3I14
 
 | |
3I13
 
 | |
3PJQ
 
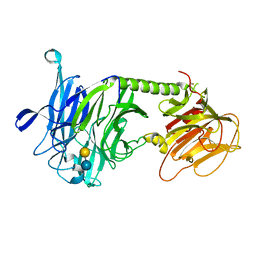 | | Trypanosoma cruzi trans-sialidase-like inactive isoform (including the natural mutation Tyr342His) in complex with lactose | | Descriptor: | Trans-sialidase, beta-D-galactopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Oppezzo, P, Baraibar, M, Obal, G, Pritsch, O, Alzari, P.M, Buschiazzo, A. | | Deposit date: | 2010-11-10 | | Release date: | 2011-06-08 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of an enzymatically inactive trans-sialidase-like lectin from Trypanosoma cruzi: the carbohydrate binding mechanism involves residual sialidase activity.
Biochim.Biophys.Acta, 1814, 2011
|
|
3PG1
 
 | |
5OU5
 
 | |
4UHS
 
 | |
4UHT
 
 | |
6B0D
 
 | | An E. coli DPS protein from ferritin superfamily | | Descriptor: | DNA protection during starvation protein, FORMIC ACID, SODIUM ION | | Authors: | Rui, W, Ruslan, S, Ronan, K, Adam, J.S. | | Deposit date: | 2017-09-14 | | Release date: | 2018-09-19 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | SIMBAD: a sequence-independent molecular-replacement pipeline.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
6B6M
 
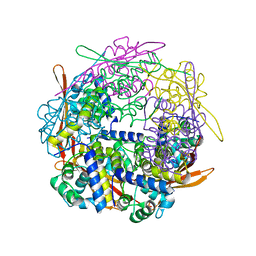 | | Cyanase from Serratia proteamaculans | | Descriptor: | Cyanate hydratase | | Authors: | Xu, Y. | | Deposit date: | 2017-10-02 | | Release date: | 2017-10-25 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | SIMBAD: a sequence-independent molecular-replacement pipeline.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
4UHJ
 
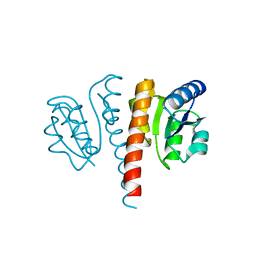 | |
4UHK
 
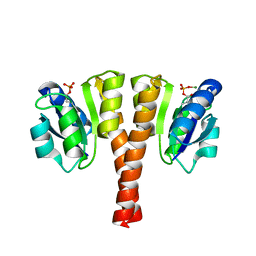 | |
5LFK
 
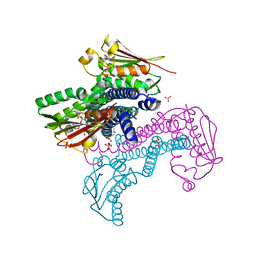 | | Crystal structure of CpxAHDC (hemiphosphorylated form) | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, SULFATE ION, ... | | Authors: | Mechaly, A.E, Alzari, P.M. | | Deposit date: | 2016-07-01 | | Release date: | 2017-06-07 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.094 Å) | | Cite: | Structural Coupling between Autokinase and Phosphotransferase Reactions in a Bacterial Histidine Kinase.
Structure, 25, 2017
|
|
