8CZL
 
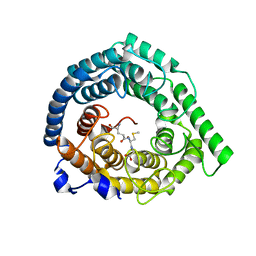 | | Human LanCL1 bound to methyl glutathione (MeGSH) | | Descriptor: | Glutathione S-transferase LANCL1, L-GAMMA-GLUTAMYL-S-METHYLCYSTEINYLGLYCINE, ZINC ION | | Authors: | Ongpipattanakul, C, Nair, S.K. | | Deposit date: | 2022-05-24 | | Release date: | 2023-01-25 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | The mechanism of thia-Michael addition catalyzed by LanC enzymes.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
8D19
 
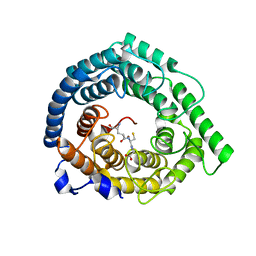 | | Human LanCL1 bound to GSH | | Descriptor: | GLUTATHIONE, Glutathione S-transferase LANCL1, ZINC ION | | Authors: | Ongpipattanakul, C, Nair, S.K. | | Deposit date: | 2022-05-26 | | Release date: | 2023-01-25 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | The mechanism of thia-Michael addition catalyzed by LanC enzymes.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
8D0V
 
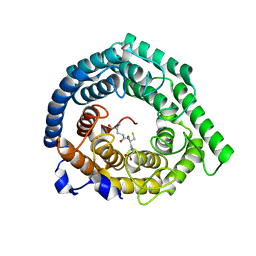 | | Human LanCL1 C264A mutant bound to GSH | | Descriptor: | GLUTATHIONE, Glutathione S-transferase LANCL1, ZINC ION | | Authors: | Ongpipattanakul, C, Nair, S.K. | | Deposit date: | 2022-05-26 | | Release date: | 2023-01-25 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | The mechanism of thia-Michael addition catalyzed by LanC enzymes.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
7RTY
 
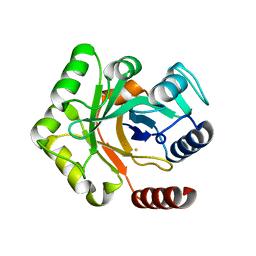 | |
8CWX
 
 | |
7MSK
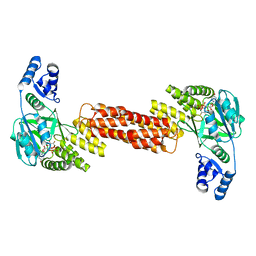 
 | | ThuS glycosin S-glycosyltransferase | | Descriptor: | Glyco_trans_2-like domain-containing protein, MAGNESIUM ION, URIDINE-5'-DIPHOSPHATE-2-DEOXY-2-FLUORO-ALPHA-D-GLUCOSE | | Authors: | Garg, N, Nair, S.K. | | Deposit date: | 2021-05-11 | | Release date: | 2022-04-13 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Structural and mechanistic investigations of protein S-glycosyltransferases.
Cell Chem Biol, 28, 2021
|
|
7MSN
 
 | | SunS glycosin S-glycosyltransferase | | Descriptor: | SPbeta prophage-derived glycosyltransferase SunS, URIDINE-5'-DIPHOSPHATE-GLUCOSE | | Authors: | Garg, N, Nair, S.K. | | Deposit date: | 2021-05-11 | | Release date: | 2022-04-13 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural and mechanistic investigations of protein S-glycosyltransferases.
Cell Chem Biol, 28, 2021
|
|
7MSP
 
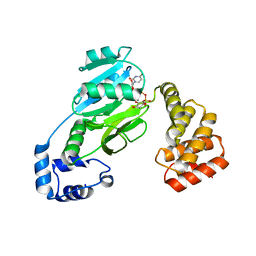 | | SunS glycosin S-glycosyltransferase | | Descriptor: | MAGNESIUM ION, SPbeta prophage-derived glycosyltransferase SunS, URIDINE-5'-DIPHOSPHATE-GLUCOSE | | Authors: | Garg, N, Nair, S.K. | | Deposit date: | 2021-05-11 | | Release date: | 2022-04-13 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural and mechanistic investigations of protein S-glycosyltransferases.
Cell Chem Biol, 28, 2021
|
|
6UAK
 
