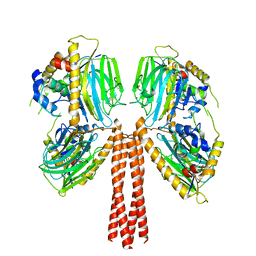
8JP4
 
 | |
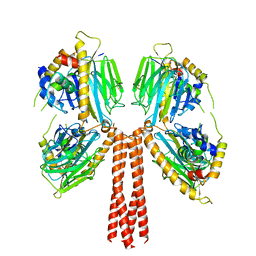
8JP5
 
 | |
5B6X
 
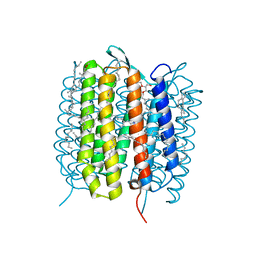 | | A three dimensional movie of structural changes in bacteriorhodopsin: structure obtained 760 ns after photoexcitation | | Descriptor: | 2,3-DI-PHYTANYL-GLYCEROL, Bacteriorhodopsin, DECANE, ... | | Authors: | Royant, A, Nango, E, Nakane, T, Tanaka, T, Arima, T, Neutze, R, Iwata, S. | | Deposit date: | 2016-06-02 | | Release date: | 2016-12-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A three-dimensional movie of structural changes in bacteriorhodopsin
Science, 354, 2016
|
|
7VGU
 
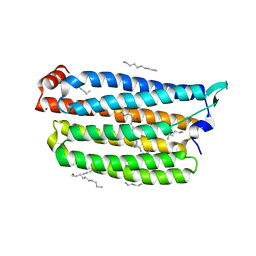 | | Time-resolved serial femtosecond crystallography structure of light-driven chloride ion-pumping rhodopsin, NM-R3 : structure obtained 1 msec after photoexcitation with bromide ion | | Descriptor: | BROMIDE ION, Chloride pumping rhodopsin, DECANE, ... | | Authors: | Hosaka, T, Nango, E, Nakane, T, Luo, F, Kimura-Someya, T, Shirouzu, M. | | Deposit date: | 2021-09-18 | | Release date: | 2022-02-16 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Conformational alterations in unidirectional ion transport of a light-driven chloride pump revealed using X-ray free electron lasers.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7VGT
 
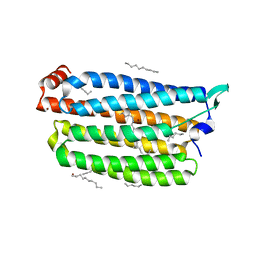 | | Time-resolved serial femtosecond crystallography structure of light-driven chloride ion-pumping rhodopsin, NM-R3: resting state structure with bromide ion | | Descriptor: | BROMIDE ION, Chloride pumping rhodopsin, DECANE, ... | | Authors: | Hosaka, T, Nango, E, Nakane, T, Luo, F, Kimura-Someya, T, Shirouzu, M. | | Deposit date: | 2021-09-18 | | Release date: | 2022-02-16 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Conformational alterations in unidirectional ion transport of a light-driven chloride pump revealed using X-ray free electron lasers.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7VGV
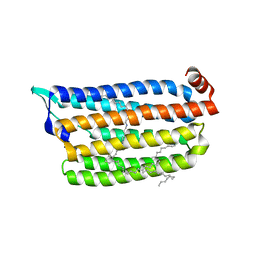 
 | | Anion free form of light-driven chloride ion-pumping rhodopsin, NM-R3, structure determined by serial femtosecond crystallography at SACLA | | Descriptor: | CHLORIDE ION, Chloride pumping rhodopsin, HEXADECANE, ... | | Authors: | Hosaka, T, Nango, E, Nakane, T, Luo, F, Kimura-Someya, T, Shirouzu, M. | | Deposit date: | 2021-09-18 | | Release date: | 2022-02-16 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Conformational alterations in unidirectional ion transport of a light-driven chloride pump revealed using X-ray free electron lasers.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
8JA3
 
