1CKL
 
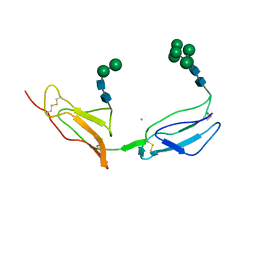 | | N-TERMINAL TWO DOMAINS OF HUMAN CD46 (MEMBRANE COFACTOR PROTEIN, MCP) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Casasnovas, J, Larvie, M, Stehle, T. | | Deposit date: | 1999-04-22 | | Release date: | 1999-06-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal structure of two CD46 domains reveals an extended measles virus-binding surface.
EMBO J., 18, 1999
|
|
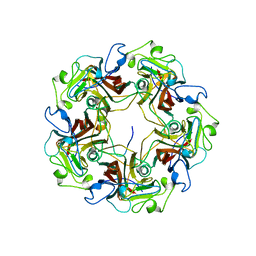
1CN3
 | |
3MK4
 
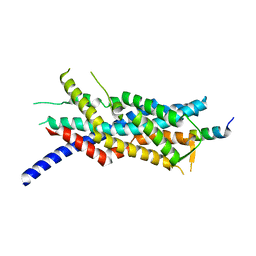 | | X-Ray structure of human PEX3 in complex with a PEX19 derived peptide | | Descriptor: | Peroxisomal biogenesis factor 19, Peroxisomal biogenesis factor 3 | | Authors: | Schmidt, F, Treiber, N, Dodt, G, Stehle, T. | | Deposit date: | 2010-04-14 | | Release date: | 2010-06-30 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Insights into peroxisome function from the structure of PEX3 in complex with a soluble fragment of PEX19
J.Biol.Chem., 285, 2010
|
|
3EXW
 
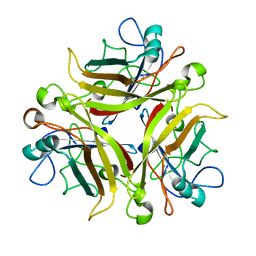 | |
7QD7
 
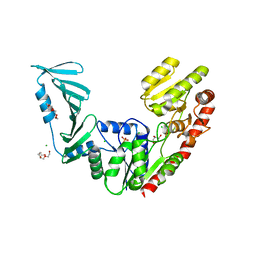 | | TarM(Se)_G117R | | Descriptor: | CHLORIDE ION, GLYCEROL, PENTAETHYLENE GLYCOL, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2021-11-26 | | Release date: | 2023-05-10 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Invasive Staphylococcus epidermidis uses a unique processive wall teichoic acid glycosyltransferase to evade immune recognition.
Sci Adv, 9, 2023
|
|
7QH9
 
 | | TarM(Se)_G117R-4RboP | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, TarM(Se)_G117R-4RboP, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2021-12-10 | | Release date: | 2023-05-10 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.689 Å) | | Cite: | Invasive Staphylococcus epidermidis uses a unique processive wall teichoic acid glycosyltransferase to evade immune recognition.
Sci Adv, 9, 2023
|
|
7QNT
 
 | | TarM(Se) native | | Descriptor: | 1,2-ETHANEDIOL, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2021-12-22 | | Release date: | 2023-05-10 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (3.21 Å) | | Cite: | Invasive Staphylococcus epidermidis uses a unique processive wall teichoic acid glycosyltransferase to evade immune recognition.
Sci Adv, 9, 2023
|
|
3I4X
 
 | | Crystal structure of the dimethylallyl tryptophan synthase FgaPT2 from Aspergillus fumigatus in complex with Trp and DMSPP | | Descriptor: | DIMETHYLALLYL S-THIOLODIPHOSPHATE, GLYCEROL, TRYPTOPHAN, ... | | Authors: | Schall, C, Zocher, G, Stehle, T. | | Deposit date: | 2009-07-03 | | Release date: | 2009-09-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The structure of dimethylallyl tryptophan synthase reveals a common architecture of aromatic prenyltransferases in fungi and bacteria
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
3I4Z
 
 | | Crystal structure of the dimethylallyl tryptophan synthase FgaPT2 from Aspergillus fumigatus | | Descriptor: | 1,3-BUTANEDIOL, GLYCEROL, Tryptophan dimethylallyltransferase | | Authors: | Schall, C, Zocher, G, Stehle, T. | | Deposit date: | 2009-07-03 | | Release date: | 2009-09-01 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | The structure of dimethylallyl tryptophan synthase reveals a common architecture of aromatic prenyltransferases in fungi and bacteria
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
3F0Y
 
