1VL3
 
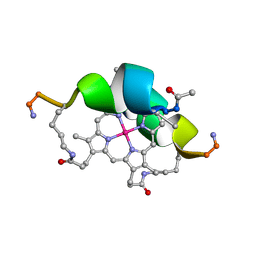 | | DESIGN OF NEW MIMOCHROMES WITH UNIQUE TOPOLOGY | | Descriptor: | CO(III)-(DEUTEROPORPHYRIN IX), GLU-SER-GLN-LEU-HIS-SER-ASN-LYS-ARG | | Authors: | Lombardi, A, Nastri, F, Marasco, D, Maglio, O, De Sanctis, G, Sinibaldi, F, Santucci, R, Coletta, M, Pavone, V. | | Deposit date: | 2004-07-05 | | Release date: | 2004-07-20 | | Last modified: | 2024-10-09 | | Method: | SOLUTION NMR | | Cite: | Design of a New Mimochrome with Unique Topology.
Chemistry, 9, 2003
|
|
3PY9
 
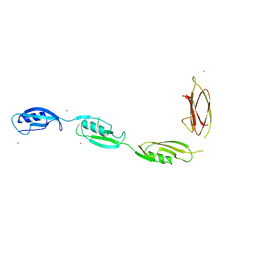 | | X-ray structural studies of the entire extra-cellular region of the Ser/Thr kinase PrkC from Staphylococcus aureus | | Descriptor: | EUROPIUM ION, Protein kinase | | Authors: | Ruggiero, A, Squeglia, F, Marasco, D, Marchetti, R, Molinaro, A, Berisio, R. | | Deposit date: | 2010-12-12 | | Release date: | 2011-01-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray structural studies of the entire extracellular region of the serine/threonine kinase PrkC from Staphylococcus aureus.
Biochem.J., 435, 2011
|
|
2MGV
 
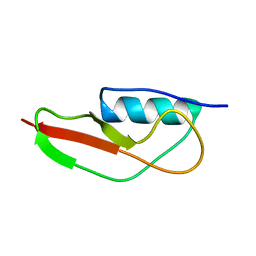 | | NMR structure of PASTA domain of PonA2 from Mycobacterium tuberculosis | | Descriptor: | Bifunctional membrane-associated penicillin-binding protein 1A/1B ponA2 | | Authors: | Calvanese, L, Falcigno, L, Maglione, C, Marasco, D, Ruggiero, A, Squeglia, F, Berisio, R, D'Auria, G. | | Deposit date: | 2013-11-11 | | Release date: | 2013-12-25 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural and binding properties of the PASTA domain of PonA2, a key penicillin binding protein from Mycobacterium tuberculosis.
Biopolymers, 101, 2014
|
|
2J5H
 | | NMR analysis of mouse CRIPTO CFC domain | | Descriptor: | TERATOCARCINOMA-DERIVED GROWTH FACTOR | | Authors: | Calvanese, L, Saporito, A, Marasco, D, D'Auria, G, Minchiotti, G, Pedone, C, Paolillo, L, Falcigno, L, Ruvo, M. | | Deposit date: | 2006-09-18 | | Release date: | 2006-10-02 | | Last modified: | 2018-05-09 | | Method: | SOLUTION NMR | | Cite: | Solution structure of mouse Cripto CFC domain and its inactive variant Trp107Ala.
J. Med. Chem., 49, 2006
|
|
4Z46
 
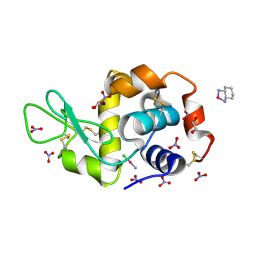 | |
4ZEE
 
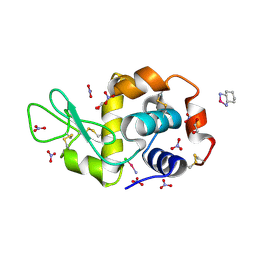 | |
4QY9
 
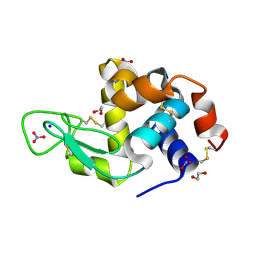 | | X-ray structure of the adduct between hen egg white lysozyme and Auoxo3, a cytotoxic gold(III) compound | | Descriptor: | 1,2-ETHANEDIOL, GOLD ION, Lysozyme C, ... | | Authors: | Russo Krauss, I, Merlino, A. | | Deposit date: | 2014-07-24 | | Release date: | 2014-11-05 | | Last modified: | 2014-11-19 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Interactions of gold-based drugs with proteins: the structure and stability of the adduct formed in the reaction between lysozyme and the cytotoxic gold(iii) compound Auoxo3.
Dalton Trans, 43, 2014
|
|
8BKE
 
