8BXO
 
 | | fragment-linked stabilizer for ERa - 14-3-3 interaction (1075287) | | Descriptor: | 14-3-3 protein sigma, ERalpha peptide, ~{N}-[3-(5-carbamimidoylthiophen-3-yl)phenyl]-4,4-bis(fluoranyl)-1-phenoxy-cyclohexane-1-carboxamide | | Authors: | Visser, E.J, Vandenboorn, E.M.F, Ottmann, C. | | Deposit date: | 2022-12-09 | | Release date: | 2023-08-02 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | From Tethered to Freestanding Stabilizers of 14-3-3 Protein-Protein Interactions through Fragment Linking.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8BXS
 
 | |
8BYC
 
 | | fragment-linked stabilizer for ERa - 14-3-3 interaction (1075316) | | Descriptor: | 14-3-3 protein sigma, ERalpha peptide, MAGNESIUM ION, ... | | Authors: | Visser, E.J, Vandenboorn, E.M.F, Ottmann, C. | | Deposit date: | 2022-12-12 | | Release date: | 2023-08-02 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | From Tethered to Freestanding Stabilizers of 14-3-3 Protein-Protein Interactions through Fragment Linking.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8BYF
 
 | | fragment-linked stabilizer for ERa - 14-3-3 interaction (1047455) | | Descriptor: | 14-3-3 protein sigma, PHE-PRO-ALA-TPO-VAL, ~{N}-[3-(5-carbamimidoylthiophen-3-yl)phenyl]-2-(4-chloranylphenoxy)-2-methyl-propanamide | | Authors: | Visser, E.J, Sijbesma, E, Ottmann, C. | | Deposit date: | 2022-12-12 | | Release date: | 2023-08-02 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | From Tethered to Freestanding Stabilizers of 14-3-3 Protein-Protein Interactions through Fragment Linking.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8BWJ
 
 | |
8BWZ
 
 | |
8BX0
 
 | |
8BXN
 
 | |
8BXQ
 
 | |
8BYB
 
 | |
8BXI
 
 | | fragment-linked stabilizer for ERa - 14-3-3 interaction (1074361) | | Descriptor: | 14-3-3 protein sigma, ERalpha peptide, MAGNESIUM ION, ... | | Authors: | Visser, E.J, Vandenboorn, E.M.F, Ottmann, C. | | Deposit date: | 2022-12-08 | | Release date: | 2023-08-02 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | From Tethered to Freestanding Stabilizers of 14-3-3 Protein-Protein Interactions through Fragment Linking.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8BWX
 
 | | Fragment-linked stabilizer for 14-3-3 and ERa (1075298) | | Descriptor: | 14-3-3 protein sigma, 4-azanyl-~{N}-[3-(5-carbamimidoylthiophen-3-yl)phenyl]-1-(4-chloranylphenoxy)cyclohexane-1-carboxamide, ERalpha peptide | | Authors: | Visser, E.J, Vandenboorn, E.M.F, Ottmann, C. | | Deposit date: | 2022-12-07 | | Release date: | 2023-08-02 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | From Tethered to Freestanding Stabilizers of 14-3-3 Protein-Protein Interactions through Fragment Linking.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8BYE
 
 | |
8BXM
 
 | | fragment-linked stabilizer for ERa - 14-3-3 interaction (1074397) | | Descriptor: | 14-3-3 protein sigma, ERalpha peptide, MAGNESIUM ION, ... | | Authors: | Visser, E.J, Vandenboorn, E.M.F, Ottmann, C. | | Deposit date: | 2022-12-09 | | Release date: | 2023-08-02 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | From Tethered to Freestanding Stabilizers of 14-3-3 Protein-Protein Interactions through Fragment Linking.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8BX3
 
 | | fragment-linked stabilizer for ERa - 14-3-3 interaction (1074372) | | Descriptor: | 14-3-3 protein sigma, 2-(4-bromanylphenoxy)-~{N}-[3-(5-carbamimidoylthiophen-3-yl)phenyl]-2-methyl-propanamide, ERalpha peptide, ... | | Authors: | Visser, E.J, Vandenboorn, E.M.F, Ottmann, C. | | Deposit date: | 2022-12-07 | | Release date: | 2023-08-02 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | From Tethered to Freestanding Stabilizers of 14-3-3 Protein-Protein Interactions through Fragment Linking.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8BYD
 