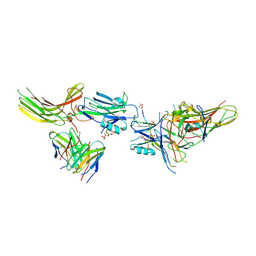 | | Structure of beta-arrestin1 in complex with C3aRpp | | Descriptor: | Beta-arrestin-1, C3a anaphylatoxin chemotactic receptor, Fab30 heavy chain, ... | | Authors: | Maharana, J, Sarma, P, Yadav, M.K, Chami, M, Banerjee, R, Shukla, A.K. | | Deposit date: | 2023-05-05 | | Release date: | 2023-12-27 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (3.94 Å) | | Cite: | Molecular insights into atypical modes of beta-arrestin interaction with seven transmembrane receptors.
Science, 383, 2024
|
|
5B6Y
 
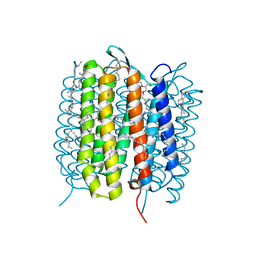 | | A three dimensional movie of structural changes in bacteriorhodopsin: structure obtained 36.2 us after photoexcitation | | Descriptor: | 2,3-DI-PHYTANYL-GLYCEROL, Bacteriorhodopsin, DECANE, ... | | Authors: | Royant, A, Nango, E, Nakane, T, Tanaka, T, Arima, T, Neutze, R, Iwata, S. | | Deposit date: | 2016-06-02 | | Release date: | 2016-12-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A three-dimensional movie of structural changes in bacteriorhodopsin
Science, 354, 2016
|
|
5B35
 
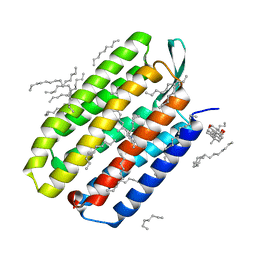 | | Serial Femtosecond Crystallography (SFX) of Ground State Bacteriorhodopsin Crystallized from Bicelles Determined Using 7-keV X-ray Free Electron Laser (XFEL) at SACLA | | Descriptor: | (3R,5S,7R,8R,9S,10S,12S,13R,14S,17R)-10,13-dimethyl-17-[(2R)-pentan-2-yl]-2,3,4,5,6,7,8,9,11,12,14,15,16,17-tetradecahydro-1H-cyclopenta[a]phenanthrene-3,7,12-triol, Bacteriorhodopsin, DECANE, ... | | Authors: | Mizohata, E, Nakane, T, Suzuki, M. | | Deposit date: | 2016-02-10 | | Release date: | 2016-11-09 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Membrane protein structure determination by SAD, SIR, or SIRAS phasing in serial femtosecond crystallography using an iododetergent
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
8YVG
 
 | | canine immunoproteasome 20S subunit in complex with compound 1 | | Descriptor: | Proteasome subunit alpha type, Proteasome subunit beta, Proteasome subunit beta type-8, ... | | Authors: | Kashima, A, Arai, Y. | | Deposit date: | 2024-03-28 | | Release date: | 2024-07-31 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (2 Å) | | Cite: | Optimization of alpha-amido boronic acids via cryo-electron microscopy analysis: Discovery of a novel highly selective immunoproteasome subunit LMP7 ( beta 5i)/LMP2 ( beta 1i) dual inhibitor.
Bioorg.Med.Chem., 109, 2024
|
|
8YSX
 
 | | canine immunoproteasome 20S subunit in complex with compound 2 | | Descriptor: | Proteasome subunit alpha type, Proteasome subunit beta, Proteasome subunit beta type-8, ... | | Authors: | Kashima, A, Arai, Y. | | Deposit date: | 2024-03-24 | | Release date: | 2024-07-31 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (2.2 Å) | | Cite: | Optimization of alpha-amido boronic acids via cryo-electron microscopy analysis: Discovery of a novel highly selective immunoproteasome subunit LMP7 ( beta 5i)/LMP2 ( beta 1i) dual inhibitor.
Bioorg.Med.Chem., 109, 2024
|
|
8YVP
 
 | | canine immunoproteasome 20S subunit in complex with compound 1 | | Descriptor: | Proteasome subunit alpha type-1, Proteasome subunit alpha type-2, Proteasome subunit alpha type-3, ... | | Authors: | Kashima, A, Arai, Y. | | Deposit date: | 2024-03-29 | | Release date: | 2024-07-31 | | Last modified: | 2024-10-30 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Optimization of alpha-amido boronic acids via cryo-electron microscopy analysis: Discovery of a novel highly selective immunoproteasome subunit LMP7 ( beta 5i)/LMP2 ( beta 1i) dual inhibitor.
Bioorg.Med.Chem., 109, 2024
|
|
8YPK
 