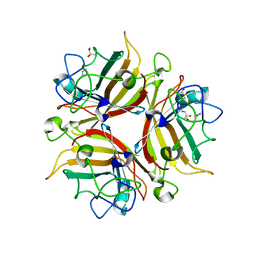 | | Crystal structure of the human Adenovirus type 14 fiber knob | | Descriptor: | Fiber protein, GLYCEROL, IMIDAZOLE | | Authors: | Persson, B.D, Reiter, D.M, Arnberg, N, Stehle, T. | | Deposit date: | 2008-10-27 | | Release date: | 2008-11-25 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | An arginine switch in the species B adenovirus knob determines high-affinity engagement of cellular receptor CD46
J.Virol., 83, 2009
|
|
3INB
 
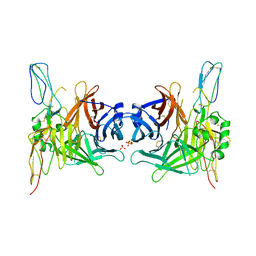 | | Structure of the measles virus hemagglutinin bound to the CD46 receptor | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin glycoprotein, Membrane cofactor protein, ... | | Authors: | Santiago, C, Celma, M.L, Stehle, T, Casasnovas, J.M. | | Deposit date: | 2009-08-12 | | Release date: | 2009-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structure of the measles virus hemagglutinin bound to the CD46 receptor
Nat.Struct.Mol.Biol., 17, 2010
|
|
8OQX
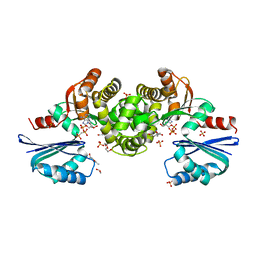 
 | | Crystal structure of Tannerella forsythia MurNAc kinase MurK with a phosphate analogue | | Descriptor: | 1,2-ETHANEDIOL, ATPase, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Gogler, K, Fink, P, Stasiak, A.C, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-12 | | Release date: | 2023-08-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
8OQW
 
 | | Crystal structure of Tannerella forsythia MurNAc kinase MurK | | Descriptor: | ATPase, GLYCEROL, SULFATE ION | | Authors: | Gogler, K, Fink, P, Stasiak, A.C, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-12 | | Release date: | 2023-08-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
8OW7
 
 | | Crystal structure of Tannerella forsythia sugar kinase K1058 in complex with N-acetylmuramic acid (MurNAc) | | Descriptor: | N-acetyl-beta-muramic acid, N-acetylglucosamine kinase, SULFATE ION | | Authors: | Stasiak, A.C, Gogler, K, Fink, P, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-08-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.06 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
8OQK
 
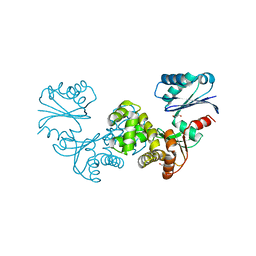 | | Crystal structure of Tannerella forsythia sugar kinase K1058 | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, N-acetylglucosamine kinase | | Authors: | Gogler, K, Fink, P, Stasiak, A.C, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-12 | | Release date: | 2023-08-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
8OW9
 
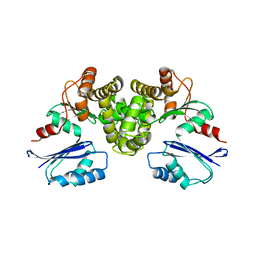 | | Crystal structure of Tannerella forsythia MurNAc kinase MurK in complex with N-acetylmuramic acid (MurNAc) | | Descriptor: | N-acetyl-beta-muramic acid, Putative novel MurNAc kinase | | Authors: | Stasiak, A.C, Gogler, K, Fink, P, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-08-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
1VSC
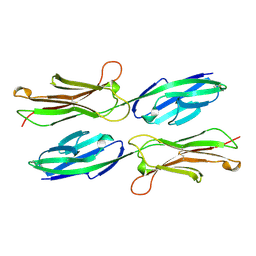 
 | | VCAM-1 | | Descriptor: | VASCULAR CELL ADHESION MOLECULE-1 | | Authors: | Wang, J, Stehle, T, Osborn, L. | | Deposit date: | 1995-04-27 | | Release date: | 1996-06-20 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The crystal structure of an N-terminal two-domain fragment of vascular cell adhesion molecule 1 (VCAM-1): a cyclic peptide based on the domain 1 C-D loop can inhibit VCAM-1-alpha 4 integrin interaction.
Proc.Natl.Acad.Sci.USA, 92, 1995
|
|
1U8C
 
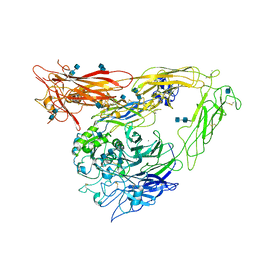 | | A novel adaptation of the integrin PSI domain revealed from its crystal structure | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Xiong, J.P, Stehle, T, Goodman, S.L, Arnaout, M.A. | | Deposit date: | 2004-08-05 | | Release date: | 2004-12-28 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | A novel adaptation of the integrin PSI domain revealed from its crystal structure.
J.Biol.Chem., 279, 2004
|
|
4MJ0
 
