1Y1N
 
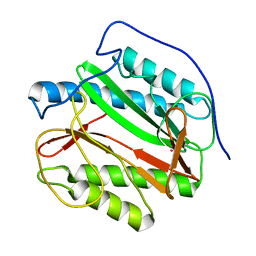 | | Identification of SH3 motif in M. Tuberculosis methionine aminopeptidase suggests a mode of interaction with the ribosome | | Descriptor: | Methionine aminopeptidase 1B, POTASSIUM ION | | Authors: | Addlagatta, A, Quillin, M.L, Omotoso, O, Liu, J.O, Matthews, B.W. | | Deposit date: | 2004-11-18 | | Release date: | 2005-05-24 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.51 Å) | | Cite: | Identification of an SH3-Binding Motif in a New Class of Methionine Aminopeptidases from Mycobacterium tuberculosis Suggests a Mode of Interaction with the Ribosome
Biochemistry, 44, 2005
|
|
1Z9G
 
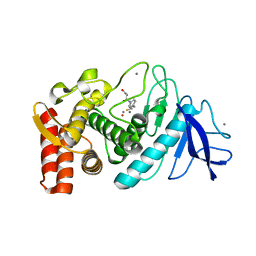 | | Crystal Structure Analysis of Thermolysin Complexed with the Inhibitor (R)-retro-thiorphan | | Descriptor: | (R)-RETRO-THIORPHAN, CALCIUM ION, Thermolysin, ... | | Authors: | Roderick, S.L, Fournie-Zaluski, M.C, Roques, B.P, Matthews, B.W. | | Deposit date: | 2005-04-01 | | Release date: | 2005-04-19 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Thiorphan and retro-thiorphan display equivalent interactions when bound to crystalline thermolysin
Biochemistry, 28, 1989
|
|
1OWZ
 
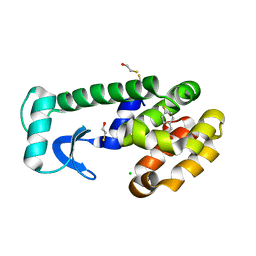 | | T4 Lysozyme Cavity Mutant L99A/M102Q Bound With 4-FluoroPhenEthyl Alcohol | | Descriptor: | 4-FLUOROPHENETHYL ALCOHOL, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Wei, B.Q, Baase, W.A, Weaver, L.H, Matthews, B.W, Shoichet, B.K. | | Deposit date: | 2003-03-31 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Testing a Flexible-receptor Docking Algorithm in a Model Binding Site
J.Mol.Biol., 337, 2004
|
|
1OVH
 
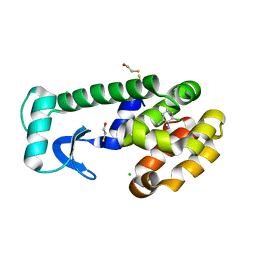 | | T4 Lysozyme Cavity Mutant L99A/M102Q Bound With 2-Chloro-6-Methyl-Aniline | | Descriptor: | 2-CHLORO-6-METHYL-ANILINE, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Wei, B.Q, Baase, W.A, Weaver, L.H, Matthews, B.W, Shoichet, B.K. | | Deposit date: | 2003-03-26 | | Release date: | 2004-04-06 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Testing a Flexible-receptor Docking Algorithm in a Model Binding Site
J.Mol.Biol., 337, 2004
|
|
1OWY
 
 | | T4 Lysozyme Cavity Mutant L99A/M102Q Bound With 2-Propyl-Aniline | | Descriptor: | 2-PROPYL-ANILINE, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Wei, B.Q, Baase, W.A, Weaver, L.H, Matthews, B.W, Shoichet, B.K. | | Deposit date: | 2003-03-31 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Testing a Flexible-receptor Docking Algorithm in a Model Binding Site
J.Mol.Biol., 337, 2004
|
|
1OVJ
 
 | | T4 Lysozyme Cavity Mutant L99A/M102Q Bound with 3-Fluoro-2-Methyl_Aniline | | Descriptor: | 3-FLUORO-2-METHYL-ANILINE, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Wei, B.Q, Baase, W.A, Weaver, L.H, Matthews, B.W, Shoichet, B.K. | | Deposit date: | 2003-03-26 | | Release date: | 2004-04-06 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Testing a Flexible-receptor Docking Algorithm in a Model Binding Site
J.Mol.Biol., 337, 2004
|
|
1PE5
 
 | | Thermolysin with tricyclic inhibitor | | Descriptor: | (6-METHYL-3,4-DIHYDRO-2H-CHROMEN-2-YL)METHYLPHOSPHINATE, 3-METHYLBUTAN-1-AMINE, CALCIUM ION, ... | | Authors: | Juers, D, Holland, D, Morgan, B.P, Bartlett, P.A, Matthews, B.W. | | Deposit date: | 2003-05-21 | | Release date: | 2004-06-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Conformational Constraint and Structural Complementarity in Thermolysin Inhibitors: Structures of Enzyme Complexes and Conclusions
To be Published
|
|
1PE7
 
