7X7N
 
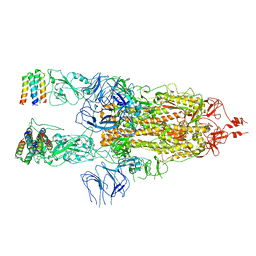 | | 3D model of the 3-RBD up single trimeric spike protein of SARS-CoV2 in the presence of synthetic peptide SIH-5. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, Synthetic peptide SIH-5 | | Authors: | Khatri, B, Pramanick, I, Malladi, S.K, Rajmani, R.S, Kumar, S, Ghosh, P, Sengupta, N, Rahisuddin, R, Kumaran, S, Ringe, R.P, Varadarajan, R, Dutta, S, Chatterjee, J. | | Deposit date: | 2022-03-10 | | Release date: | 2022-04-27 | | Last modified: | 2022-11-16 | | Method: | ELECTRON MICROSCOPY (4.47 Å) | | Cite: | A dimeric proteomimetic prevents SARS-CoV-2 infection by dimerizing the spike protein.
Nat.Chem.Biol., 18, 2022
|
|
4N6A
 
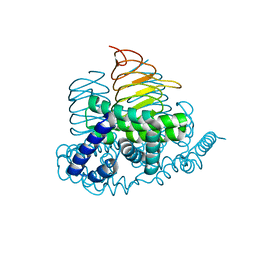 | | Soybean Serine Acetyltransferase Apoenzyme | | Descriptor: | PHOSPHATE ION, Serine Acetyltransferase Apoenzyme | | Authors: | Yi, H, Dey, S, Kumaran, S, Krishnan, H.B, Jez, J.M. | | Deposit date: | 2013-10-11 | | Release date: | 2013-11-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure of soybean serine acetyltransferase and formation of the cysteine regulatory complex as a molecular chaperone.
J.Biol.Chem., 288, 2013
|
|
4N6B
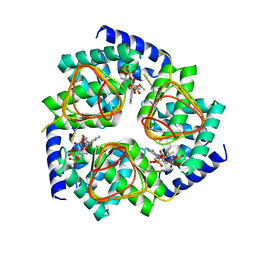 
 | | Soybean Serine Acetyltransferase Complexed with CoA | | Descriptor: | COENZYME A, Serine Acetyltransferase Apoenzyme | | Authors: | Yi, H, Dey, S, Kumaran, S, Krishnan, H.B, Jez, J.M. | | Deposit date: | 2013-10-11 | | Release date: | 2013-11-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.005 Å) | | Cite: | Structure of soybean serine acetyltransferase and formation of the cysteine regulatory complex as a molecular chaperone.
J.Biol.Chem., 288, 2013
|
|
6LCN
 
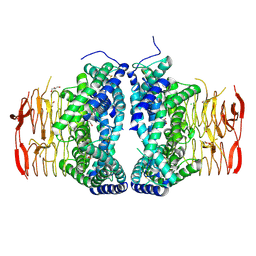 | | Crystal structure of Serine Acetyltransferase from Planctomyces limnophilus at 2.15A | | Descriptor: | CHLORIDE ION, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Kumar, N, Singh, R.P, Singh, A.K, Kumaran, S. | | Deposit date: | 2019-11-19 | | Release date: | 2020-12-02 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Understanding Mechanics of competitive-allostery Using Engineered Cysteine Synthase Assembly
To Be Published
|
|
4RUH
 
