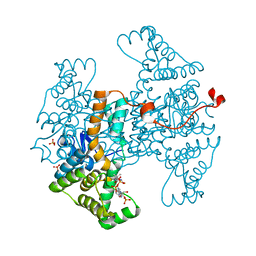
6HXP
 
 | |
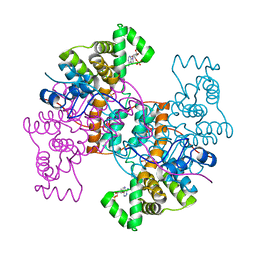
6HXN
 
 | |
6HXH
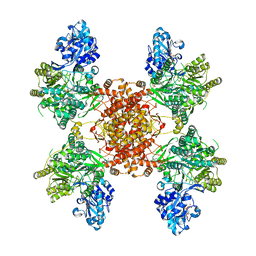 
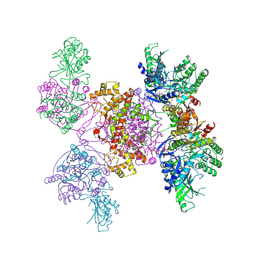
 | | Structure of the human ATP citrate lyase holoenzyme in complex with citrate, coenzyme A and Mg.ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ATP-citrate synthase,Human ATP citrate lyase, CITRATE ANION, ... | | Authors: | Verstraete, K, Verschueren, K. | | Deposit date: | 2018-10-17 | | Release date: | 2019-04-10 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of ATP citrate lyase and the origin of citrate synthase in the Krebs cycle.
Nature, 568, 2019
|
|
6HXI
 
 | |
7NR1
 
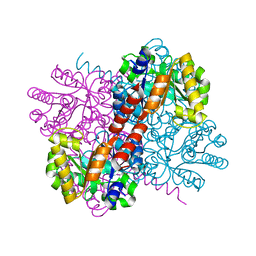 | | Crystal structure of holo-S116A mutant of Hydroxy ketone aldolase (SwHKA) from Sphingomonas wittichii RW1 | | Descriptor: | HpcH/HpaI aldolase, MAGNESIUM ION | | Authors: | Justo, I, Marsden, S.R, Hanefeld, U, Bento, I. | | Deposit date: | 2021-03-02 | | Release date: | 2022-09-21 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Substrate Induced Movement of the Metal Cofactor between Active and Resting State.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
7NUJ
 
 | | Crystal structure of holo-SwHPA-Mg (hydroxy ketone aldolase) from Sphingomonas wittichii RW1 | | Descriptor: | HpcH/HpaI aldolase, MAGNESIUM ION | | Authors: | Laustsen, J, Justo, I, Marsden, S.R, Hanefeld, U, Bento, I. | | Deposit date: | 2021-03-12 | | Release date: | 2022-09-21 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Substrate Induced Movement of the Metal Cofactor between Active and Resting State.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
7O5R
 
 | | Crystal structure of holo-SwHPA-Mn (hydroxyketoacid aldolase) from Sphingomonas wittichii RW1 | | Descriptor: | BROMIDE ION, MANGANESE (II) ION, POTASSIUM ION, ... | | Authors: | Laustsen, J, Justo, I, Marsden, S.R, Hanefeld, U, Bento, I. | | Deposit date: | 2021-04-09 | | Release date: | 2022-10-26 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Substrate Induced Movement of the Metal Cofactor between Active and Resting State.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
7O9R
 
 | | Crystal structure of holo-H44A mutant of Hydroxy ketone aldolase (SwHKA) from Sphingomonas wittichii RW1 | | Descriptor: | BROMIDE ION, DI(HYDROXYETHYL)ETHER, HpcH/HpaI aldolase, ... | | Authors: | Laustsen, J, Justo, I, Marsden, S.R, Hanefeld, U, Bento, I. | | Deposit date: | 2021-04-16 | | Release date: | 2022-10-26 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Substrate Induced Movement of the Metal Cofactor between Active and Resting State.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
7O87
 