 | | Thermolysin with bicyclic inhibitor | | Descriptor: | 2-(4-METHYLPHENOXY)ETHYLPHOSPHINATE, 3-METHYLBUTAN-1-AMINE, CALCIUM ION, ... | | Authors: | Juers, D, Yusuff, N, Bartlett, P.A, Matthews, B.W. | | Deposit date: | 2003-05-21 | | Release date: | 2004-06-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Conformational Constraint and Structural Complementarity in Thermolysin Inhibitors: Structures of Enzyme Complexes and Conclusions
To be Published
|
|
1PE8
 
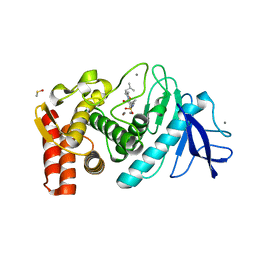 | | Thermolysin with monocyclic inhibitor | | Descriptor: | 2-ETHOXYETHYLPHOSPHINATE, 3-METHYLBUTAN-1-AMINE, CALCIUM ION, ... | | Authors: | Juers, D, Pyun, H.-J, Bartlett, P.A, Matthews, B.W. | | Deposit date: | 2003-05-21 | | Release date: | 2004-06-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Conformational Constraint and Structural Complementarity in Thermolysin Inhibitors: Structures of Enzyme Complexes and Conclusions
To be Published
|
|
1OVK
 
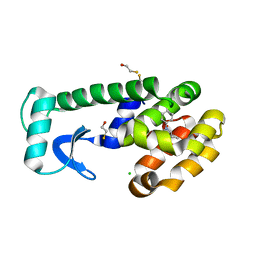 | | T4 Lysozyme Cavity Mutant L99A/M102Q Bound with N-Allyl-Aniline | | Descriptor: | BETA-MERCAPTOETHANOL, CHLORIDE ION, Lysozyme, ... | | Authors: | Wei, B.Q, Baase, W.A, Weaver, L.H, Matthews, B.W, Shoichet, B.K. | | Deposit date: | 2003-03-26 | | Release date: | 2004-04-06 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Testing a Flexible-receptor Docking Algorithm in a Model Binding Site
J.Mol.Biol., 337, 2004
|
|
1OYU
 
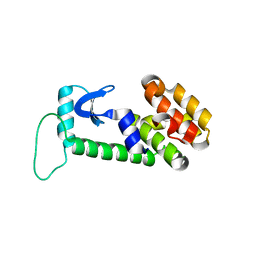 | |
1OV7
 
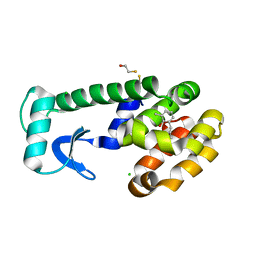 | | T4 Lysozyme Cavity Mutant L99A/M102Q Bound with 2-Allyl-6-Methyl-Phenol | | Descriptor: | 2-ALLYL-6-METHYL-PHENOL, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Wei, B.Q, Baase, W.A, Weaver, L.H, Matthews, B.W, Shoichet, B.K. | | Deposit date: | 2003-03-25 | | Release date: | 2004-04-06 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Testing a Flexible-receptor Docking Algorithm in a Model Binding Site
J.Mol.Biol., 337, 2004
|
|
1OV5
 
 | | T4 Lysozyme Cavity Mutant L99a/M102Q Bound With 2-Allylphenol | | Descriptor: | 2-ALLYLPHENOL, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Wei, B.Q, Baase, W.A, Weaver, L.H, Matthews, B.W, Shoichet, B.K. | | Deposit date: | 2003-03-25 | | Release date: | 2004-04-06 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Testing a Flexible-receptor Docking Algorithm in a Model Binding Site
J.Mol.Biol., 337, 2004
|
|
3BBZ
 | | Structure of the nucleocapsid-binding domain from the mumps virus phosphoprotein | | Descriptor: | BROMIDE ION, FORMIC ACID, P protein | | Authors: | Kingston, R.L, Gay, L.S, Baase, W.S, Matthews, B.W. | | Deposit date: | 2007-11-11 | | Release date: | 2008-05-27 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of the nucleocapsid-binding domain from the mumps virus polymerase; an example of protein folding induced by crystallization
J.Mol.Biol., 379, 2008
|
|
3B2P
 
 | | Crystal structure of E. coli Aminopeptidase N in complex with arginine | | Descriptor: | ARGININE, Aminopeptidase N, GLYCEROL, ... | | Authors: | Anthony, A, Leslie, G, Matthews, B.W. | | Deposit date: | 2007-10-18 | | Release date: | 2008-05-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis for the unusual specificity of Escherichia coli aminopeptidase N.
Biochemistry, 47, 2008
|
|
3B2X
 
