3ALY
 
 | |
2YV0
 
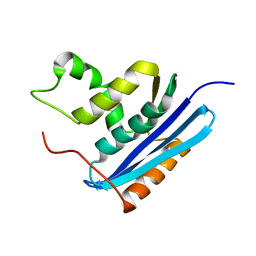
 | | Structural and Thermodynamic Analyses of E. coli ribonuclease HI Variant with Quintuple Thermostabilizing Mutations | | Descriptor: | Ribonuclease HI | | Authors: | Haruki, M, Motegi, T, Tadokoro, T, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2007-04-06 | | Release date: | 2008-03-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural and thermodynamic analyses of Escherichia coli RNase HI variant with quintuple thermostabilizing mutations.
Febs J., 274, 2007
|
|
2Z8Z
 
 | | Crystal structure of a platinum-bound S445C mutant of Pseudomonas sp. MIS38 lipase | | Descriptor: | CALCIUM ION, Lipase, PLATINUM (II) ION, ... | | Authors: | Angkawidjaja, C, You, D.J, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2007-09-13 | | Release date: | 2007-10-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a family I.3 lipase from Pseudomonas sp. MIS38 in a closed conformation
FEBS Lett., 581, 2007
|
|
2ZJ6
 
 | |
2ZJ7
 
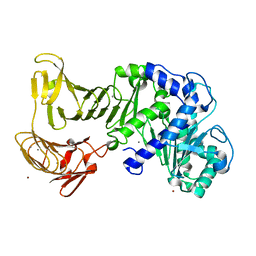 | |
3A70
 
 | | Crystal structure of Pseudomonas sp. MIS38 lipase in complex with diethyl phosphate | | Descriptor: | ACETATE ION, CALCIUM ION, DIETHYL PHOSPHONATE, ... | | Authors: | Angkawidjaja, C, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2009-09-10 | | Release date: | 2010-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | X-ray Crystallographic and MD Simulation Studies on the Mechanism of Interfacial Activation of a Family I.3 Lipase with Two Lids
J.Mol.Biol., 2010
|
|
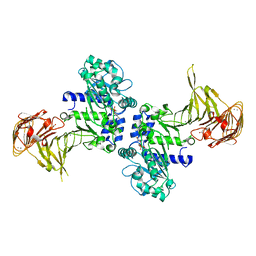
2ZF5
 
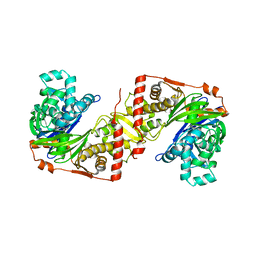 | | Crystal Structure of highly thermostable glycerol kinase from a hyperthermophilic archaeon | | Descriptor: | Glycerol kinase | | Authors: | Koga, Y, Katsumi, R, You, D.-J, Matsumura, H, Takano, K, Kanaya, S. | | Deposit date: | 2007-12-20 | | Release date: | 2008-05-06 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of highly thermostable glycerol kinase from a hyperthermophilic archaeon in a dimeric form
Febs J., 275, 2008
|
|
2ZVD
 
 | | Crystal structure of Pseudomonas sp. MIS38 lipase in an open conformation | | Descriptor: | CALCIUM ION, Lipase | | Authors: | Angkawidjaja, C, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2008-11-05 | | Release date: | 2009-11-10 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | X-ray Crystallographic and MD Simulation Studies on the Mechanism of Interfacial Activation of a Family I.3 Lipase with Two Lids
J.Mol.Biol., 2010
|
|
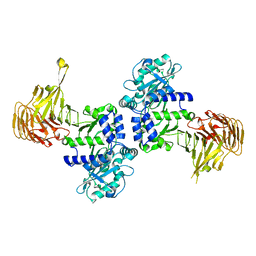
2ZQB
 
 | |
2Z8X
 
 | | Crystal structure of extracellular lipase from Pseudomonas sp. MIS38 | | Descriptor: | CALCIUM ION, Lipase, ZINC ION | | Authors: | Angkawidjaja, C, You, D.J, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2007-09-11 | | Release date: | 2007-10-30 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Crystal structure of a family I.3 lipase from Pseudomonas sp. MIS38 in a closed conformation
FEBS Lett., 581, 2007
|
|
3A6Z
 
 | | Crystal structure of Pseudomonas sp. MIS38 lipase (PML) in the open conformation following dialysis against Ca-free buffer | | Descriptor: | CALCIUM ION, Lipase | | Authors: | Angkawidjaja, C, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2009-09-10 | | Release date: | 2010-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | X-ray Crystallographic and MD Simulation Studies on the Mechanism of Interfacial Activation of a Family I.3 Lipase with Two Lids
J.Mol.Biol., 2010
|
|
3AIM
 