 | |
8BY9
 
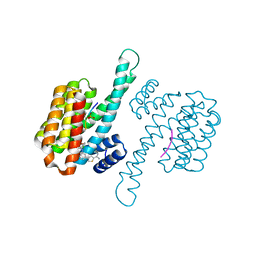 | | fragment-linked stabilizer for ERa - 14-3-3 interaction (1075292) | | Descriptor: | 14-3-3 protein sigma, ERalpha peptide, ~{N}-[3-(5-carbamimidoylthiophen-3-yl)phenyl]-1-(4-chloranylphenoxy)-4,4-bis(fluoranyl)cyclohexane-1-carboxamide | | Authors: | Visser, E.J, Vandenboorn, E.M.F, Ottmann, C. | | Deposit date: | 2022-12-12 | | Release date: | 2023-08-02 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | From Tethered to Freestanding Stabilizers of 14-3-3 Protein-Protein Interactions through Fragment Linking.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8BYG
 
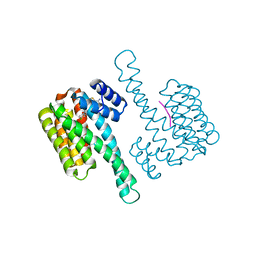 | | fragment-linked stabilizer for ERa - 14-3-3 interaction (1047648) | | Descriptor: | 14-3-3 protein sigma, ERalpha peptide, ~{N}-[2-[(2-carbamimidoyl-1-benzothiophen-4-yl)-methyl-amino]ethyl]-2-(4-chloranylphenoxy)-~{N},2-dimethyl-propanamide | | Authors: | Visser, E.J, Sijbesma, E, Ottmann, C. | | Deposit date: | 2022-12-12 | | Release date: | 2023-08-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | From Tethered to Freestanding Stabilizers of 14-3-3 Protein-Protein Interactions through Fragment Linking.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
5SQT
 
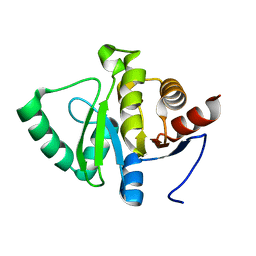 | | PanDDA analysis group deposition -- Crystal structure of SARS-CoV-2 NSP3 macrodomain in complex with ZINC000833624464 - (R,R) and (S,S) isomers | | Descriptor: | (3R,4R)-4-methyl-1-(2-oxo-2,3-dihydro-1,3-benzoxazole-7-carbonyl)pyrrolidine-3-carboxylic acid, (3S,4S)-4-methyl-1-(2-oxo-2,3-dihydro-1,3-benzoxazole-7-carbonyl)pyrrolidine-3-carboxylic acid, Non-structural protein 3 | | Authors: | Correy, G.J, Fraser, J.S. | | Deposit date: | 2022-06-09 | | Release date: | 2022-07-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Iterative computational design and crystallographic screening identifies potent inhibitors targeting the Nsp3 macrodomain of SARS-CoV-2.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
5SR2
 
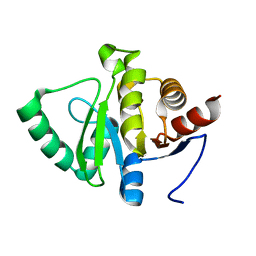 | |
5SQ4
 
 | |
5SQI
 
 | | PanDDA analysis group deposition -- Crystal structure of SARS-CoV-2 NSP3 macrodomain in complex with Z5016127255 - (R,R) and (S,S) isomers | | Descriptor: | (1R,2R)-4-hydroxy-1-[4-(methylcarbamamido)benzamido]-2,3-dihydro-1H-indene-2-carboxylic acid, (1S,2S)-4-hydroxy-1-[4-(methylcarbamamido)benzamido]-2,3-dihydro-1H-indene-2-carboxylic acid, Non-structural protein 3 | | Authors: | Correy, G.J, Fraser, J.S. | | Deposit date: | 2022-06-09 | | Release date: | 2022-07-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Iterative computational design and crystallographic screening identifies potent inhibitors targeting the Nsp3 macrodomain of SARS-CoV-2.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
5SR1
 
 | |
5SR5
 
 | |
5SPZ
 
 | |
