3KRN
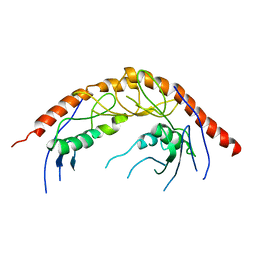 
 | | Crystal Structure of C. elegans cell-death-related nuclease 5(CRN-5) | | Descriptor: | Protein C14A4.5, confirmed by transcript evidence | | Authors: | Yang, C.-C, Wang, Y.-T, Hsiao, Y.-Y, Doudeva, L.G, Chow, S.Y, Yuan, H.S. | | Deposit date: | 2009-11-19 | | Release date: | 2010-01-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.918 Å) | | Cite: | Structural and biochemical characterization of CRN-5 and Rrp46: an exosome component participating in apoptotic DNA degradation
Rna, 16, 2010
|
|
4Y1H
 
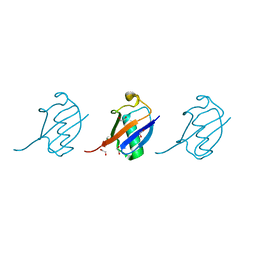 | | Crystal structure of K33 linked tri-Ubiquitin | | Descriptor: | 1,2-ETHANEDIOL, Ubiquitin-40S ribosomal protein S27a | | Authors: | Kristariyanto, Y.A, Abdul Rehman, S.A, Choi, S.Y, Ritorto, S, Campbell, D.G, Morrice, N.A, Toth, R, Kulathu, Y. | | Deposit date: | 2015-02-07 | | Release date: | 2015-03-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Assembly and structure of Lys33-linked polyubiquitin reveals distinct conformations.
Biochem.J., 467, 2015
|
|
4XYZ
 
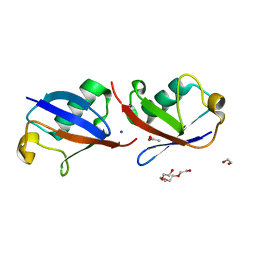 | | Crystal structure of K33 linked di-Ubiquitin | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, IODIDE ION, ... | | Authors: | Kristariyanto, Y.A, Abdul Rehman, S.A, Choi, S.Y, Ritorto, S, Campbell, D.G, Morrice, N.A, Toth, R, Kulathu, Y. | | Deposit date: | 2015-02-03 | | Release date: | 2015-03-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Assembly and structure of Lys33-linked polyubiquitin reveals distinct conformations.
Biochem.J., 467, 2015
|
|
5XJH
 
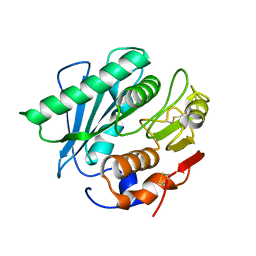 | |
7BYK
 
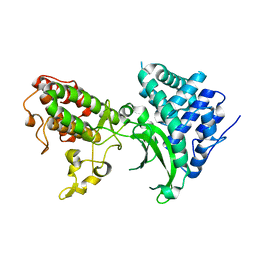 | |
5F20
 
 | |
5F1Z
 
 | |
6IJ6
 
 | |
6IJ4
 
 | |
6IJ5
 
 | |
6IJ3
 
 | |
5YNS
 
 | |
5JKN
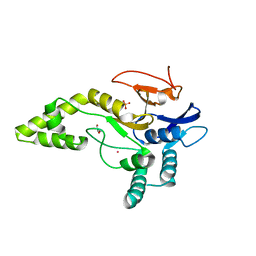 
 | | Crystal structure of deubiquitinase MINDY-1 | | Descriptor: | DI(HYDROXYETHYL)ETHER, MERCURY (II) ION, PHOSPHATE ION, ... | | Authors: | Abdul Rehman, S.A, Kulathu, Y. | | Deposit date: | 2016-04-26 | | Release date: | 2016-06-22 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | MINDY-1 Is a Member of an Evolutionarily Conserved and Structurally Distinct New Family of Deubiquitinating Enzymes.
Mol.Cell, 63, 2016
|
|
8SX3
 