 | | R267E mutant of a HSL-like carboxylesterase from Sulfolobus tokodaii | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, 303aa long hypothetical esterase, ... | | Authors: | Angkawidjaja, C, Kanaya, S. | | Deposit date: | 2010-05-16 | | Release date: | 2011-06-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure and stability of a thermostable carboxylesterase from the thermoacidophilic archaeon Sulfolobustokodaii
Febs J., 279, 2012
|
|
3AIL
 
 | |
3AIK
 
 | | Crystal structure of a HSL-like carboxylesterase from Sulfolobus tokodaii | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, 303aa long hypothetical esterase, ... | | Authors: | Angkawidjaja, C, Kanaya, S. | | Deposit date: | 2010-05-16 | | Release date: | 2011-06-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure and stability of a thermostable carboxylesterase from the thermoacidophilic archaeon Sulfolobustokodaii
Febs J., 279, 2012
|
|
5X7K
 
 | |
1RDB
 
 | |
5NEN
 | |
1RDA
 | |
1RDC
 
 | |
3AOV
 
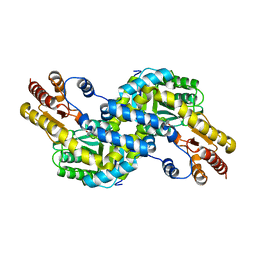 | | Crystal structure of Pyrococcus horikoshii kynurenine aminotransferase in complex with PLP | | Descriptor: | PYRIDOXAL-5'-PHOSPHATE, Putative uncharacterized protein PH0207 | | Authors: | Okada, K, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2010-10-07 | | Release date: | 2011-10-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Characterization of kynurenine aminotransferase from hyperthermophilic archaeon: enzymatic activity for conversion to kynurenic acid is allosterically regulated by alpha-ketoglutaric acid cooperatively with kynurenine
To be Published
|
|
3ATH
 
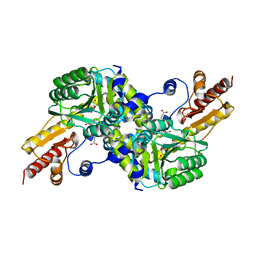 | | Crystal structure of Pyrococcus horikoshii kynurenine aminotransferase in complex with four AKGs as substrates and allosteric effectors | | Descriptor: | 2-OXOGLUTARIC ACID, PYRIDOXAL-5'-PHOSPHATE, Putative uncharacterized protein PH0207 | | Authors: | Okada, K, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2011-01-05 | | Release date: | 2012-01-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Characterization of kynurenine aminotransferase from hyperthermophilic archaeon: enzymatic activity for conversion to kynurenic acid is allosterically regulated by alpha-ketoglutaric acid cooperatively with kynurenine
To be Published
|
|
3AV7
 
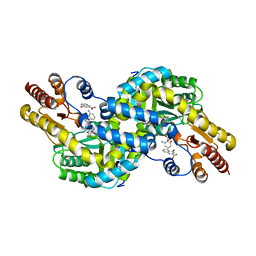 | | Crystal structure of Pyrococcus horikoshii kynurenine aminotransferase in complex with PMP, KYN as substrates and KYA as products | | Descriptor: | (2S)-2-amino-4-(2-aminophenyl)-4-oxobutanoic acid, 4'-DEOXY-4'-AMINOPYRIDOXAL-5'-PHOSPHATE, 4-hydroxyquinoline-2-carboxylic acid, ... | | Authors: | Okada, K, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2011-02-23 | | Release date: | 2012-03-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Structural insight into kynurenic acid excretion mechanisms of kynurenine aminotransferase in the hyperthermophilic archaeon Pyrococcus horikoshii
To be Published
|
|
6K76
 
 | |
6K79
 
 | | Glycerol kinase form Thermococcus kodakarensis, complex structure with substrate. | | Descriptor: | GLYCEROL, Glycerol kinase, TRIETHYLENE GLYCOL | | Authors: | Koga, Y, Angkawidjaja, C, Matsumura, H, Hokao, R. | | Deposit date: | 2019-06-06 | | Release date: | 2020-06-10 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Structural analysis of hexameric structure of glycerol kinase from Thermococcus kodakaraeinsis KOD1
To Be Published
|
|
6K78
 
 | | Glycerol kinase form Thermococcus kodakarensis, complex structure with substrate. | | Descriptor: | GLYCEROL, Glycerol kinase, TRIETHYLENE GLYCOL | | Authors: | Koga, Y, Angkawidjaja, C, Matsumura, H, Hokao, R. | | Deposit date: | 2019-06-06 | | Release date: | 2020-06-10 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.301 Å) | | Cite: | Structural analysis of hexameric structure of glycerol kinase from Thermococcus kodakaraeinsis KOD1
To Be Published
|
|
