6FG7
 
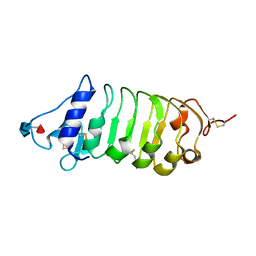 | | Crystal structure of the BIR2 ectodomain from Arabidopsis thaliana. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Inactive LRR receptor-like serine/threonine-protein kinase BIR2, ... | | Authors: | Hothorn, M, Hohmann, U. | | Deposit date: | 2018-01-10 | | Release date: | 2018-01-24 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The SERK3 elongated allele defines a role for BIR ectodomains in brassinosteroid signalling.
Nat Plants, 4, 2018
|
|
6FG8
 
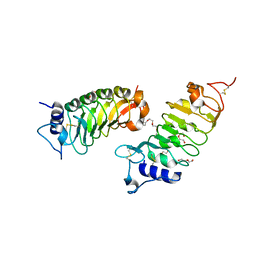 | | Crystal structure of the BIR3 - SERK1 complex from Arabidopsis thaliana. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DI(HYDROXYETHYL)ETHER, PENTAETHYLENE GLYCOL, ... | | Authors: | Hothorn, M, Hohmann, U. | | Deposit date: | 2018-01-10 | | Release date: | 2018-01-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | The SERK3 elongated allele defines a role for BIR ectodomains in brassinosteroid signalling.
Nat Plants, 4, 2018
|
|
6G3W
 
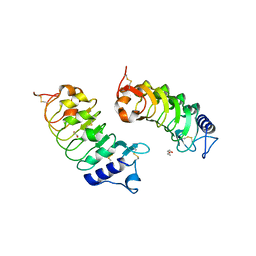 | |
3G3T
 
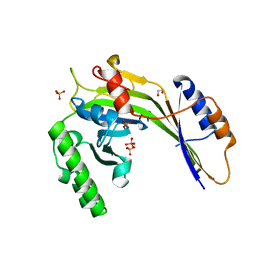 | | Crystal structure of a eukaryotic polyphosphate polymerase in complex with orthophosphate | | Descriptor: | 1,2-ETHANEDIOL, PHOSPHATE ION, Vacuolar transporter chaperone 4 | | Authors: | Lenherr, E.D, Hothorn, M, Scheffzek, K. | | Deposit date: | 2009-02-02 | | Release date: | 2009-05-05 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Catalytic core of a membrane-associated eukaryotic polyphosphate polymerase.
Science, 324, 2009
|
|
1WM0
 
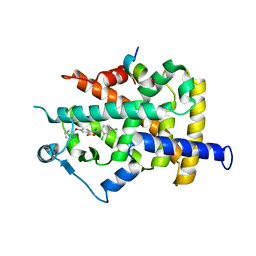 | | PPARgamma in complex with a 2-BABA compound | | Descriptor: | 14-mer from Nuclear receptor coactivator 2, 2-[(2,4-DICHLOROBENZOYL)AMINO]-5-(PYRIMIDIN-2-YLOXY)BENZOIC ACID, Peroxisome proliferator activated receptor gamma | | Authors: | Ostberg, T, Svensson, S, Selen, G, Uppenberg, J, Thor, M, Sundbom, M, Sydow-Backman, M, Gustavsson, A.L, Jendeberg, L. | | Deposit date: | 2004-07-01 | | Release date: | 2004-09-07 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | A new class of peroxisome proliferator-activated receptor agonists with a novel binding epitope shows antidiabetic effects
J.Biol.Chem., 279, 2004
|
|
6R1H
 
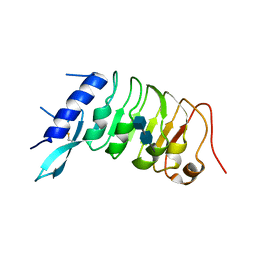 | |
2FXK
 
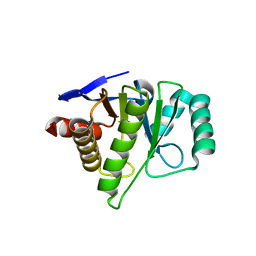 | | Crystal structure of the macro-domain of human core histone variant macroH2A1.1 (form A) | | Descriptor: | H2A histone family, member Y isoform 1 | | Authors: | Kustatscher, G, Hothorn, M, Pugieux, C, Scheffzek, K, Ladurner, A.G. | | Deposit date: | 2006-02-06 | | Release date: | 2006-02-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Splicing regulates NAD metabolite binding to histone macroH2A.
Nat.Struct.Mol.Biol., 12, 2005
|
|
6HX3
 