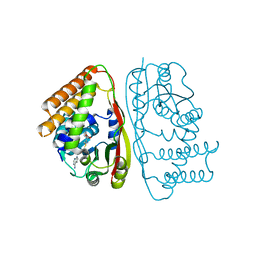 | | LahSb - C-terminal methyltransferase involved in RiPP biosynthesis | | Descriptor: | S-ADENOSYL-L-HOMOCYSTEINE, SAM dependent methyltransferase LahSB | | Authors: | Nair, S.K, Estrada, P. | | Deposit date: | 2019-09-10 | | Release date: | 2019-12-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Characterization of a Dehydratase and Methyltransferase in the Biosynthesis of Ribosomally Synthesized and Post-translationally Modified Peptides in Lachnospiraceae.
Chembiochem, 21, 2020
|
|
4QED
 
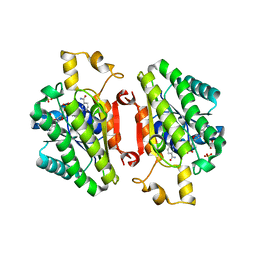 | | ElxO Y152F with NADPH Bound | | Descriptor: | ElxO, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, SULFATE ION | | Authors: | Garg, N, Nair, S.K. | | Deposit date: | 2014-05-15 | | Release date: | 2014-07-02 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Substrate Specificity of the Lanthipeptide Peptidase ElxP and the Oxidoreductase ElxO.
Acs Chem.Biol., 9, 2014
|
|
6B9T
 
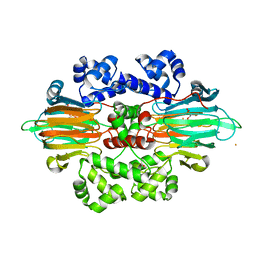 | |
6B9S
 
 | |
6B9R
 
 | |
6C0Y
 
 | |
6MPZ
 
 | | Crystal structure of a double glycine motif protease from AMS/PCAT transporter in complex with the leader peptide | | Descriptor: | 3,6,9,12,15,18-HEXAOXAICOSANE, Double Glycine Motif Protease domain from AMS/PCAT Transporter, peptide aldehyde inhibitor 1 based on the ProcA2.8 leader peptide | | Authors: | Dong, S.-H, Nair, S.K. | | Deposit date: | 2018-10-09 | | Release date: | 2019-02-06 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Insights into AMS/PCAT transporters from biochemical and structural characterization of a double Glycine motif protease.
Elife, 8, 2019
|
|
6M7Y
 
 | |
6C0G
 
 | |
6C0H
 
 | |
6EC8
 
 | |
6EC7
 
 | |
3OU7
 
 | | DhpI-SAM-HEP complex | | Descriptor: | (2-hydroxyethyl)phosphonic acid, S-ADENOSYLMETHIONINE, SAM-dependent methyltransferase, ... | | Authors: | Bae, B, Nair, S.K. | | Deposit date: | 2010-09-14 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Characterization and structure of DhpI, a phosphonate O-methyltransferase involved in dehydrophos biosynthesis.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3OU6
 
 | | DhpI-SAM complex | | Descriptor: | S-ADENOSYLMETHIONINE, SAM-dependent methyltransferase, SULFATE ION | | Authors: | Bae, B, Nair, S.K. | | Deposit date: | 2010-09-14 | | Release date: | 2010-10-27 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Characterization and structure of DhpI, a phosphonate O-methyltransferase involved in dehydrophos biosynthesis.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3OU2
 
 | | DhpI-SAH complex structure | | Descriptor: | S-ADENOSYL-L-HOMOCYSTEINE, SAM-dependent methyltransferase | | Authors: | Bae, B, Nair, S.K. | | Deposit date: | 2010-09-14 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Characterization and structure of DhpI, a phosphonate O-methyltransferase involved in dehydrophos biosynthesis.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3RZZ
 
 | |
4E5P
 
 | |
