4H8P
 
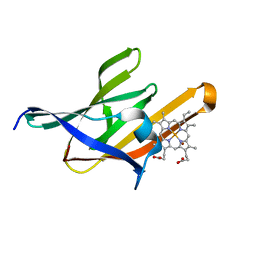 | | NEAT5 domain of IsdX2, a B. anthracis hemophore in complex with heme | | Descriptor: | Iron Transport-associated domain protein, PROTOPORPHYRIN IX CONTAINING FE, ZINC ION | | Authors: | Owens, C.P, Goulding, C.W. | | Deposit date: | 2012-09-23 | | Release date: | 2013-02-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | The Near-iron Transporter (NEAT) Domains of the Anthrax Hemophore IsdX2 Require a Critical Glutamine to Extract Heme from Methemoglobin.
J.Biol.Chem., 288, 2013
|
|
4H8Q
 
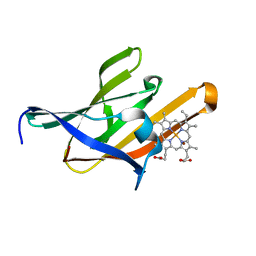 | | Structure of the Q29T IsdX2-NEAT5 mutant in complex with heme | | Descriptor: | Iron Transport-associated domain protein, PROTOPORPHYRIN IX CONTAINING FE, ZINC ION | | Authors: | Owens, C.P, Goulding, C.W. | | Deposit date: | 2012-09-23 | | Release date: | 2013-02-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Near-iron Transporter (NEAT) Domains of the Anthrax Hemophore IsdX2 Require a Critical Glutamine to Extract Heme from Methemoglobin.
J.Biol.Chem., 288, 2013
|
|
4IHU
 
 | |
3QLL
 
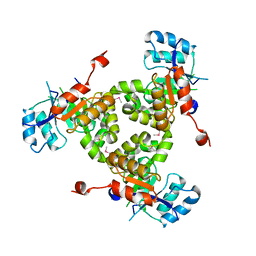 | | Crystal Structure of RipC from Yersinia pestis | | Descriptor: | Citrate lyase | | Authors: | Torres, R, Goulding, C.W. | | Deposit date: | 2011-02-02 | | Release date: | 2012-01-11 | | Last modified: | 2018-05-30 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structural insights into RipC, a putative citrate lyase beta subunit from a Yersinia pestis virulence operon
Acta Crystallogr.,Sect.F, 68, 2012
|
|
3QB9
 
 | |
6PLE
 
 | |
6PWZ
 
 | |
3SIK
 
 | |
3S8D
 | |
3QLK
 
 | | Crystal Structure of RipA from Yersinia pestis | | Descriptor: | Coenzyme A transferase | | Authors: | Torres, R, Goulding, C.W. | | Deposit date: | 2011-02-02 | | Release date: | 2012-01-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Biochemical, structural and molecular dynamics analyses of the potential virulence factor RipA from Yersinia pestis.
Plos One, 6, 2011
|
|
3QLI
 
 | | Crystal Structure of RipA from Yersinia pestis | | Descriptor: | ACETATE ION, Coenzyme A transferase | | Authors: | Torres, R, Goulding, C.W. | | Deposit date: | 2011-02-02 | | Release date: | 2012-01-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Biochemical, structural and molecular dynamics analyses of the potential virulence factor RipA from Yersinia pestis.
Plos One, 6, 2011
|
|
8EY3
 
 | | Contact-dependent growth inhibition (CDI) immunity protein from E. coli O32:H37 | | Descriptor: | Cys_rich_CPCC domain-containing protein, FE (III) ION, SODIUM ION | | Authors: | Michalska, K, Stols, L, Eschenfeldt, W, Goulding, C.W, Hayes, C.S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2022-10-26 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | Contact-dependent growth inhibition (CDI) immunity protein from E. coli O32:H37
To Be Published
|
|
8EY4
 
 | | Contact-dependent growth inhibition toxin-immunity protein complex from E. coli O32:H37 | | Descriptor: | Cys_rich_CPCC domain-containing protein, FE (III) ION, PT-VENN domain-containing protein | | Authors: | Michalska, K, Stols, L, Eschenfeldt, W, Goulding, C.W, Hayes, C.S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2022-10-26 | | Release date: | 2023-11-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Contact-dependent growth inhibition toxin-immunity protein complex from E. coli O32:H37
To Be Published
|
|
4Q7O
 