 | | mouse proteasome 20S subunit in complex with compound 1 | | Descriptor: | Proteasome subunit alpha type-1, Proteasome subunit alpha type-2, Proteasome subunit alpha type-3, ... | | Authors: | Kashima, A, Arai, Y. | | Deposit date: | 2024-03-17 | | Release date: | 2024-07-31 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Optimization of alpha-amido boronic acids via cryo-electron microscopy analysis: Discovery of a novel highly selective immunoproteasome subunit LMP7 ( beta 5i)/LMP2 ( beta 1i) dual inhibitor.
Bioorg.Med.Chem., 109, 2024
|
|
7W9W
 
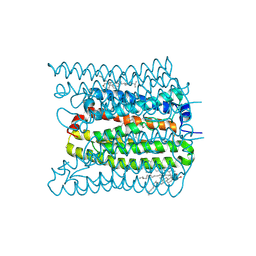 | | 2.02 angstrom cryo-EM structure of the pump-like channelrhodopsin ChRmine | | Descriptor: | CHOLESTEROL, ChRmine, PALMITIC ACID, ... | | Authors: | Kishi, K.E, Kim, Y, Fukuda, M, Yamashita, K, Deisseroth, K, Kato, H.E. | | Deposit date: | 2021-12-11 | | Release date: | 2022-02-02 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (2.02 Å) | | Cite: | Structural basis for channel conduction in the pump-like channelrhodopsin ChRmine.
Cell, 185, 2022
|
|
8KDN
 
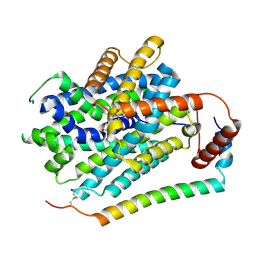 | | Structure of LAT1-CD98hc in complex with L-Phe, focused on TMD | | Descriptor: | 4F2 cell-surface antigen heavy chain, Large neutral amino acids transporter small subunit 1, PHENYLALANINE | | Authors: | Lee, Y. | | Deposit date: | 2023-08-09 | | Release date: | 2025-02-12 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (4.12 Å) | | Cite: | Structural basis of anticancer drug recognition and amino acid transport by LAT1.
Nat Commun, 16, 2025
|
|
8KDJ
 
 | | Structure of apo inward-open LAT1-CD98h in nanodisc, focused on TMD | | Descriptor: | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine, 4F2 cell-surface antigen heavy chain, CHOLESTEROL, ... | | Authors: | Lee, Y. | | Deposit date: | 2023-08-09 | | Release date: | 2025-02-12 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.73 Å) | | Cite: | Structural basis of anticancer drug recognition and amino acid transport by LAT1.
Nat Commun, 16, 2025
|
|
8KDH
 
 | | Structure of LAT1-CD98hc in complex with BCH, focused on TMD | | Descriptor: | (1~{S},2~{R},4~{R})-2-azanylbicyclo[2.2.1]heptane-2-carboxylic acid, 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine, 4F2 cell-surface antigen heavy chain, ... | | Authors: | Lee, Y. | | Deposit date: | 2023-08-09 | | Release date: | 2025-02-12 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.78 Å) | | Cite: | Structural basis of anticancer drug recognition and amino acid transport by LAT1.
Nat Commun, 16, 2025
|
|
8KDD
 
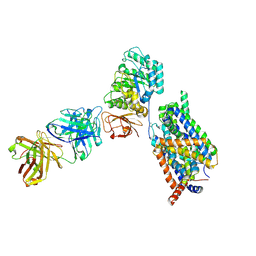 | | Structure of LAT1-CD98hc-Fab170 in complex with JPH203, consensus map | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4F2 cell-surface antigen heavy chain, Fab170 heavy chain, ... | | Authors: | Lee, Y. | | Deposit date: | 2023-08-09 | | Release date: | 2025-02-12 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.83 Å) | | Cite: | Structural basis of anticancer drug recognition and amino acid transport by LAT1.
Nat Commun, 16, 2025
|
|
8KDP
 
