9ATX
 
 | |
9P6P
 
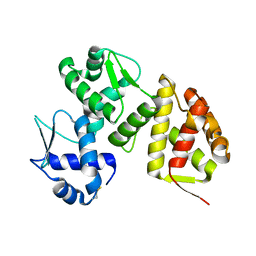
 | | Crystal Structure of the SARS-CoV-2 2'-O-Methyltransferase with (m7GpppA)pUpU (Cap-0) and S-Adenosyl-L-homocysteine (SAH). | | Descriptor: | 2'-O-methyltransferase, 7N-METHYL-8-HYDROGUANOSINE-5'-TRIPHOSPHATE, CHLORIDE ION, ... | | Authors: | Minasov, G, Shuvalova, L, Maltseva, N, Kim, Y, Kiryukhina, O, Joachimiak, A, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2025-06-19 | | Release date: | 2025-07-02 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure of the SARS-CoV-2 2'-O-Methyltransferase with (m7GpppA)pUpU (Cap-0) and S-Adenosyl-L-homocysteine (SAH).
To Be Published
|
|
9OVI
 
 | | Crystal Structure of SH3-like_bac-type domain (79-145) of Conserved domain protein GBAA_2967 from Bacillus anthracis Ames ancestor | | Descriptor: | Conserved domain protein, FORMIC ACID, MAGNESIUM ION | | Authors: | Minasov, G, Dubrovska, I, Winsor, J, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2025-05-30 | | Release date: | 2025-06-11 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of SH3-like_bac-type domain (79-145) of Conserved domain protein GBAA_2967 from Bacillus anthracis Ames ancestor.
To Be Published
|
|
9OW2
 
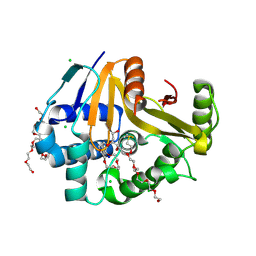 | | Crystal Structure of the Surface Protein (CD630_07380) from Clostridium difficile Strain 630 | | Descriptor: | 1,2-ETHANEDIOL, 3,6,9,12,15,18,21,24,27,30,33,36,39-TRIDECAOXAHENTETRACONTANE-1,41-DIOL, CHLORIDE ION, ... | | Authors: | Minasov, G, Shuvalova, L, Kiryukhina, O, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2025-06-02 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal Structure of the Surface Protein (CD630_07380) from Clostridium difficile Strain 630.
To Be Published
|
|
9OWG
 
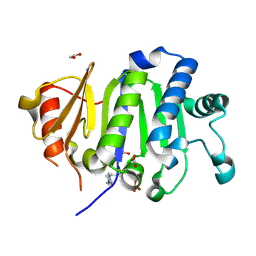 | | Crystal Structure of the Immunity Protein (52 domain-containing protein) from Pseudomonas aeruginosa PAO1. | | Descriptor: | 1,2-ETHANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CARBONATE ION, ... | | Authors: | Minasov, G, Wawrzak, Z, Skarina, T, Stogios, P.J, Savchenko, A, Satchell, K.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2025-06-02 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal Structure of the Immunity Protein (52 domain-containing protein) from Pseudomonas aeruginosa PAO1.
To Be Published
|
|
9NZB
 
 | | Crystal Structure of acyl-CoA lyase subunit beta from Pseudomonas aeruginosa PAO1 | | Descriptor: | Acyl-CoA lyase beta chain | | Authors: | Minasov, G, Shukla, S, Shuvalova, L, Brunzelle, J.S, Warwzak, Z, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2025-03-31 | | Release date: | 2025-07-09 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal Structure of acyl-CoA lyase subunit beta from Pseudomonas aeruginosa PAO1
To Be Published
|
|
9P0F
 