 | | The crystal structure of an immunity protein NMB0503 from Neisseria meningitidis MC58 | | Descriptor: | BROMIDE ION, FORMIC ACID, Immunity protein | | Authors: | Tan, K, Stols, L, Eschenfeldt, W, Babnigg, G, Low, D.A, Hayes, C.S, Goulding, C.W, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Structure-Function Analysis of Polymorphic CDI Toxin-Immunity Protein Complexes (UC4CDI) | | Deposit date: | 2014-04-25 | | Release date: | 2014-05-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | The structure of a contact-dependent growth-inhibition (CDI) immunity protein from Neisseria meningitidis MC58.
Acta Crystallogr F Struct Biol Commun, 71, 2015
|
|
5HKQ
 
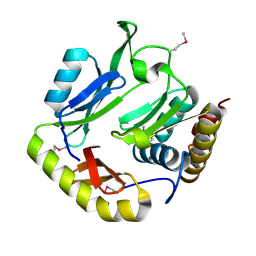 | | Crystal structure of CDI complex from Escherichia coli STEC_O31 | | Descriptor: | CdiI immunity protein, Contact-dependent inhibitor A | | Authors: | Michalska, K, Stols, L, Eschenfeldt, W, Goulding, C.W, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Structure-Function Analysis of Polymorphic CDI Toxin-Immunity Protein Complexes (UC4CDI) | | Deposit date: | 2016-01-14 | | Release date: | 2017-01-18 | | Last modified: | 2020-03-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Functional plasticity of antibacterial EndoU toxins.
Mol.Microbiol., 109, 2018
|
|
5FFP
 
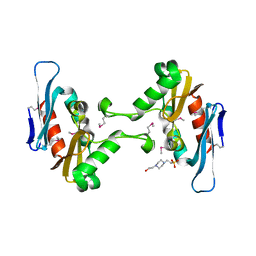 | | Crystal structure of CdiI from Burkholderia dolosa AUO158 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Immunity 23 family protein | | Authors: | Michalska, K, Stols, L, Eschenfeldt, W, Goulding, C.W, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Structure-Function Analysis of Polymorphic CDI Toxin-Immunity Protein Complexes (UC4CDI) | | Deposit date: | 2015-12-18 | | Release date: | 2016-01-20 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of CdiI from Burkholderia dolosa AUO158
To Be Published
|
|
4NTQ
 
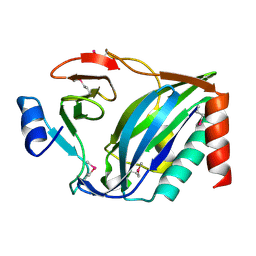 | |
5I4R
 
 | | Contact-dependent inhibition system from Escherichia coli NC101 - ternary CdiA/CdiI/EF-Tu complex (trypsin-modified) | | Descriptor: | Contact-dependent inhibitor A, Contact-dependent inhibitor I, Elongation factor Tu, ... | | Authors: | Michalska, K, Stols, L, Eschenfeldt, W, Hayes, C.S, Goulding, C.W, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Structure-Function Analysis of Polymorphic CDI Toxin-Immunity Protein Complexes (UC4CDI) | | Deposit date: | 2016-02-12 | | Release date: | 2017-06-28 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of a novel antibacterial toxin that exploits elongation factor Tu to cleave specific transfer RNAs.
Nucleic Acids Res., 45, 2017
|
|
5I4Q
 
