9G6K
 
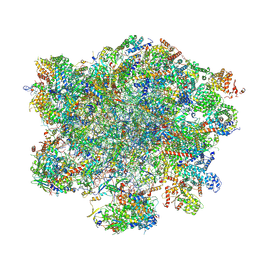 | | LSU structure derived from the LSU sample of the mitoribosome from T. gondii. | | Descriptor: | AMP-dependent synthetase/ligase domain-containing protein, AP2 domain transcription factor AP2IV-1, AP2 domain transcription factor AP2VIIb-2, ... | | Authors: | Rocha, R.E.O, Barua, S, Boissier, F, Nguyen, T.T, Hashem, Y. | | Deposit date: | 2024-07-18 | | Release date: | 2024-12-11 | | Last modified: | 2025-01-01 | | Method: | ELECTRON MICROSCOPY (2.89 Å) | | Cite: | Apicomplexan mitoribosome from highly fragmented rRNAs to a functional machine.
Nat Commun, 15, 2024
|
|
9FIA
 
 | | SSU(body) structure derived from the SSU sample of the mitoribosome from T. gondii. | | Descriptor: | 30S ribosomal protein S12, putative, 30S ribosomal protein S5, ... | | Authors: | Rocha, R.E.O, Barua, S, Boissier, F, Nguyen, T.T, Hashem, Y. | | Deposit date: | 2024-05-28 | | Release date: | 2024-12-11 | | Last modified: | 2025-01-01 | | Method: | ELECTRON MICROSCOPY (3.29 Å) | | Cite: | Apicomplexan mitoribosome from highly fragmented rRNAs to a functional machine.
Nat Commun, 15, 2024
|
|
7YCV
 
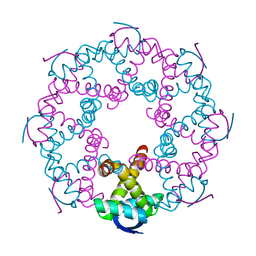 | |
7YCW
 
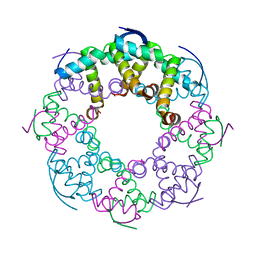 | |
7YCS
 
 | |
7YCU
 
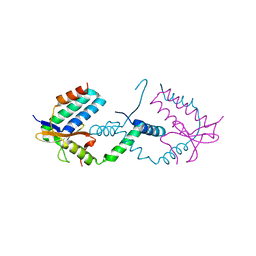 | |
8KGE
 
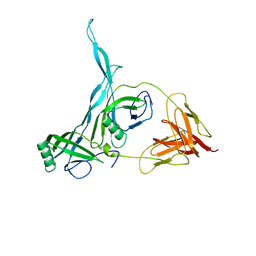 | |
8JVM
 
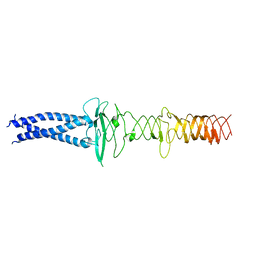 | | AHS-CSF domains of phage lambda tail | | Descriptor: | Tip attachment protein J | | Authors: | Wang, J. | | Deposit date: | 2023-06-28 | | Release date: | 2023-10-18 | | Last modified: | 2024-01-24 | | Method: | ELECTRON MICROSCOPY (3.86 Å) | | Cite: | Architecture of the bacteriophage lambda tail.
Structure, 32, 2024
|
|
9KRQ
 