 | | 10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10 | | Descriptor: | 10E8 Fab heavy chain, 10E8 light chain, 10E8-GT10.2 immunogen, ... | | Authors: | Huang, J, Ozorowski, G, Ward, A.B. | | Deposit date: | 2023-05-19 | | Release date: | 2024-05-22 | | Last modified: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Vaccination induces broadly neutralizing antibody precursors to HIV gp41.
Nat.Immunol., 25, 2024
|
|
8U03
 
 | |
8TZN
 
 | |
8TZW
 
 | |
8U08
 
 | |
4IKC
 
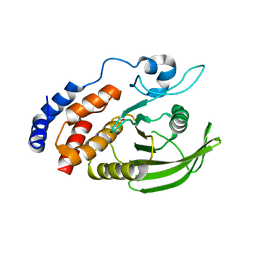 | | Crystal Structure of catalytic domain of PTPRQ | | Descriptor: | CHLORIDE ION, Phosphotidylinositol phosphatase PTPRQ, SULFATE ION | | Authors: | Yu, K.R, Ryu, S.E, Kim, S.J. | | Deposit date: | 2012-12-26 | | Release date: | 2013-07-31 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Structural basis for the dephosphorylating activity of PTPRQ towards phosphatidylinositide substrates
Acta Crystallogr.,Sect.D, 69, 2013
|
|
7JU2
 
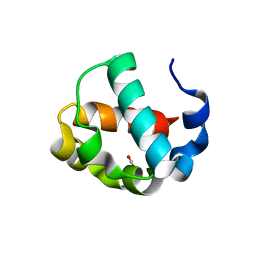 | | Crystal structure of the monomeric ETV6 PNT domain | | Descriptor: | FORMIC ACID, Transcription factor ETV6 | | Authors: | Gerak, C.A.N, Kolesnikov, M, Murphy, M.E.P, McIntosh, L.P. | | Deposit date: | 2020-08-19 | | Release date: | 2021-01-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.85002184 Å) | | Cite: | Biophysical characterization of the ETV6 PNT domain polymerization interfaces.
J.Biol.Chem., 296, 2021
|
|
5HZ2
 
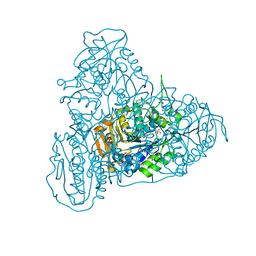 | | Crystal structure of PhaC1 from Ralstonia eutropha | | Descriptor: | GLYCEROL, Poly-beta-hydroxybutyrate polymerase, SULFATE ION | | Authors: | Kim, J, Kim, K.-J. | | Deposit date: | 2016-02-02 | | Release date: | 2016-12-07 | | Last modified: | 2017-04-05 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Ralstonia eutropha polyhydroxyalkanoate synthase C-terminal domain and reaction mechanisms.
Biotechnol J, 12, 2017
|
|
9EZL
 
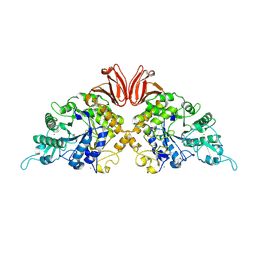 | |
5JQS
 
 | |
3HKM
 
 | | Crystal Structure of rice(Oryza sativa) Rrp46 | | Descriptor: | Os03g0854200 protein | | Authors: | Yang, C.-C, Wang, Y.-T, Hsiao, Y.-Y, Doudeva, L.G, Yuan, H.S. | | Deposit date: | 2009-05-25 | | Release date: | 2010-01-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9845 Å) | | Cite: | Structural and biochemical characterization of CRN-5 and Rrp46: an exosome component participating in apoptotic DNA degradation
Rna, 16, 2010
|
|
7NFW
 
 | |
