6WQN
 
 | |
6V81
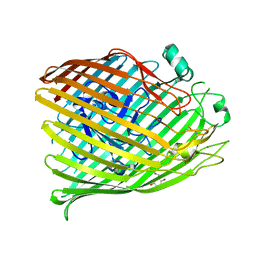 
 | | The crystal structure of the outer-membrane transporter YncD | | Descriptor: | CALCIUM ION, Probable TonB-dependent receptor YncD, octyl beta-D-glucopyranoside | | Authors: | Grinter, R. | | Deposit date: | 2019-12-10 | | Release date: | 2020-05-06 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.957 Å) | | Cite: | The crystal structure of the TonB-dependent transporter YncD reveals a positively charged substrate-binding site.
Acta Crystallogr D Struct Biol, 76, 2020
|
|
6V4V
 
 | |
6WQQ
 
 | |
6WRU
 
 | |
6WRS
 
 | |
6B05
 
 | |
6B03
 
 | |
5VTG
 
 | |
6BPM
 
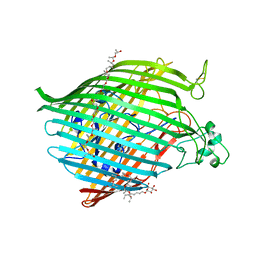 | | The crystal structure of the Ferric-Catecholate import receptor Fiu from K12 E. coli: Closed form (C21) | | Descriptor: | (20S)-2,5,8,11,14,17-HEXAMETHYL-3,6,9,12,15,18-HEXAOXAHENICOSANE-1,20-DIOL, Catecholate siderophore receptor Fiu, octyl beta-D-glucopyranoside | | Authors: | Grinter, R. | | Deposit date: | 2017-11-23 | | Release date: | 2018-11-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The structure of the bacterial iron-catecholate transporter Fiu suggests that it imports substrates via a two-step mechanism.
J.Biol.Chem., 2019
|
|
6BRS
 
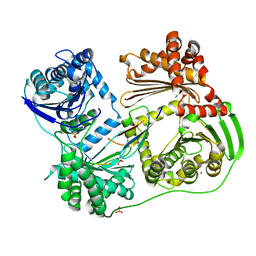 | |
6BPN
 
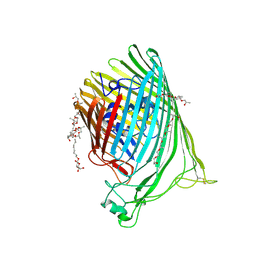 | | The crystal structure of the Ferric-Catecholate import receptor Fiu from E. coli K12: Open form (C2221) | | Descriptor: | (20S)-2,5,8,11,14,17-HEXAMETHYL-3,6,9,12,15,18-HEXAOXAHENICOSANE-1,20-DIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, ... | | Authors: | Grinter, R. | | Deposit date: | 2017-11-23 | | Release date: | 2018-11-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The structure of the bacterial iron-catecholate transporter Fiu suggests that it imports substrates via a two-step mechanism.
J.Biol.Chem., 2019
|
|
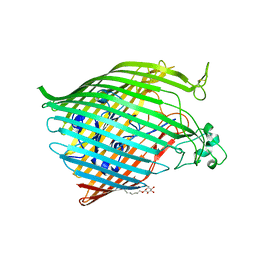
6BPO
 | |