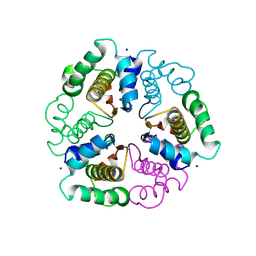 | |
9DWP
 
 | | Crystal Structure of C4-Dicarboxylate-Binding Protein (PA0884) of Tripartite ATP-independent Periplasmic Transporter Family from Pseudomonas aeruginosa PAO1 in Complex with Mesaconic Acid | | Descriptor: | (2E)-2-METHYLBUT-2-ENEDIOIC ACID, C4-dicarboxylate-binding protein, CHLORIDE ION, ... | | Authors: | Minasov, G, Shukla, S, Shuvalova, L, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2024-10-09 | | Release date: | 2025-07-09 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Crystal Structure of C4-Dicarboxylate-Binding Protein (PA0884) of Tripartite ATP-independent Periplasmic Transporter Family from Pseudomonas aeruginosa PAO1 in Complex with Mesaconic Acid
To Be Published
|
|
9DWO
 
 | |
9DY3
 
 | | Crystal Structure of C4-Dicarboxylate-Binding Periplasmic Protein (PA5167) of Tripartite ATP-independent Periplasmic Transporter Family from Pseudomonas aeruginosa PAO1 in Complex with L-Malate | | Descriptor: | (2S)-2-hydroxybutanedioic acid, C4-dicarboxylate-binding periplasmic protein DctP, ZINC ION | | Authors: | Minasov, G, Shukla, S, Shuvalova, L, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2024-10-12 | | Release date: | 2025-07-09 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal Structure of C4-Dicarboxylate-Binding Periplasmic Protein (PA5167) of Tripartite ATP-independent Periplasmic Transporter Family from Pseudomonas aeruginosa PAO1 in Complex with L-Malate.
To Be Published
|
|
8VDK
 
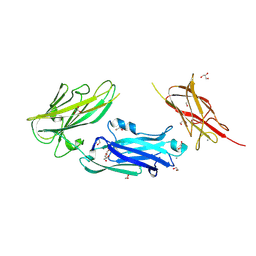 | | Crystal Structure of LPXTG-motif Cell Wall Anchor Domain Protein MSCRAMM_SdrD from Staphylococcus aureus | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, GLYCEROL, ... | | Authors: | Kim, Y, Tan, A, Endres, M, Joachimiak, A, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2023-12-15 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of LPXTG-motif Cell Wall Anchor Domain Protein MSCRAMM_SdrD from Staphylococcus aureus
To Be Published
|
|
8G1Y
 
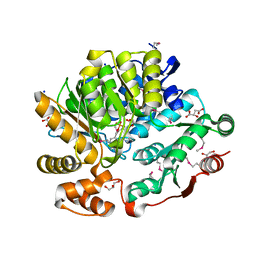 | | Crystal Structure of the Threonine Synthase from Streptococcus pneumoniae in complex with Pyridoxal 5-phosphate. | | Descriptor: | CHLORIDE ION, D-MALATE, GLYCEROL, ... | | Authors: | Minasov, G, Shuvalova, L, Kiryukhina, O, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID), Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2023-02-03 | | Release date: | 2023-02-15 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Crystal Structure of the Threonine Synthase from Streptococcus pneumoniae in complex with Pyridoxal 5-phosphate.
To Be Published
|
|
8G28
 
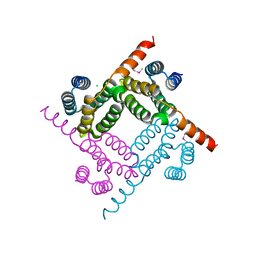 | | Crystal Structure of the C-terminal Fragment of AAA ATPase from Streptococcus pneumoniae. | | Descriptor: | ATPase, AAA family, CHLORIDE ION | | Authors: | Minasov, G, Shuvalova, L, Brunzelle, J.S, Kiryukhina, O, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID), Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2023-02-03 | | Release date: | 2023-02-15 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal Structure of the C-terminal Fragment of AAA ATPase from Streptococcus pneumoniae.
To Be Published
|
|
8G22
 