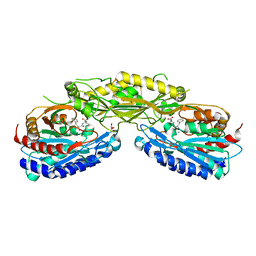 | | Crystal structure of Human Carnosinase-2 (CN2) in complex with inhibitor, Bestatin at 2.25 A | | Descriptor: | 2-(3-AMINO-2-HYDROXY-4-PHENYL-BUTYRYLAMINO)-4-METHYL-PENTANOIC ACID, Cytosolic non-specific dipeptidase, GLYCEROL, ... | | Authors: | pandya, V, Kaushik, A, Singh, A.K, Singh, R.P, Kumaran, S. | | Deposit date: | 2014-11-19 | | Release date: | 2015-11-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal structure of Human Carnosinase-2 (CN2) in complex with inhibitor, Bestatin at 2.25 A
TO BE PUBLISHED
|
|
5DBH
 
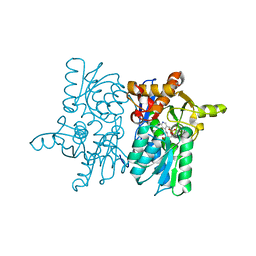 | |
5DBE
 
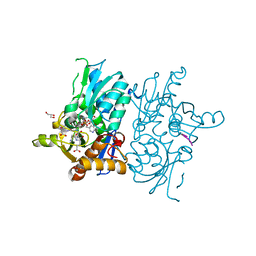 | | Crystal structure of O-acetylserine sulfhydrylase from Haemophilus influenzae in complex with pre-reactive O-acetyl serine, alpha-aminoacrylate reaction intermediate and peptide inhibitor at the resolution of 2.25A | | Descriptor: | 2-{[(E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}prop-2-enoic acid, C-terminal peptide from Serine acetyltransferase, Cysteine synthase, ... | | Authors: | Singh, A.K, Kaushik, A, Ekka, M.K, Kumaran, S. | | Deposit date: | 2015-08-21 | | Release date: | 2015-09-09 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal Structure Of O-Acetylserine Sulfhydrylase From Haemophilus Influenzae In Complex With Pre-Reactive O-Acetyl Serine, Alpha-Aminoacrylate Reactionintermediate And Peptide Inhibitor At The Resolution Of 2.25A
To Be Published
|
|
8I7E
 
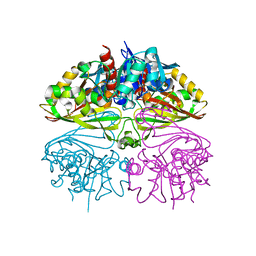 | | Crystal structure of Glyceraldehyde 3-phosphate dehydrogenase from Salmonella typhi at 2.05A | | Descriptor: | Glyceraldehyde-3-phosphate dehydrogenase | | Authors: | Kumar, N, Dilawari, R, Chaubey, G.K, Modanwal, R, Talukdar, S, Dhiman, A, Chaudhary, S, Patidar, A, Kumar, A, Raje, C.I, Raje, M, Kumaran, S. | | Deposit date: | 2023-01-31 | | Release date: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Crystal structure of Glyceraldehyde 3-phosphate dehydrogenase from Salmonella typhi at 2.05A
To Be Published
|
|
5XCW
 
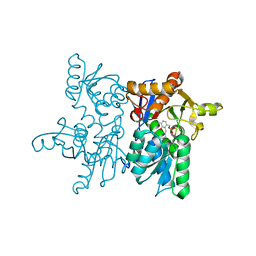 | |
5XCP
 
 | |
5XCN
 
 | |
7CM8
 
 | | High resolution crystal structure of M92A mutant of O-acetyl-L-serine sulfhydrylase from Haemophilus influenzae | | Descriptor: | Cysteine synthase, GLYCEROL, SODIUM ION | | Authors: | Kaushik, A, Rahisuddin, R, Saini, N, Kumaran, S. | | Deposit date: | 2020-07-25 | | Release date: | 2020-08-19 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular mechanism of selective substrate engagement and inhibitor disengagement of cysteine synthase.
J.Biol.Chem., 296, 2020
|
|
7C35
 
