5D1H
 
 | |
5EOV
 
 | |
4OX9
 
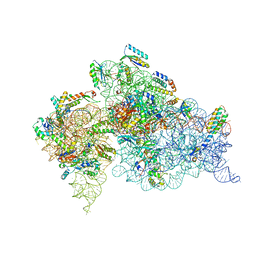 | | Crystal structure of the aminoglycoside resistance methyltransferase NpmA bound to the 30S ribosomal subunit | | Descriptor: | 16S rRNA, 16S rRNA (adenine(1408)-N(1))-methyltransferase, 30S ribosomal protein S10, ... | | Authors: | Dunkle, J.A, Conn, G.L, Dunham, C.M. | | Deposit date: | 2014-02-04 | | Release date: | 2014-04-09 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.8035 Å) | | Cite: | Molecular recognition and modification of the 30S ribosome by the aminoglycoside-resistance methyltransferase NpmA.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
404D
 
 | |
4RX1
 
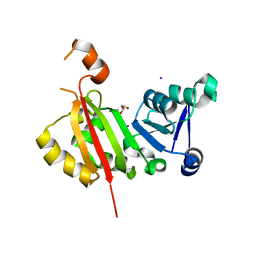 | | Crystal Structure of antibiotic-resistance methyltransferase Kmr | | Descriptor: | GLYCEROL, IODIDE ION, Putative rRNA methyltransferase | | Authors: | Savic, M. | | Deposit date: | 2014-12-08 | | Release date: | 2015-03-11 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.472 Å) | | Cite: | 30S Subunit-Dependent Activation of the Sorangium cellulosum So ce56 Aminoglycoside Resistance-Conferring 16S rRNA Methyltransferase Kmr.
Antimicrob.Agents Chemother., 59, 2015
|
|
3PYJ
 
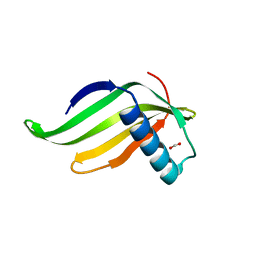 | | Reduced sweetness of a monellin (MNEI) mutant results from increased protein flexibility and disruption of a distant poly-(L-proline) II helix | | Descriptor: | FORMIC ACID, Monellin chain B/Monellin chain A chimeric protein | | Authors: | Hobbs, J.R, Conn, G.C, Munger, S.D, Templeton, C.M. | | Deposit date: | 2010-12-13 | | Release date: | 2011-04-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Reduced Sweetness of a Monellin (MNEI) Mutant Results from Increased Protein Flexibility and Disruption of a Distant Poly-(L-Proline) II Helix.
Chem Senses, 36, 2011
|
|
3MQ2
 
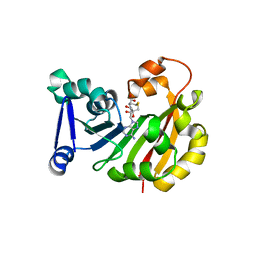 | | Crystal Structure of 16S rRNA Methyltranferase KamB | | Descriptor: | 16S rRNA methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Macmaster, R.A. | | Deposit date: | 2010-04-27 | | Release date: | 2010-12-08 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Structural insights into the function of aminoglycoside-resistance A1408 16S rRNA methyltransferases from antibiotic-producing and human pathogenic bacteria.
Nucleic Acids Res., 38, 2010
|
|
4MCE
 
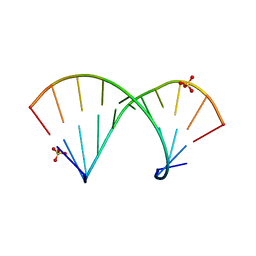 | | Crystal structure of the Gas5 GRE Mimic | | Descriptor: | Gas5 GREM Fwd, Gas5 GREM Rev, SULFATE ION | | Authors: | Hudson, W.H, Ortlund, E.A. | | Deposit date: | 2013-08-21 | | Release date: | 2014-09-24 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.205 Å) | | Cite: | Conserved sequence-specific lincRNA-steroid receptor interactions drive transcriptional repression and direct cell fate.
Nat Commun, 5, 2014
|
|
4MCF
 
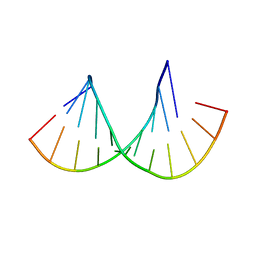 | | Crystal structure of the Gas5 GRE Mimic | | Descriptor: | Gas5 GREM Fwd, Gas5 GREM Rev, SULFATE ION | | Authors: | Hudson, W.H, Ortlund, E.A. | | Deposit date: | 2013-08-21 | | Release date: | 2014-11-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.899 Å) | | Cite: | Conserved sequence-specific lincRNA-steroid receptor interactions drive transcriptional repression and direct cell fate.
Nat Commun, 5, 2014
|
|
4RWZ
 
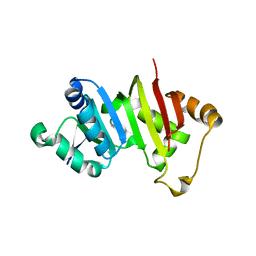 | | Crystal structure of the antibiotic-resistance methyltransferase Kmr | | Descriptor: | Putative rRNA methyltransferase | | Authors: | Savic, M. | | Deposit date: | 2014-12-08 | | Release date: | 2015-03-11 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.801 Å) | | Cite: | 30S Subunit-Dependent Activation of the Sorangium cellulosum So ce56 Aminoglycoside Resistance-Conferring 16S rRNA Methyltransferase Kmr.
Antimicrob.Agents Chemother., 59, 2015
|
|
1OLN
 
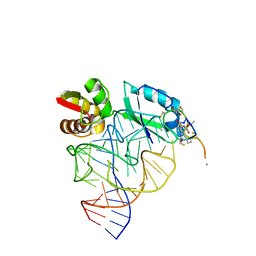 | | Model for thiostrepton antibiotic binding to L11 substrate from 50S ribosomal RNA | | Descriptor: | 50S RIBOSOMAL PROTEIN L11, RNA, THIOSTREPTON | | Authors: | Lentzen, G, Klinck, R, Matassova, N, Aboul-Ela, F, Murchie, A.I.H. | | Deposit date: | 2003-08-08 | | Release date: | 2003-09-11 | | Last modified: | 2019-08-21 | | Method: | SOLUTION NMR, THEORETICAL MODEL | | Cite: | Structural Basis for Contrasting Activities of Ribosome Binding Thiazole Antibiotics
Chem.Biol., 10, 2003
|
|