 | | Structure of apo outward-open LAT1-CD98h in nanodisc, focused on TMD | | Descriptor: | 4F2 cell-surface antigen heavy chain, Large neutral amino acids transporter small subunit 1 | | Authors: | Lee, Y. | | Deposit date: | 2023-08-09 | | Release date: | 2025-02-12 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (4.12 Å) | | Cite: | Structural basis of anticancer drug recognition and amino acid transport by LAT1.
Nat Commun, 16, 2025
|
|
8KDI
 
 | | Structure of apo inward-open LAT1-CD98hc-Fab170 in nanodisc, consensus map | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4F2 cell-surface antigen heavy chain, Fab170 heavy chain, ... | | Authors: | Lee, Y. | | Deposit date: | 2023-08-09 | | Release date: | 2025-02-12 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.58 Å) | | Cite: | Structural basis of anticancer drug recognition and amino acid transport by LAT1.
Nat Commun, 16, 2025
|
|
8KDF
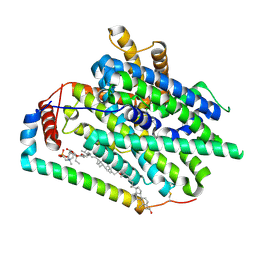 
 | | Structure of LAT1-CD98hc in complex with JPH203, focused on TMD | | Descriptor: | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine, 4F2 cell-surface antigen heavy chain, CHOLESTEROL, ... | | Authors: | Lee, Y. | | Deposit date: | 2023-08-09 | | Release date: | 2025-02-12 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.89 Å) | | Cite: | Structural basis of anticancer drug recognition and amino acid transport by LAT1.
Nat Commun, 16, 2025
|
|
8KDG
 
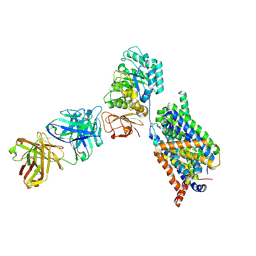 | | Structure of LAT1-CD98hc-Fab170 in complex with BCH, consensus map | | Descriptor: | (1~{S},2~{R},4~{R})-2-azanylbicyclo[2.2.1]heptane-2-carboxylic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose, 4F2 cell-surface antigen heavy chain, ... | | Authors: | Lee, Y. | | Deposit date: | 2023-08-09 | | Release date: | 2025-02-12 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.68 Å) | | Cite: | Structural basis of anticancer drug recognition and amino acid transport by LAT1.
Nat Commun, 16, 2025
|
|
8KDO
 
 | | Structure of LAT1-CD98hc in complex with melphalan, focused on TMD | | Descriptor: | (2~{S})-2-azanyl-3-[4-[bis(2-chloroethyl)amino]phenyl]propanoic acid, 4F2 cell-surface antigen heavy chain, Large neutral amino acids transporter small subunit 1 | | Authors: | Lee, Y. | | Deposit date: | 2023-08-09 | | Release date: | 2025-02-12 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (4.12 Å) | | Cite: | Structural basis of anticancer drug recognition and amino acid transport by LAT1.
Nat Commun, 16, 2025
|
|
5B6W
 
 | | A three dimensional movie of structural changes in bacteriorhodopsin: structure obtained 16 ns after photoexcitation | | Descriptor: | 2,3-DI-PHYTANYL-GLYCEROL, Bacteriorhodopsin, DECANE, ... | | Authors: | Royant, A, Nango, E, Nakane, T, Tanaka, T, Arima, T, Neutze, R, Iwata, S. | | Deposit date: | 2016-06-02 | | Release date: | 2016-12-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A three-dimensional movie of structural changes in bacteriorhodopsin
Science, 354, 2016
|
|
5B6Z
 
 | | A three dimensional movie of structural changes in bacteriorhodopsin: structure obtained 1.725 ms us after photoexcitation | | Descriptor: | 2,3-DI-PHYTANYL-GLYCEROL, Bacteriorhodopsin, DECANE, ... | | Authors: | Royant, A, Nango, E, Nakane, T, Tanaka, T, Arima, T, Neutze, R, Iwata, S. | | Deposit date: | 2016-06-02 | | Release date: | 2016-12-21 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A three-dimensional movie of structural changes in bacteriorhodopsin
Science, 354, 2016
|
|