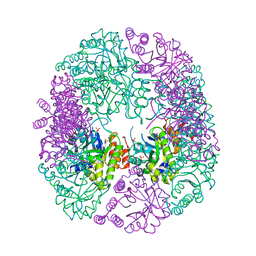 | | PDX1.2/PDX1.3 complex | | Descriptor: | Pyridoxal 5'-phosphate synthase subunit PDX1.3, Pyridoxal 5'-phosphate synthase-like subunit PDX1.2, SULFATE ION | | Authors: | Robinson, G.C, Kaufmann, M, Roux, C, Martinez-Font, J, Hothorn, M, Thore, S, Fitzpatrick, T.B. | | Deposit date: | 2018-10-15 | | Release date: | 2019-04-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of the pseudoenzyme PDX1.2 in complex with its cognate enzyme PDX1.3: a total eclipse.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6HYE
 
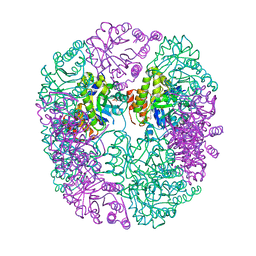 | | PDX1.2/PDX1.3 complex (PDX1.3:K97A) | | Descriptor: | Pyridoxal 5'-phosphate synthase subunit PDX1.3, Pyridoxal 5'-phosphate synthase-like subunit PDX1.2, SULFATE ION | | Authors: | Robinson, G.C, Kaufmann, M, Roux, C, Martinez-Font, J, Hothorn, M, Thore, S, Fitzpatrick, T.B. | | Deposit date: | 2018-10-20 | | Release date: | 2019-04-24 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Crystal structure of the pseudoenzyme PDX1.2 in complex with its cognate enzyme PDX1.3: a total eclipse.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6HXG
 
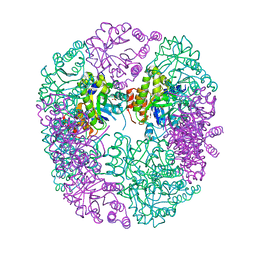 | | PDX1.2/PDX1.3 complex (intermediate) | | Descriptor: | Pyridoxal 5'-phosphate synthase subunit PDX1.3, Pyridoxal 5'-phosphate synthase-like subunit PDX1.2, SULFATE ION | | Authors: | Robinson, G.C, Kaufmann, M, Roux, C, Martinez-Font, J, Hothorn, M, Thore, S, Fitzpatrick, T.B. | | Deposit date: | 2018-10-17 | | Release date: | 2019-04-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the pseudoenzyme PDX1.2 in complex with its cognate enzyme PDX1.3: a total eclipse.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
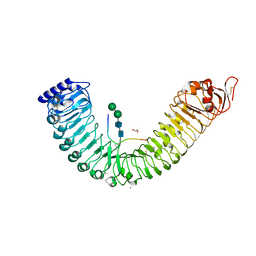
5IXT
 
 | |
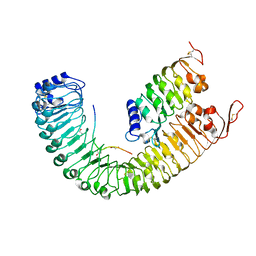
5IYX
 
 | |
5IYV
 
 | |
5IXQ
 
 | |
5IXO
 
 | |
6EU9
 
 | | Crystal structure of Platynereis dumerilii RAR ligand-binding domain in complex with all-trans retinoic acid | | Descriptor: | RETINOIC ACID, Retinoic acid receptor | | Authors: | Handberg-Thorsager, M, Gutierrez-Mazariegos, J, Arold, S.T, Nadendla, E.K, Bertucci, P.Y, Germain, P, Tomancak, P, Pierzchalski, K, Jones, J.W, Albalat, R, Kane, M.A, Bourguet, W, Laudet, V, Arendt, D, Schubert, M. | | Deposit date: | 2017-10-29 | | Release date: | 2018-03-14 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | The ancestral retinoic acid receptor was a low-affinity sensor triggering neuronal differentiation.
Sci Adv, 4, 2018
|
|
5LPY
 