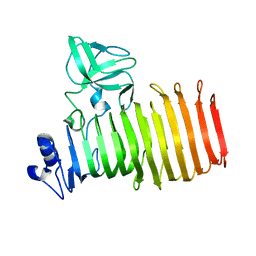 | |
3HHV
 
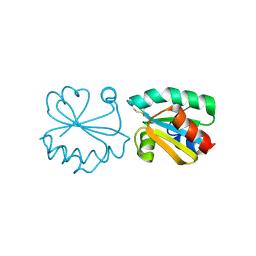 | | The crystal structure of the Thioredoxin A2 from Sulfolobus solfataricus | | Descriptor: | Thioredoxin (TrxA-2) | | Authors: | Ruggiero, A, Masullo, M, Ruocco, M.R, Arcari, P, Zagari, A, Vitagliano, L. | | Deposit date: | 2009-05-18 | | Release date: | 2009-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | The dimeric structure of Sulfolobus solfataricus thioredoxin A2 and the basis of its thermostability
Proteins, 77, 2009
|
|
3LE7
 
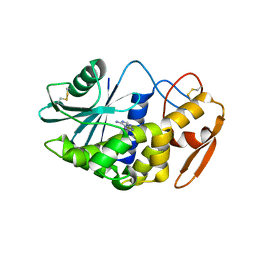 | | Crystal structure of PD-L1 from P. dioica in complex with adenine | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ADENINE, Ribosome-inactivating protein PD-L1/PD-L2 | | Authors: | Ruggiero, A, Berisio, R. | | Deposit date: | 2010-01-14 | | Release date: | 2010-04-14 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The role of the glycan moiety on the structure-function relationships of PD-L1, type 1 ribosome-inactivating protein from P. dioica leaves
Mol Biosyst, 6, 2010
|
|
3NE0
 
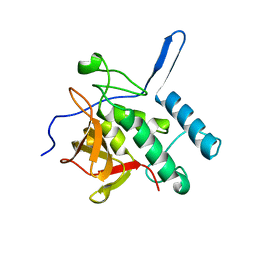 | |
5NZ9
 | | NMR structure of an EphA2-Sam fragment | | Descriptor: | Ephrin type-A receptor 2 | | Authors: | Mercurio, F.A, Leone, M. | | Deposit date: | 2017-05-12 | | Release date: | 2017-12-20 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | The Sam-Sam interaction between Ship2 and the EphA2 receptor: design and analysis of peptide inhibitors.
Sci Rep, 7, 2017
|
|
2LMR
 
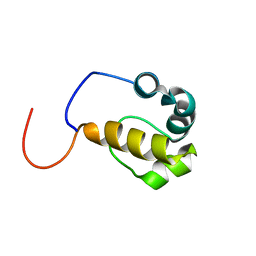 | | Solution structure of the first sam domain of odin | | Descriptor: | Ankyrin repeat and SAM domain-containing protein 1A | | Authors: | Leone, M, Mercurio, F. | | Deposit date: | 2011-12-12 | | Release date: | 2012-03-14 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the First Sam Domain of Odin and Binding Studies with the EphA2 Receptor.
Biochemistry, 51, 2012
|
|
6F7M
 
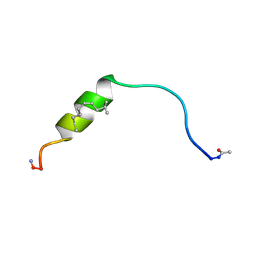 | | NMR structure of EphA2-Sam stapled peptides (S13ST) | | Descriptor: | Ephrin type-A receptor 2 | | Authors: | Mercurio, F.A, Leone, M. | | Deposit date: | 2017-12-11 | | Release date: | 2018-08-01 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Sam domain-based stapled peptides: Structural analysis and interaction studies with the Sam domains from the EphA2 receptor and the lipid phosphatase Ship2.
Bioorg. Chem., 80, 2018
|
|
6F7N
 
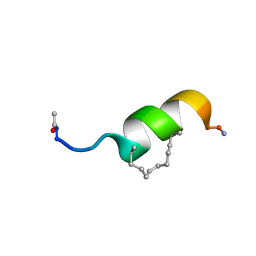 | | NMR structure of EphA2-Sam stapled peptides (S13STshort) | | Descriptor: | Ephrin type-A receptor 2 | | Authors: | Mercurio, F.A, Leone, M. | | Deposit date: | 2017-12-11 | | Release date: | 2018-08-01 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Sam domain-based stapled peptides: Structural analysis and interaction studies with the Sam domains from the EphA2 receptor and the lipid phosphatase Ship2.
Bioorg. Chem., 80, 2018
|
|
6F7O
 | | NMR structure of an Odin-Sam1 stapled peptide | | Descriptor: | Ankyrin repeat and SAM domain-containing protein 1A | | Authors: | Leone, M, Mercurio, F.A. | | Deposit date: | 2017-12-11 | | Release date: | 2018-08-01 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Sam domain-based stapled peptides: Structural analysis and interaction studies with the Sam domains from the EphA2 receptor and the lipid phosphatase Ship2.
Bioorg. Chem., 80, 2018
|
|
2MYQ
 
 | | NMR structure of an Odin-Sam1 fragment | | Descriptor: | Ankyrin repeat and SAM domain-containing protein 1A | | Authors: | Mercurio, F.A, Leone, M. | | Deposit date: | 2015-01-30 | | Release date: | 2015-07-08 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Peptide Fragments of Odin-Sam1: Conformational Analysis and Interaction Studies with EphA2-Sam.
Chembiochem, 16, 2015
|
|
5JMG
 
