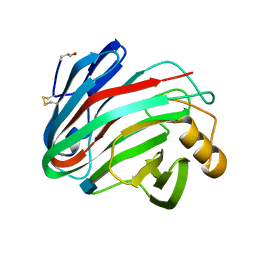 | | The X-ray Crystal Structure of the Trichoderma reesei Family 12 Endoglucanase 3, Cel12A, at 1.9 A Resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ENDO-BETA-1,4-GLUCANASE | | Authors: | Sandgren, M, Shaw, A, Ropp, T.H, Wu, S, Bott, R, Cameron, A.D, Stahlberg, J, Mitchinson, C, Jones, T.A. | | Deposit date: | 2001-02-16 | | Release date: | 2001-04-24 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The X-Ray Crystal Structure of the Trichoderma Reesei Family 12 Endoglucanase 3, Cel12A, at 1.9 A Resolution
J.Mol.Biol., 308, 2001
|
|
4Y6S
 
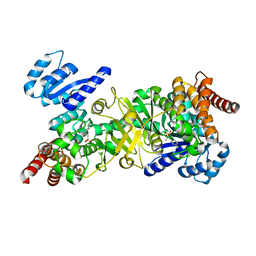 | | Structure of Plasmodium falciparum DXR in complex with a beta-substituted fosmidomycin analogue, RC134, and manganese | | Descriptor: | 1-deoxy-D-xylulose 5-phosphate reductoisomerase, apicoplast, MANGANESE (II) ION, ... | | Authors: | Sooriyaarachchi, S, Bergfors, T, Jones, T.A, Mowbray, S.L. | | Deposit date: | 2015-02-13 | | Release date: | 2015-04-01 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Synthesis and Bioactivity of beta-Substituted Fosmidomycin Analogues Targeting 1-Deoxy-d-xylulose-5-phosphate Reductoisomerase.
J.Med.Chem., 58, 2015
|
|