 | |
5XA2
 
 | |
7DJQ
 
 | |
7EWO
 
 | | Crystal Structure of D67A, E68P double mutant of O-acetyl-L-serine sulfhydrylase from Haemophilus influenzae | | Descriptor: | Cysteine synthase | | Authors: | Rahisuddin, R, Ekka, M.K, Singh, A.K, Saini, N, Patel, M, Kumar, N, Kumaran, S. | | Deposit date: | 2021-05-25 | | Release date: | 2021-06-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of D67A, E68P double mutant of O-acetyl-L-serine sulfhydrylase from Haemophilus influenzae
To Be Published
|
|
4N69
 
 | | Soybean Serine Acetyltransferase Complexed with Serine | | Descriptor: | PHOSPHATE ION, SERINE, Serine Acetyltransferase Apoenzyme | | Authors: | Yi, H, Dey, S, Kumaran, S, Krishnan, H.B, Jez, J.M. | | Deposit date: | 2013-10-11 | | Release date: | 2013-11-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of soybean serine acetyltransferase and formation of the cysteine regulatory complex as a molecular chaperone.
J.Biol.Chem., 288, 2013
|
|
4HO1
 
 | |
4ZU1
 
 | | Crystal Structure of O-Acetylserine Sulfhydrylase from Haemophilus influenzae in complex with O-acetyl serine and peptide inhibitor | | Descriptor: | C-terminal peptide from Serine acetyltransferase, Cysteine synthase, GLYCEROL, ... | | Authors: | Ekka, M.K, Singh, A.K, Kaushik, A, Kumaran, S. | | Deposit date: | 2015-05-15 | | Release date: | 2015-06-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.202 Å) | | Cite: | Crystal Structure of O-Acetylserine Sulfhydrylase from Haemophilus inuen-zae in complex with O-acetyl serine and peptide inhibitor
To Be Published
|
|
4ZU6
 
 | | Crystal Structure of O-Acetylserine Sulfhydrylase from Haemophilus influenzae in complex with pre-reactive o-acetyl serine, alpha-aminoacrylate reaction intermediate and Peptide inhibitor at the resolution of 2.25A | | Descriptor: | C-terminal peptide from Serine acetyltransferase, Cysteine synthase, O-ACETYLSERINE, ... | | Authors: | Ekka, M.K, Singh, A.K, Kaushik, A, Kumaran, S. | | Deposit date: | 2015-05-15 | | Release date: | 2015-06-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Crystal Structure of O-Acetylserine Sulfhydrylase from Haemophilus inuen-zae in complex with pre-reactive o-acetyl serine and peptide inhibitor
To Be Published
|
|
4ORE
 
 | |
4NU8
 
 | |
4G12
 
 | | Crystal structure of putative TetR family transcriptional regulator, Fad35R, from Mycobacterium tuberculosis | | Descriptor: | GLYCEROL, Probable transcriptional regulatory protein (Probably TETR-FAMILY) | | Authors: | Singh, A.K, Manjasetty, B.A, Singh, V, Mittal, M, Kumaran, S. | | Deposit date: | 2012-07-10 | | Release date: | 2013-07-10 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.44 Å) | | Cite: | Crystal structure of putative TetR family transcriptional regulator, Fad35R, from Mycobacterium tuberculosis
to be published
|
|
4G1P
 
 | | Structural and Mechanistic Basis of Substrate Recognition by Novel Di-peptidase Dug1p From Saccharomyces cerevisiae | | Descriptor: | CYSTEINE, Cys-Gly metallodipeptidase DUG1, GLYCINE, ... | | Authors: | Singh, A.K, Singh, M, Pandya, V.K, Singh, V, Mittal, M, Kumaran, S. | | Deposit date: | 2012-07-11 | | Release date: | 2013-07-17 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.547 Å) | | Cite: | Structural and Mechanistic Basis of Substrate Recognition by Novel Di-peptidase Dug1p From Saccromyces cerevesiae
To be published
|
|
4GWO
 
 | |