 | | Crystal structure of holo-F210W mutant of Hydroxy ketone aldolase (SwHKA) from Sphingomonas wittichii RW1 in complex with hydroxypyruvate | | Descriptor: | 3-HYDROXYPYRUVIC ACID, BROMIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Laustsen, J, Justo, I, Marsden, S.R, Hanefeld, U, Bento, I. | | Deposit date: | 2021-04-14 | | Release date: | 2022-10-26 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Substrate Induced Movement of the Metal Cofactor between Active and Resting State.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
7O5W
 
 | | Crystal structure of holo-F210W mutant of Hydroxy ketone aldolase (SwHKA)from Sphingomonas wittichii RW1 | | Descriptor: | BROMIDE ION, DI(HYDROXYETHYL)ETHER, HpcH/HpaI aldolase, ... | | Authors: | Laustsen, J, Justo, I, Marsden, S.R, Hanefeld, U, Bento, I. | | Deposit date: | 2021-04-09 | | Release date: | 2022-10-26 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Substrate Induced Movement of the Metal Cofactor between Active and Resting State.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
7OBU
 
 | | Crystal structure of holo-F210W mutant of Hydroxy ketone aldolase (SwHKA) from Sphingomonas wittichii RW1, with the active site in the resting and the active state | | Descriptor: | 3-HYDROXYPYRUVIC ACID, HpcH/HpaI aldolase, MAGNESIUM ION, ... | | Authors: | Laustsen, J, Justo, I, Marsden, S.R, Hanefeld, U, Bento, I. | | Deposit date: | 2021-04-23 | | Release date: | 2022-10-26 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Substrate Induced Movement of the Metal Cofactor between Active and Resting State.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
7O5I
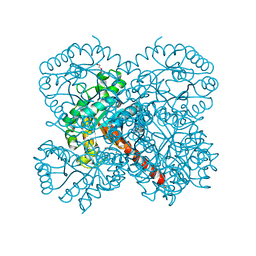 
 | | Crystal structure of apo-SwHKA (Hydroxy ketone aldolase) from Sphingomonas wittichii RW1 | | Descriptor: | BROMIDE ION, DI(HYDROXYETHYL)ETHER, HpcH/HpaI aldolase, ... | | Authors: | Laustsen, J, Justo, I, Marsden, S.R, Hanefeld, U, Bento, I. | | Deposit date: | 2021-04-08 | | Release date: | 2022-11-16 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Substrate Induced Movement of the Metal Cofactor between Active and Resting State.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
7O5V
 
 | | Crystal structure of holo-H44A mutant of Hydroxy ketone aldolase (SwHKA) from Sphingomonas wittichii RW1, in complex with Hydroxypyruvate | | Descriptor: | 3-HYDROXYPYRUVIC ACID, BROMIDE ION, HpcH/HpaI aldolase, ... | | Authors: | Laustsen, J, Justo, I, Marsden, S.R, Hanefeld, U, Bento, I. | | Deposit date: | 2021-04-09 | | Release date: | 2022-11-16 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Substrate Induced Movement of the Metal Cofactor between Active and Resting State.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
2D4M
 
 | |
2D4L
 
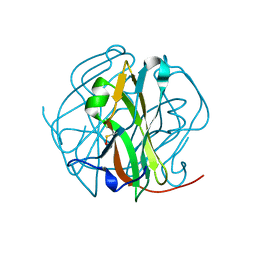 | |
2D4N
 
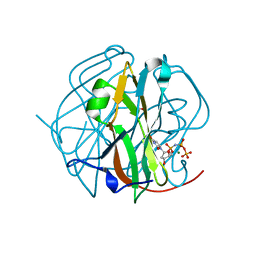 | | Crystal Structure of M-PMV dUTPase complexed with dUPNPP, substrate analogue | | Descriptor: | 2'-DEOXYURIDINE 5'-ALPHA,BETA-IMIDO-TRIPHOSPHATE, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, DU, ... | | Authors: | Nemeth, V, Barabas, O, Vertessy, G.B. | | Deposit date: | 2005-10-20 | | Release date: | 2006-11-21 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Flexible segments modulate co-folding of dUTPase and nucleocapsid proteins.
Nucleic Acids Res., 35, 2007
|
|
3C7X
 