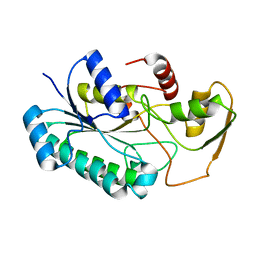 | | Crystal Structure of the dTDP-4-dehydrorhamnose Reductase from Streptococcus pneumoniae. | | Descriptor: | dTDP-4-dehydrorhamnose reductase | | Authors: | Minasov, G, Shuvalova, L, Brunzelle, J.S, Kiryukhina, O, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID), Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2023-02-03 | | Release date: | 2023-02-22 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | Crystal Structure of the dTDP-4-dehydrorhamnose Reductase from Streptococcus pneumoniae.
To Be Published
|
|
8G62
 
 | | Papain-Like Protease of SARS CoV-2 in complex with remodilin NCGC 390004 | | Descriptor: | 3-methoxy-5-(1-methylpiperidin-4-yl)-N-[4-(pyrrolidine-1-sulfonyl)phenyl]benzamide, ACETATE ION, CHLORIDE ION, ... | | Authors: | Osipiuk, J, Tesar, C, Endres, M, Jedrzejczak, R, Luci, D, Kales, S, Simeonov, A, Rai, G, Drayman, N, Tay, S, Oakes, S, Rosner, M, Chen, B, Dulin, N, Solway, J, Joachimiak, A, Center for Structural Genomics of Infectious Diseases (CSGID), Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2023-02-14 | | Release date: | 2023-02-22 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Papain-Like Protease of SARS CoV-2 in complex with remodilin NCGC 390004
To Be Published
|
|
9DSB
 
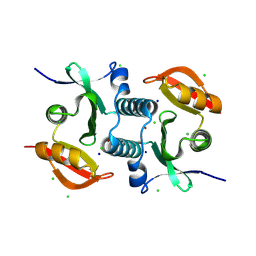 | | Crystal Structure of Spermin/spermidine N-Acetyltransferase from Enterococcus faecalis V583 | | Descriptor: | ACETATE ION, CHLORIDE ION, POTASSIUM ION, ... | | Authors: | Kim, Y, Maltseva, N, Endres, M, Kuhn, M, Joachimiak, A, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2024-09-26 | | Release date: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of Spermin/spermidine N-Acetyltransferase from Enterococcus faecalis V583
To Be Published
|
|
8F8O
 
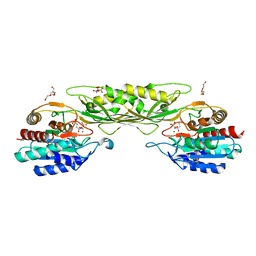 | | Crystal Structure of the Succinyl-diaminopimelate Desuccinylase (DapE) from Acinetobacter baumannii in complex with Succinic and L-Lactic Acids | | Descriptor: | (2S)-2-HYDROXYPROPANOIC ACID, CITRIC ACID, SUCCINIC ACID, ... | | Authors: | Minasov, G, Shuvalova, L, Brunzelle, J.S, Dubrovska, I, Pshenychnyi, S, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID), Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2022-11-22 | | Release date: | 2022-11-30 | | Last modified: | 2023-02-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structure of the Succinyl-diaminopimelate Desuccinylase (DapE) from Acinetobacter baumannii in complex with Succinic and L-Lactic Acids
To Be Published
|
|
8SFG
 
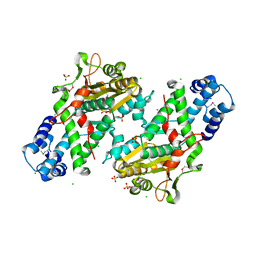 | | Crystal Structure of the Open Unbound Catalytically Inactive Makes Caterpillars Floppy-like (MCF) Effector from Vibrio vulnificus CMCP6 | | Descriptor: | Autotransporter adhesin, CHLORIDE ION, SULFATE ION | | Authors: | Minasov, G, Shuvalova, L, Rosas-Lemus, M, Herrera, A, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2023-04-11 | | Release date: | 2024-06-05 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure of the Open Unbound Catalytically Inactive Makes Caterpillars Floppy-like (MCF) Effector from Vibrio vulnificus CMCP6.
To Be Published
|
|
8VVA
 
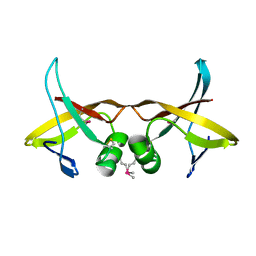 | |
7R6S
 
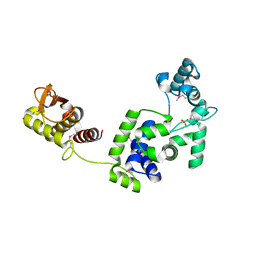 | | Crystal Structure of the Putative Bacteriophage Protein from Stenotrophomonas maltophilia | | Descriptor: | Putative bacteriophage protein, SULFATE ION | | Authors: | Minasov, G, Shuvalova, L, Kiryukhina, O, Brunzelle, J.S, Wiersum, G, Satchell, K.J.F, Center for Structural Genomics of Infectious Diseases (CSGID), Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2021-06-23 | | Release date: | 2022-11-09 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of the Putative Bacteriophage Protein from Stenotrophomonas maltophilia
To Be Published
|
|
9DSY
 