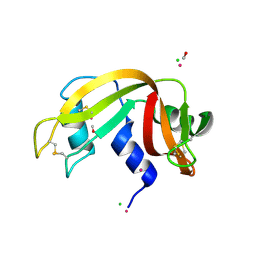 | | X-ray structure of the complex between bovine pancreatic ribonuclease and pentachlorocarbonyliridate(III) (4 days of soaking) | | Descriptor: | CARBON MONOXIDE, CHLORIDE ION, IRIDIUM ION, ... | | Authors: | Caterino, M, Petruk, A.A, Vergara, A, Ferraro, G, Merlino, A. | | Deposit date: | 2016-04-29 | | Release date: | 2016-07-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Mapping the protein-binding sites for iridium(iii)-based CO-releasing molecules.
Dalton Trans, 45, 2016
|
|
5JML
 
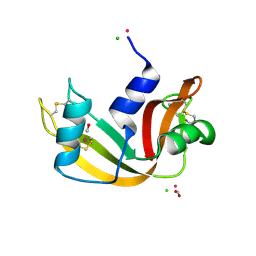 | | X-ray structure of the complex between bovine pancreatic ribonuclease and penthachlorocarbonyliridate(III) (2 months of soaking) | | Descriptor: | CARBON MONOXIDE, CHLORIDE ION, IRIDIUM ION, ... | | Authors: | Caterino, M, Petruk, A.A, Vergara, A, Ferraro, G, Merlino, A. | | Deposit date: | 2016-04-29 | | Release date: | 2016-07-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Mapping the protein-binding sites for iridium(iii)-based CO-releasing molecules.
Dalton Trans, 45, 2016
|
|
4NHS
 
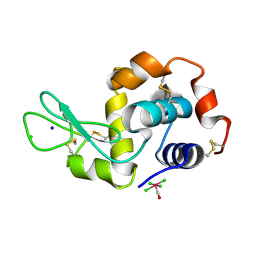 | | X-ray structure of the complex between hen egg white lysozyme and pentachlorocarbonyliridate(III) (9 days) | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Petruk, A.A, Bikiel, D.E, Vergara, A, Merlino, A. | | Deposit date: | 2013-11-05 | | Release date: | 2014-09-17 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Interaction between proteins and Ir based CO releasing molecules: mechanism of adduct formation and CO release.
Inorg.Chem., 53, 2014
|
|
4NIJ
 
 | | X-ray structure of the complex between hen egg white lysozyme and pentachlorocarbonyliridate(III) (30 days) | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Petruk, A.A, Bikiel, D.E, Vergara, A, Merlino, A. | | Deposit date: | 2013-11-06 | | Release date: | 2014-09-17 | | Last modified: | 2015-06-24 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Interaction between proteins and Ir based CO releasing molecules: mechanism of adduct formation and CO release.
Inorg.Chem., 53, 2014
|
|
4NHP
 
 | | X-ray structure of the complex between the hen egg white lysozyme and pentachlorocarbonyliridate (III) (4 days) | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Petruk, A.A, Bikiel, D.E, Vergara, A, Merlino, A. | | Deposit date: | 2013-11-05 | | Release date: | 2014-09-17 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Interaction between proteins and Ir based CO releasing molecules: mechanism of adduct formation and CO release.
Inorg.Chem., 53, 2014
|
|
4NHT
 
 | | X-ray structure of the complex between hen egg white lysozyme and pentachlorocarbonyliridate(III) (6 days) | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Petruk, A.A, Bikiel, D.E, Vergara, A, Merlino, A. | | Deposit date: | 2013-11-05 | | Release date: | 2014-09-17 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Interaction between proteins and Ir based CO releasing molecules: mechanism of adduct formation and CO release.
Inorg.Chem., 53, 2014
|
|
4N9R
 
 | | X-ray structure of the complex between hen egg white lysozyme and pentacholrocarbonyliridate(III) (1 day) | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Petruk, A.A, Bikiel, D.E, Vergara, A, Merlino, A. | | Deposit date: | 2013-10-21 | | Release date: | 2015-12-30 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Interaction between proteins and Ir based CO releasing molecules: mechanism of adduct formation and CO release.
Inorg.Chem., 53, 2014
|
|
4NHQ
 
 | | X-ray structure of the complex between hen egg white lysozyme and pentachlorocarbonyliridate(III) (5 days) | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Petruk, A.A, Bikiel, D.E, Vergara, A, Merlino, A. | | Deposit date: | 2013-11-05 | | Release date: | 2014-09-17 | | Last modified: | 2015-06-24 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Interaction between proteins and Ir based CO releasing molecules: mechanism of adduct formation and CO release.
Inorg.Chem., 53, 2014
|
|