 | | Crystal Structure of E. coli Aminopeptidase N in complex with Lysine | | Descriptor: | Aminopeptidase N, GLYCEROL, LYSINE, ... | | Authors: | Addlagatta, A, Gay, L, Matthews, B.W. | | Deposit date: | 2007-10-19 | | Release date: | 2008-05-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural basis for the unusual specificity of Escherichia coli aminopeptidase N.
Biochemistry, 47, 2008
|
|
1FA6
 
 | | CRYSTAL STRUCTURE OF THE CO(II)-BOUND GLYOXALASE I OF ESCHERICHIA COLI | | Descriptor: | COBALT (II) ION, GLYOXALASE I | | Authors: | He, M.M, Clugston, S.L, Honek, J.F, Matthews, B.W. | | Deposit date: | 2000-07-12 | | Release date: | 2000-09-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Determination of the structure of Escherichia coli glyoxalase I suggests a structural basis for differential metal activation.
Biochemistry, 39, 2000
|
|
1F9Z
 
 | | CRYSTAL STRUCTURE OF THE NI(II)-BOUND GLYOXALASE I FROM ESCHERICHIA COLI | | Descriptor: | GLYOXALASE I, NICKEL (II) ION | | Authors: | He, M.M, Clugston, S.L, Honek, J.F, Matthews, B.W. | | Deposit date: | 2000-07-11 | | Release date: | 2000-09-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Determination of the structure of Escherichia coli glyoxalase I suggests a structural basis for differential metal activation.
Biochemistry, 39, 2000
|
|
1FA8
 
 | |
1FA7
 
 | | CRYSTAL STRUCTURE OF CD(II)-BOUND GLYOXALASE I OF ESCHERICHIA COLI | | Descriptor: | CADMIUM ION, GLYOXALASE I | | Authors: | He, M.M, Clugston, S.L, Honek, J.F, Matthews, B.W. | | Deposit date: | 2000-07-12 | | Release date: | 2000-09-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Determination of the structure of Escherichia coli glyoxalase I suggests a structural basis for differential metal activation.
Biochemistry, 39, 2000
|
|
1FA5
 
 | | CRYSTAL STRUCTURE OF THE ZN(II)-BOUND GLYOXALASE I OF ESCHERICHIA COLI | | Descriptor: | GLYOXALASE I, ZINC ION | | Authors: | He, M.M, Clugston, S.L, Honek, J.F, Matthews, B.W. | | Deposit date: | 2000-07-12 | | Release date: | 2000-09-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Determination of the structure of Escherichia coli glyoxalase I suggests a structural basis for differential metal activation.
Biochemistry, 39, 2000
|
|
1G0J
 
 | | CRYSTAL STRUCTURE OF T4 LYSOZYME MUTANT T152S | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, CHLORIDE ION, PROTEIN (LYSOZYME) | | Authors: | Xu, J, Baase, W.A, Quillin, M.L, Matthews, B.W. | | Deposit date: | 2000-10-06 | | Release date: | 2001-05-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and thermodynamic analysis of the binding of solvent at internal sites in T4 lysozyme.
Protein Sci., 10, 2001
|
|
1G0M
 
 | | CRYSTAL STRUCTURE OF T4 LYSOZYME MUTANT T152I | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, CHLORIDE ION, PROTEIN (LYSOZYME) | | Authors: | Xu, J, Baase, W.A, Quillin, M.L, Matthews, B.W. | | Deposit date: | 2000-10-06 | | Release date: | 2001-05-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural and thermodynamic analysis of the binding of solvent at internal sites in T4 lysozyme.
Protein Sci., 10, 2001
|
|
1G0Q
 
 | | CRYSTAL STRUCTURE OF T4 LYSOZYME MUTANT V149I | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, CHLORIDE ION, PROTEIN (LYSOZYME) | | Authors: | Xu, J, Baase, W.A, Quillin, M.L, Matthews, B.W. | | Deposit date: | 2000-10-06 | | Release date: | 2001-05-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and thermodynamic analysis of the binding of solvent at internal sites in T4 lysozyme.
Protein Sci., 10, 2001
|
|
1G0G
 
 | | CRYSTAL STRUCTURE OF T4 LYSOZYME MUTANT T152A | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, CHLORIDE ION, PROTEIN (LYSOZYME) | | Authors: | Xu, J, Baase, W.A, Quillin, M.L, Matthews, B.W. | | Deposit date: | 2000-10-06 | | Release date: | 2001-05-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural and thermodynamic analysis of the binding of solvent at internal sites in T4 lysozyme.
Protein Sci., 10, 2001
|
|