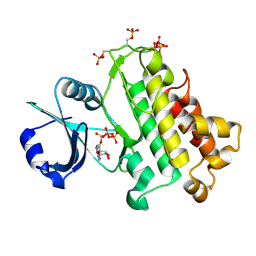 | |
3KIN
 
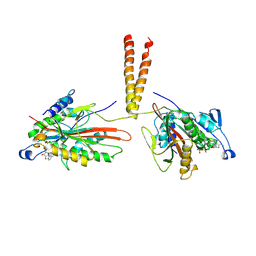 | | KINESIN (DIMERIC) FROM RATTUS NORVEGICUS | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, KINESIN HEAVY CHAIN | | Authors: | Kozielski, F, Sack, S, Marx, A, Thormahlen, M, Schonbrunn, E, Biou, V, Thompson, A, Mandelkow, E.-M, Mandelkow, E. | | Deposit date: | 1997-08-25 | | Release date: | 1998-10-14 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | The crystal structure of dimeric kinesin and implications for microtubule-dependent motility.
Cell(Cambridge,Mass.), 91, 1997
|
|
6XZM
 
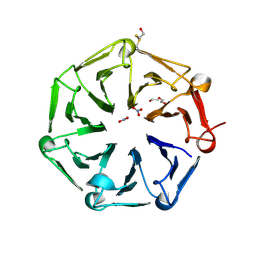 | | Arabidopsis UV-B photoreceptor UVR8 mutant D96N D107N W285A | | Descriptor: | DI(HYDROXYETHYL)ETHER, NITRATE ION, TRIETHYLENE GLYCOL, ... | | Authors: | Lau, K, Hothorn, M. | | Deposit date: | 2020-02-04 | | Release date: | 2021-01-13 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.10009313 Å) | | Cite: | A constitutively monomeric UVR8 photoreceptor confers enhanced UV-B photomorphogenesis.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6XZN
 
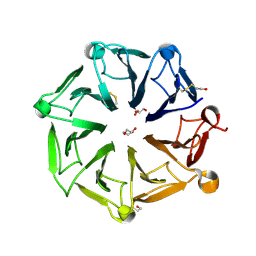 | | Arabidopsis UV-B photoreceptor UVR8 mutant G101S W285A | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, TRIETHYLENE GLYCOL, ... | | Authors: | Lau, K, Hothorn, M. | | Deposit date: | 2020-02-04 | | Release date: | 2021-01-13 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | A constitutively monomeric UVR8 photoreceptor confers enhanced UV-B photomorphogenesis.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6XZL
 
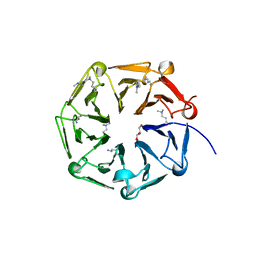 | | Arabidopsis UV-B photoreceptor UVR8 mutant D96N D107N | | Descriptor: | GLYCEROL, Ultraviolet-B receptor UVR8 | | Authors: | Lau, K, Hothorn, M. | | Deposit date: | 2020-02-04 | | Release date: | 2021-01-13 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | A constitutively monomeric UVR8 photoreceptor confers enhanced UV-B photomorphogenesis.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6GRE
 | |
6QV5
 
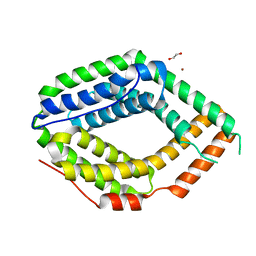 | |
6QV7
 
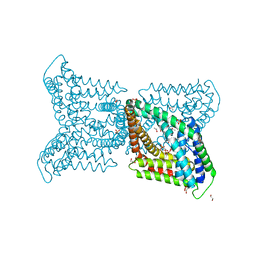 | |
6QVA
 
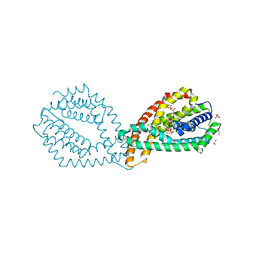 | |
