3REM
 
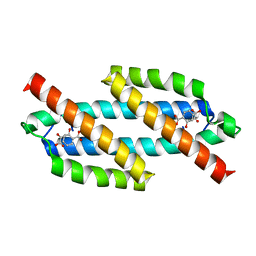 | | Structure of the Isochorismate-Pyruvate Lyase from Pseudomonas aerugionsa with Bound Salicylate and Pyruvate | | Descriptor: | 2-HYDROXYBENZOIC ACID, PYRUVIC ACID, Salicylate biosynthesis protein pchB | | Authors: | Olucha, J, Ouellette, A.N, Luo, Q, Lamb, A.L. | | Deposit date: | 2011-04-04 | | Release date: | 2011-07-27 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | pH Dependence of Catalysis by Pseudomonas aeruginosa Isochorismate-Pyruvate Lyase: Implications for Transition State Stabilization and the Role of Lysine 42.
Biochemistry, 50, 2011
|
|
3S5W
 
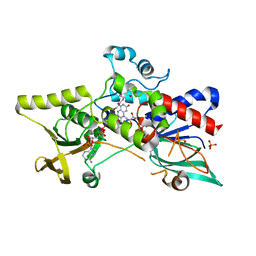 | | Ornithine Hydroxylase (PvdA) from Pseudomonas aeruginosa | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, L-ornithine 5-monooxygenase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Olucha, J, Lamb, A.L. | | Deposit date: | 2011-05-23 | | Release date: | 2011-07-13 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Two Structures of an N-Hydroxylating Flavoprotein Monooxygenase: ORNITHINE HYDROXYLASE FROM PSEUDOMONAS AERUGINOSA.
J.Biol.Chem., 286, 2011
|
|
3S61
 
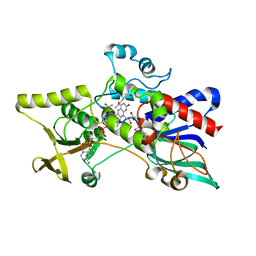 | |
3RET
 
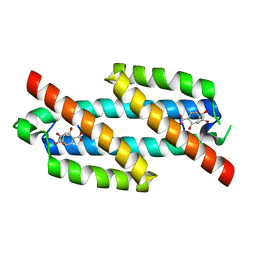 | | Salicylate and Pyruvate Bound Structure of the Isochorismate-Pyruvate Lyase K42E Mutant from Pseudomonas aerugionsa | | Descriptor: | 2-HYDROXYBENZOIC ACID, PYRUVIC ACID, Salicylate biosynthesis protein pchB | | Authors: | Olucha, J, Ouellette, A.N, Luo, Q, Lamb, A.L. | | Deposit date: | 2011-04-05 | | Release date: | 2011-07-27 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | pH Dependence of Catalysis by Pseudomonas aeruginosa Isochorismate-Pyruvate Lyase: Implications for Transition State Stabilization and the Role of Lysine 42.
Biochemistry, 50, 2011
|
|
3HGX
 
 | |
3HGW
 
 | | Apo Structure of Pseudomonas aeruginosa Isochorismate-Pyruvate Lyase I87T mutant | | Descriptor: | CALCIUM ION, Salicylate biosynthesis protein pchB | | Authors: | Luo, Q, Lamb, A.L. | | Deposit date: | 2009-05-14 | | Release date: | 2009-06-30 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure-function analyses of isochorismate-pyruvate lyase from Pseudomonas aeruginosa suggest differing catalytic mechanisms for the two pericyclic reactions of this bifunctional enzyme
Biochemistry, 48, 2009
|
|