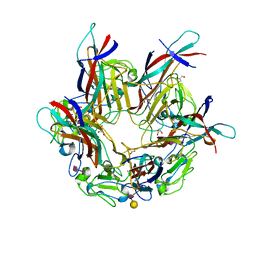 | | BK Polyomavirus VP1 pentamer in complex with GD3 oligosaccharide | | Descriptor: | CHLORIDE ION, GLYCEROL, N-acetyl-alpha-neuraminic acid-(2-8)-N-acetyl-alpha-neuraminic acid, ... | | Authors: | Neu, U, Stroh, L.J, Stehle, T. | | Deposit date: | 2013-09-03 | | Release date: | 2013-11-06 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A Structure-Guided Mutation in the Major Capsid Protein Retargets BK Polyomavirus.
Plos Pathog., 9, 2013
|
|
3O2K
 
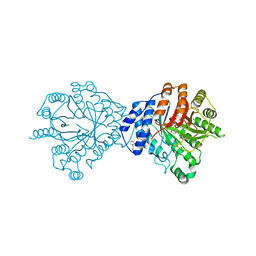 | | Crystal Structure of Brevianamide F Prenyltransferase Complexed with Brevianamide F and Dimethylallyl S-thiolodiphosphate | | Descriptor: | (3S,8aS)-3-(1H-indol-3-ylmethyl)hexahydropyrrolo[1,2-a]pyrazine-1,4-dione, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Brevianamide F prenyltransferase, ... | | Authors: | Jost, M, Zocher, G.E, Stehle, T. | | Deposit date: | 2010-07-22 | | Release date: | 2010-12-08 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure-function analysis of an enzymatic prenyl transfer reaction identifies a reaction chamber with modifiable specificity.
J.Am.Chem.Soc., 132, 2010
|
|
3PH5
 
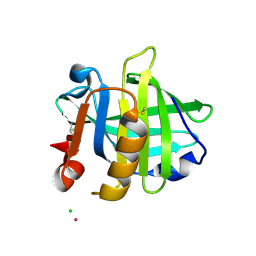 | |
3PH6
 
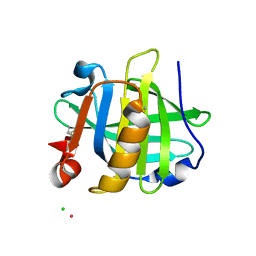 | |
6H5E
 
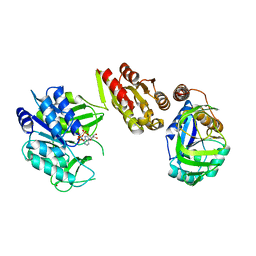 | | Crystal Structure of the GatD/MurT Enzyme Complex from Staphylococcus aureus with bound AMPPNP | | Descriptor: | DI(HYDROXYETHYL)ETHER, DUF1727 domain-containing protein, GLYCEROL, ... | | Authors: | Noeldeke, E.R, Niemann, V, Stoerk, E, Stehle, T. | | Deposit date: | 2018-07-24 | | Release date: | 2018-09-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.139 Å) | | Cite: | Structural basis of cell wall peptidoglycan amidation by the GatD/MurT complex of Staphylococcus aureus.
Sci Rep, 8, 2018
|
|
3O8E
 
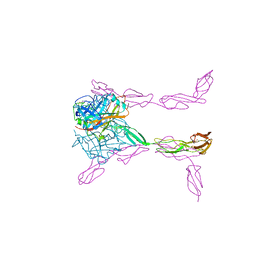 | | Structure of extracelllar portion of CD46 in complex with Adenovirus type 11 knob | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DITHIANE DIOL, Fiber 36.1 kDa protein, ... | | Authors: | Persson, B.D, Schmitz, N.B, Casasnovas, J.M, Stehle, T. | | Deposit date: | 2010-08-03 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Structure of the extracellular portion of CD46 provides insights into its interactions with complement proteins and pathogens.
Plos Pathog., 6, 2010
|
|
6GS2
 
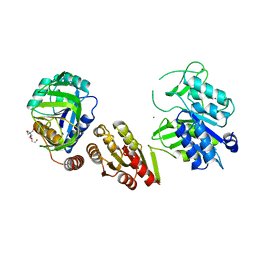 | | Crystal Structure of the GatD/MurT Enzyme Complex from Staphylococcus aureus | | Descriptor: | DI(HYDROXYETHYL)ETHER, MAGNESIUM ION, SA1707 protein, ... | | Authors: | Muckenfuss, L.M, Noeldeke, E.R, Niemann, V, Zocher, G, Stehle, T. | | Deposit date: | 2018-06-13 | | Release date: | 2018-09-05 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Structural basis of cell wall peptidoglycan amidation by the GatD/MurT complex of Staphylococcus aureus.
Sci Rep, 8, 2018
|
|