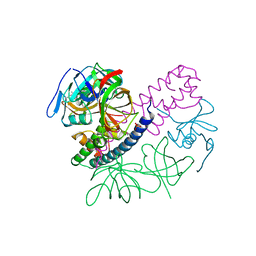 | | Contact-dependent inhibition system from Escherichia coli NC101 - ternary CdiA/CdiI/EF-Tu complex (domains 2 and 3) | | Descriptor: | CHLORIDE ION, Contact-dependent inhibitor A, Contact-dependent inhibitor I, ... | | Authors: | Michalska, K, Stols, L, Eschenfeldt, W, Hayes, C.S, Goulding, C.W, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Structure-Function Analysis of Polymorphic CDI Toxin-Immunity Protein Complexes (UC4CDI) | | Deposit date: | 2016-02-12 | | Release date: | 2017-06-28 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structure of a novel antibacterial toxin that exploits elongation factor Tu to cleave specific transfer RNAs.
Nucleic Acids Res., 45, 2017
|
|
4NL5
 
 | | Mycobacterium tuberculosis heme-degrading protein MhuD in complex with heme and cyanide | | Descriptor: | ACETATE ION, CYANIDE ION, Heme-degrading monooxygenase HmoB, ... | | Authors: | Morse, R.P, Chao, A, Goulding, C.W. | | Deposit date: | 2013-11-13 | | Release date: | 2014-06-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystallographic and Spectroscopic Insights into Heme Degradation by Mycobacterium tuberculosis MhuD.
Inorg.Chem., 53, 2014
|
|
4IVH
 
 | | Crystal structure of QKLVFFAED nonapeptide segment from amyloid beta incorporated into a macrocyclic beta-sheet template | | Descriptor: | SODIUM ION, TERTIARY-BUTYL ALCOHOL, cyclo[Gln-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-(delta-linked-Orn)-Hao-Lys-Hao-(p-bromoPhe)-Thr-(delta-linked-Orn)] | | Authors: | Pham, J.D, Chim, N, Goulding, C.W, Nowick, J.S. | | Deposit date: | 2013-01-22 | | Release date: | 2013-07-31 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structures of beta-Amyloid Peptide Oligomers
J.Am.Chem.Soc., 2013
|
|
4LMP
 
 | | Mycobacterium tuberculosis L-alanine dehydrogenase x-ray structure in complex with N6-methyl adenosine | | Descriptor: | Alanine dehydrogenase, GLYCEROL, N-methyladenosine, ... | | Authors: | Kim, H.-B, Hung, L.-W, Goulding, C.W, Terwilliger, T.C, Kim, C.-Y, Structures of Mtb Proteins Conferring Susceptibility to Known Mtb Inhibitors (MTBI) | | Deposit date: | 2013-07-10 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Drug target analysis by dye-ligand affinity chromatography
To be Published
|
|
4LPN
 
 | | Frog M-ferritin with cobalt, D127E mutant | | Descriptor: | CHLORIDE ION, COBALT (II) ION, Ferritin, ... | | Authors: | Torres, R, Behera, R, Goulding, C.W. | | Deposit date: | 2013-07-16 | | Release date: | 2014-07-16 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | D127E ion channel exit modification in ferritin nanocages entraps Fe(II) and impairs its distribution to diiron catalytic centers
To be Published
|
|
4LPM
 
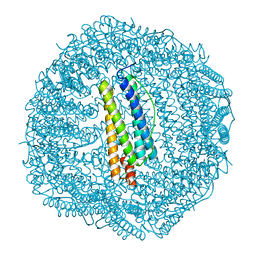 | | Frog M-ferritin with magnesium, D127E mutant | | Descriptor: | CHLORIDE ION, Ferritin, middle subunit, ... | | Authors: | Torres, R, Behera, R, Goulding, C.W, Theil, E.C. | | Deposit date: | 2013-07-16 | | Release date: | 2014-07-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | D127E ion channel exit modification in ferritin nanocages entraps Fe(II) and impairs its distribution to diiron catalytic centers
To be Published
|
|
4N8I
 
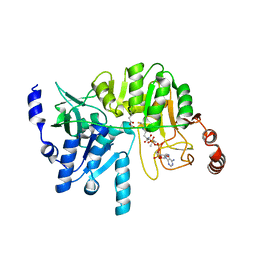 | | M31G mutant, RipA structure | | Descriptor: | 4-hydroxybutyrate coenzyme A transferase, ACETATE ION, COENZYME A | | Authors: | Torres, R, Goulding, C.W. | | Deposit date: | 2013-10-17 | | Release date: | 2014-04-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.011 Å) | | Cite: | Structural snapshots along the reaction pathway of Yersinia pestis RipA, a putative butyryl-CoA transferase.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