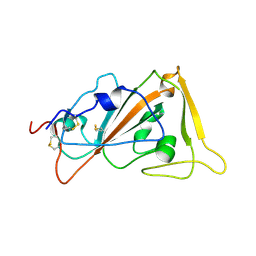 | | Crystal structure of ZXC21 RBD | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein | | Authors: | Lan, J, Wang, C.H. | | Deposit date: | 2024-11-28 | | Release date: | 2025-05-14 | | Method: | X-RAY DIFFRACTION (3.08 Å) | | Cite: | Structural insights into the receptor-binding domain of bat coronavirus ZXC21.
Structure, 2025
|
|
7VI9
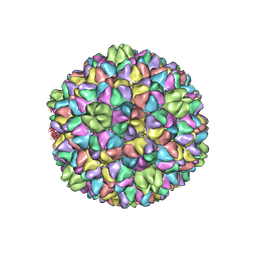 
 | |
7VII
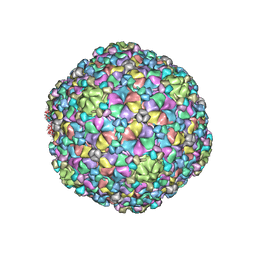 
 | |
7VIA
 
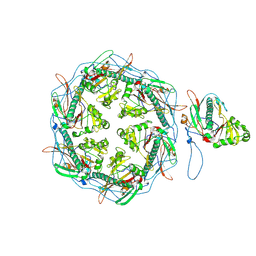 | |
7VIK
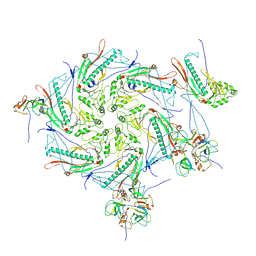 
 | |
7PO5
 
 | |
7PNM
 
 | |
7PNQ
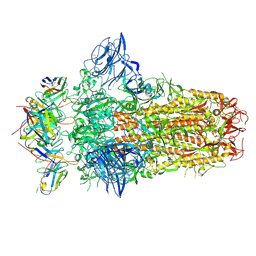 
 | |
8XY9
 
 | | Crystal structure of SARS-CoV-2 BF.7 RBD and human ACE2 complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Processed angiotensin-converting enzyme 2, ... | | Authors: | Lan, J, Wang, C.H. | | Deposit date: | 2024-01-19 | | Release date: | 2025-01-22 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (3.64 Å) | | Cite: | Receptor binding mechanism and immune evasion capacity of SARS-CoV-2 BQ.1.1 lineage.
Virology, 600, 2024
|
|
8XYE
 
 | | Crystal structure of SARS-CoV-2 BA.4 RBD and human ACE2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Processed angiotensin-converting enzyme 2, ... | | Authors: | Lan, J, Wang, C.H. | | Deposit date: | 2024-01-19 | | Release date: | 2025-01-22 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (3.32 Å) | | Cite: | Receptor binding mechanism and immune evasion capacity of SARS-CoV-2 BQ.1.1 lineage.
Virology, 600, 2024
|
|
8XYG
 
 | | Crystal structure of SARS-CoV-2 BQ.1.1 RBD and human ACE2 | | Descriptor: | Processed angiotensin-converting enzyme 2, Spike protein S1 | | Authors: | Lan, J, Wang, C.H. | | Deposit date: | 2024-01-19 | | Release date: | 2025-01-22 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (3.64 Å) | | Cite: | Receptor binding mechanism and immune evasion capacity of SARS-CoV-2 BQ.1.1 lineage.
Virology, 600, 2024
|
|
5XN6
 
 | | Heterodimer crystal structure of geranylgeranyl diphosphate synthases 1 with GGPPS Recruiting Protein(OsGRP) from Oryza sativa | | Descriptor: | Os02g0668100 protein, Os07g0580900 protein | | Authors: | Wang, C, Zhou, F, Lu, S, Zhang, P. | | Deposit date: | 2017-05-18 | | Release date: | 2017-06-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.598 Å) | | Cite: | A recruiting protein of geranylgeranyl diphosphate synthase controls metabolic flux toward chlorophyll biosynthesis in rice
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
5XN5
 
 | |
6DJ4
 
 | |
8WJB
 
 | |
6KI2
 
 | |
6KI1
 
 | |