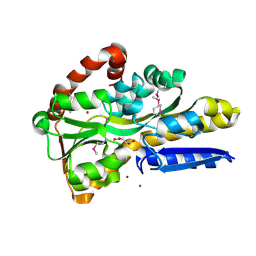 | | Crystal Structure of C4-Dicarboxylate-Binding Periplasmic Protein (PA5167) of Tripartite ATP-independent Periplasmic Transporter Family from Pseudomonas aeruginosa PAO1 in Complex with Succinic Acid | | Descriptor: | C4-dicarboxylate-binding periplasmic protein DctP, SUCCINIC ACID, ZINC ION | | Authors: | Minasov, G, Shukla, S, Shuvalova, L, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2024-09-30 | | Release date: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal Structure of C4-Dicarboxylate-Binding Periplasmic Protein (PA5167) of Tripartite ATP-independent Periplasmic Transporter Family from Pseudomonas aeruginosa PAO1 in Complex with Succinic Acid
To Be Published
|
|
9DTL
 
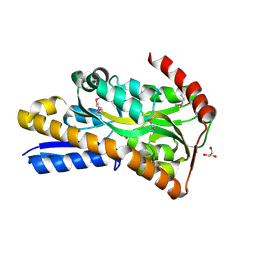 | | Crystal Structure of C4-Dicarboxylate-Binding Protein (PA0884) of Tripartite ATP-independent Periplasmic Transporter Family from Pseudomonas aeruginosa PAO1 in Complex with Succinic Acid | | Descriptor: | C4-dicarboxylate-binding periplasmic protein, GLYCEROL, SUCCINIC ACID | | Authors: | Minasov, G, Shukla, S, Shuvalova, L, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2024-10-01 | | Release date: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal Structure of C4-Dicarboxylate-Binding Protein (PA0884) of Tripartite ATP-independent Periplasmic Transporter Family from Pseudomonas aeruginosa PAO1 in Complex with Succinic Acid
To Be Published
|
|
9BPW
 
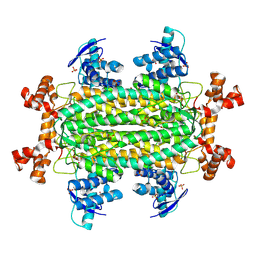 | | Crystal Structure of Fumarate Hydratase FumC from Staphylococcus aureus | | Descriptor: | CHLORIDE ION, FORMIC ACID, FUMARIC ACID, ... | | Authors: | Kim, Y, Tesar, C, Endres, M, Babnigg, G, Wong, T, Joachimiak, A, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2024-05-08 | | Release date: | 2025-05-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of Fumarate Hydratase FumC from Staphylococcus aureus
To Be Published
|
|
9C3D
 
 | | Crystal structure of metallo-beta-lactamase superfamily protein CcrA-like_MBL-B1 from Dyadobacter fermentans | | Descriptor: | Metallo-beta-lactamase type 2, ZINC ION | | Authors: | Kim, Y, Maltseva, N, Welk, L, Endres, M, Joachimiak, A, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2024-05-31 | | Release date: | 2024-07-17 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure of metallo-beta-lactamase superfamily protein CcrA-like_MBL-B1 from Dyadobacter fermentans
To Be Published
|
|
9DCG
 
 | | Crystal Structure of the Thiol:Disulfide Interchange Protein DsbC from Vibrio cholerae | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, FORMIC ACID, ... | | Authors: | Kim, Y, Maltseva, N, Shatsman, S, Joachimiak, A, Center for Structural Biology of Infectious Diseases (CSBID) | | Deposit date: | 2024-08-26 | | Release date: | 2024-09-04 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Crystal Structure of the Thiol:Disulfide Interchange Protein DsbC from Vibrio cholerae
To Be Published
|
|