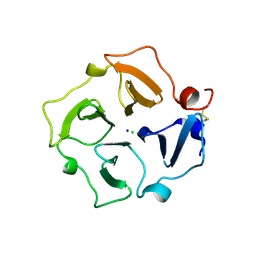 | | Hemopexin-like domain of matrix metalloproteinase 14 | | Descriptor: | CHLORIDE ION, Matrix metalloproteinase-14, SODIUM ION | | Authors: | Tochowicz, A, Itoh, Y, Maskos, K, Bode, W, Goettig, P. | | Deposit date: | 2008-02-08 | | Release date: | 2009-02-10 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The dimer interface of the membrane type 1 matrix metalloproteinase hemopexin domain: crystal structure and biological functions
J.Biol.Chem., 286, 2011
|
|
6EU6
 
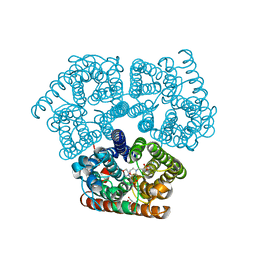 | | Sensor Amt Protein | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, AMMONIUM ION, Ammonium transporter, ... | | Authors: | Pflueger, T, Hernandez, C, Andrade, S.L.A. | | Deposit date: | 2017-10-28 | | Release date: | 2018-01-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Signaling ammonium across membranes through an ammonium sensor histidine kinase.
Nat Commun, 9, 2018
|
|
4MT7
 
 | | Crystal structure of collybistin I | | Descriptor: | Rho guanine nucleotide exchange factor 9 | | Authors: | Schneeberger, D, Schindelin, H. | | Deposit date: | 2013-09-19 | | Release date: | 2014-08-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | A conformational switch in collybistin determines the differentiation of inhibitory postsynapses.
Embo J., 33, 2014
|
|
4MT6
 
 | |
3PNY
 
 | |
3PNV
 
 | |
1FD9
 
 | | CRYSTAL STRUCTURE OF THE MACROPHAGE INFECTIVITY POTENTIATOR PROTEIN (MIP) A MAJOR VIRULENCE FACTOR FROM LEGIONELLA PNEUMOPHILA | | Descriptor: | PROTEIN (MACROPHAGE INFECTIVITY POTENTIATOR PROTEIN), ZINC ION | | Authors: | Riboldi-Tunnicliffe, A, Jessen, S, Konig, B, Rahfeld, J, Hacker, J, Fischer, G, Hilgenfeld, R. | | Deposit date: | 2000-07-20 | | Release date: | 2001-07-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Crystal structure of Mip, a prolylisomerase from Legionella pneumophila
Nat.Struct.Biol., 8, 2001
|
|
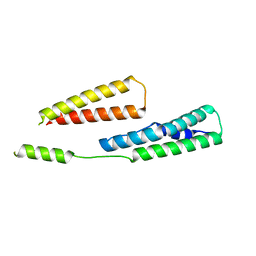
6GV8
 | |
3SY8
 
 | | Crystal structure of the response regulator RocR | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, MAGNESIUM ION, RocR | | Authors: | Chen, M.W, Kotaka, M, Vonrhein, C, Bricogne, G, Lescar, J. | | Deposit date: | 2011-07-16 | | Release date: | 2012-07-18 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insights into the regulatory mechanism of the response regulator RocR from Pseudomonas aeruginosa in cyclic Di-GMP signaling.
J.Bacteriol., 194, 2012
|
|
