1U2D
 
 | | Structre of transhydrogenaes (dI.NADH)2(dIII.NADPH)1 asymmetric complex | | Descriptor: | GLYCEROL, NAD(P) transhydrogenase subunit alpha part 1, NAD(P) transhydrogenase subunit beta, ... | | Authors: | Mather, O.C, Singh, A, van Boxel, G.I, White, S.A, Jackson, J.B. | | Deposit date: | 2004-07-19 | | Release date: | 2005-01-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Active-site conformational changes associated with hydride transfer in proton-translocating transhydrogenase.
Biochemistry, 43, 2004
|
|
1U2G
 
 | | transhydrogenase (dI.ADPr)2(dIII.NADPH)1 asymmetric complex | | Descriptor: | ADENOSINE-5-DIPHOSPHORIBOSE, GLYCEROL, NAD(P) transhydrogenase subunit alpha part 1, ... | | Authors: | Mather, O.C, Singh, A, van Boxel, G.I, White, S.A, Jackson, J.B. | | Deposit date: | 2004-07-19 | | Release date: | 2005-01-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Active-site conformational changes associated with hydride transfer in proton-translocating transhydrogenase.
Biochemistry, 43, 2004
|
|
1U28
 
 | | R. rubrum transhydrogenase asymmetric complex (dI.NAD+)2(dIII.NADP+)1 | | Descriptor: | GLYCEROL, NAD(P) transhydrogenase subunit alpha part 1, NAD(P) transhydrogenase subunit beta, ... | | Authors: | Mather, O.C, Singh, A, van Boxel, G.I, White, S.A, Jackson, J.B. | | Deposit date: | 2004-07-16 | | Release date: | 2005-01-18 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Active-site conformational changes associated with hydride transfer in proton-translocating transhydrogenase.
Biochemistry, 43, 2004
|
|
1U31
 
 | | recombinant human heart transhydrogenase dIII bound with NADPH | | Descriptor: | GLYCEROL, NAD(P) transhydrogenase, mitochondrial, ... | | Authors: | Mather, O.C, Singh, A, van Boxel, G.I, White, S.A, Jackson, J.B. | | Deposit date: | 2004-07-20 | | Release date: | 2005-01-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Active-site conformational changes associated with hydride transfer in proton-translocating transhydrogenase.
Biochemistry, 43, 2004
|
|
2FR8
 
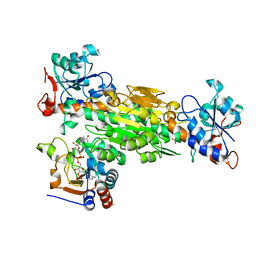 | | Structure of transhydrogenase (dI.R127A.NAD+)2(dIII.NADP+)1 asymmetric complex | | Descriptor: | NAD(P) transhydrogenase subunit alpha part 1, NAD(P) transhydrogenase subunit beta, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Brondijk, T.H, van Boxel, G.I, Mather, O.C, Quirk, P.G, White, S.A, Jackson, J.B. | | Deposit date: | 2006-01-19 | | Release date: | 2006-02-28 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Role of Invariant Amino Acid Residues at the Hydride Transfer Site of Proton-translocating Transhydrogenase.
J.Biol.Chem., 281, 2006
|
|
2FRD
 
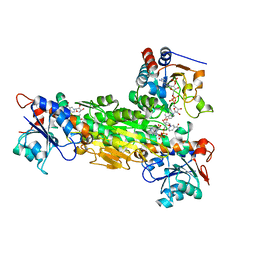 | | Structure of Transhydrogenase (dI.S138A.NADH)2(dIII.NADPH)1 asymmetric complex | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, NAD(P) transhydrogenase subunit alpha part 1, NAD(P) transhydrogenase subunit beta, ... | | Authors: | Brondijk, T.H, van Boxel, G.I, Mather, O.C, Quirk, P.G, White, S.A, Jackson, J.B. | | Deposit date: | 2006-01-19 | | Release date: | 2006-02-28 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | The Role of Invariant Amino Acid Residues at the Hydride Transfer Site of Proton-translocating Transhydrogenase.
J.Biol.Chem., 281, 2006
|
|
2FSV
 
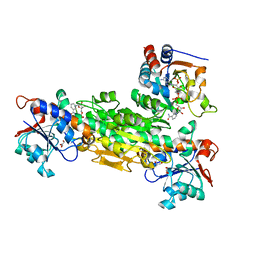 | | Structure of transhydrogenase (dI.D135N.NAD+)2(dIII.E155W.NADP+)1 asymmetric complex | | Descriptor: | GLYCEROL, NAD(P) transhydrogenase subunit alpha part 1, NAD(P) transhydrogenase subunit beta, ... | | Authors: | Brondijk, T.H, van Boxel, G.I, Mather, O.C, Quirk, P.G, White, S.A, Jackson, J.B. | | Deposit date: | 2006-01-23 | | Release date: | 2006-02-28 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Role of Invariant Amino Acid Residues at the Hydride Transfer Site of Proton-translocating Transhydrogenase.
J.Biol.Chem., 281, 2006
|
|
2QB7
 
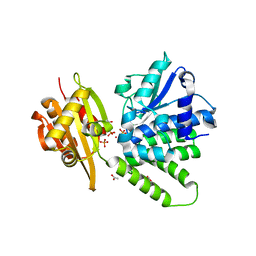 | | Saccharomyces cerevisiae cytosolic exopolyphosphatase, phosphate complex | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, COBALT (II) ION, ... | | Authors: | White, S.A, Ugochukwu, E. | | Deposit date: | 2007-06-16 | | Release date: | 2007-12-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The crystal structure of the cytosolic exopolyphosphatase from Saccharomyces cerevisiae reveals the basis for substrate specificity.
J.Mol.Biol., 371, 2007
|
|
2QB6
 
 | | Saccharomyces cerevisiae cytosolic exopolyphosphatase, sulfate complex | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Exopolyphosphatase, ... | | Authors: | White, S.A, Ugochukwu, E. | | Deposit date: | 2007-06-16 | | Release date: | 2007-12-11 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The crystal structure of the cytosolic exopolyphosphatase from Saccharomyces cerevisiae reveals the basis for substrate specificity.
J.Mol.Biol., 371, 2007
|
|
2QB8
 
 | |
3H7U
 
 | | Crystal structure of the plant stress-response enzyme AKR4C9 | | Descriptor: | ACETATE ION, Aldo-keto reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | White, S.A, Simpson, P.J, Ride, J.P. | | Deposit date: | 2009-04-28 | | Release date: | 2009-08-04 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Characterization of two novel aldo-keto reductases from Arabidopsis: expression patterns, broad substrate specificity, and an open active-site structure suggest a role in toxicant metabolism following stress.
J.Mol.Biol., 392, 2009
|
|
3H7R
 
 | | Crystal structure of the plant stress-response enzyme AKR4C8 | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Aldo-keto reductase, ... | | Authors: | White, S.A, Simpson, P.J, Ride, J.P. | | Deposit date: | 2009-04-28 | | Release date: | 2009-08-04 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Characterization of two novel aldo-keto reductases from Arabidopsis: expression patterns, broad substrate specificity, and an open active-site structure suggest a role in toxicant metabolism following stress.
J.Mol.Biol., 392, 2009
|
|
