2H21
 
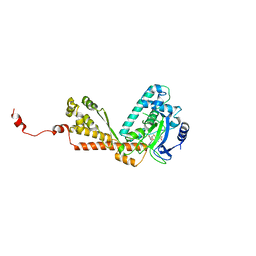 | | Structure of Rubisco LSMT bound to AdoMet | | Descriptor: | Ribulose-1,5 bisphosphate carboxylase/oxygenase, S-ADENOSYLMETHIONINE | | Authors: | Couture, J.F, Hauk, G, Trievel, R.C. | | Deposit date: | 2006-05-17 | | Release date: | 2006-05-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Catalytic Roles for Carbon-Oxygen Hydrogen Bonding in SET Domain Lysine Methyltransferases.
J.Biol.Chem., 281, 2006
|
|
2H23
 
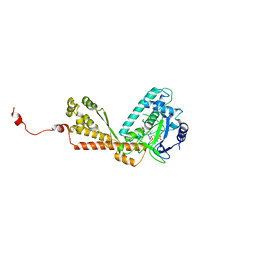 | | Structure of Rubisco LSMT bound to Trimethyllysine and AdoHcy | | Descriptor: | N-TRIMETHYLLYSINE, Ribulose-1,5 bisphosphate carboxylase/oxygenase large subunit N-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Couture, J.F, Hauk, G, Trievel, R.C. | | Deposit date: | 2006-05-17 | | Release date: | 2006-05-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Catalytic Roles for Carbon-Oxygen Hydrogen Bonding in SET Domain Lysine Methyltransferases.
J.Biol.Chem., 281, 2006
|
|
1EMS
 
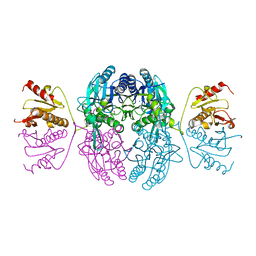 | | CRYSTAL STRUCTURE OF THE C. ELEGANS NITFHIT PROTEIN | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ETHYL MERCURY ION, NIT-FRAGILE HISTIDINE TRIAD FUSION PROTEIN, ... | | Authors: | Pace, H.C, Hodawadekar, S.C, Draganescu, A, Huang, J, Bieganowski, P, Pekarsky, Y, Croce, C.M, Brenner, C. | | Deposit date: | 2000-03-17 | | Release date: | 2000-07-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the worm NitFhit Rosetta Stone protein reveals a Nit tetramer binding two Fhit dimers.
Curr.Biol., 10, 2000
|
|
5TJD
 
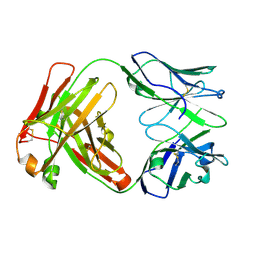 | | Computer-based rational design of improved functionality for antibody catalysts toward organophosphorus compounds | | Descriptor: | FAB A.17 L47K mutant HEAVY CHAIN, FAB A.17 L47K mutant Light Chain | | Authors: | Chatziefthimiou, S, Stepanova, A, Smirnov, I, Gabibov, A, Wilmanns, M. | | Deposit date: | 2016-10-04 | | Release date: | 2018-05-02 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Multiscale computation delivers organophosphorus reactivity and stereoselectivity to immunoglobulin scavengers.
Proc.Natl.Acad.Sci.USA, 2020
|
|
1JKN
 
 | | Solution Structure of the Nudix Enzyme Diadenosine Tetraphosphate Hydrolase from Lupinus angustifolius Complexed with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, diadenosine 5',5'''-P1,P4-tetraphosphate hydrolase | | Authors: | Fletcher, J.I, Swarbrick, J.D, Maksel, D, Gayler, K.R, Gooley, P.R. | | Deposit date: | 2001-07-12 | | Release date: | 2002-02-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The structure of Ap(4)A hydrolase complexed with ATP-MgF(x) reveals the basis of substrate binding.
Structure, 10, 2002
|
|
