3VQM
 
 | |
3A6M
 
 | |
3WCU
 
 | | The structure of a deoxygenated 400 kda hemoglobin provides a more accurate description of the cooperative mechanism of giant hemoglobins: Deoxygenated form | | Descriptor: | A1 globin chain of giant V2 hemoglobin, A2 globin chain of giant V2 hemoglobin, B1 globin chain of giant V2 hemoglobin, ... | | Authors: | Numoto, N, Nakagawa, T, Ohara, R, Hasegawa, T, Kita, A, Yoshida, T, Maruyama, T, Imai, K, Fukumori, Y, Miki, K. | | Deposit date: | 2013-06-01 | | Release date: | 2014-06-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The structure of a deoxygenated 400 kDa haemoglobin reveals ternary- and quaternary-structural changes of giant haemoglobins
Acta Crystallogr.,Sect.D, 70, 2014
|
|
3WCV
 
 | | The structure of a deoxygenated 400 kda hemoglobin provides a more accurate description of the cooperative mechanism of giant hemoglobins: CA bound form | | Descriptor: | A1 globin chain of giant V2 hemoglobin, A2 globin chain of giant V2 hemoglobin, B1 globin chain of giant V2 hemoglobin, ... | | Authors: | Numoto, N, Nakagawa, T, Ohara, R, Hasegawa, T, Kita, A, Yoshida, T, Maruyama, T, Imai, K, Fukumori, Y, Miki, K. | | Deposit date: | 2013-06-01 | | Release date: | 2014-06-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The structure of a deoxygenated 400 kDa haemoglobin reveals ternary- and quaternary-structural changes of giant haemoglobins
Acta Crystallogr.,Sect.D, 70, 2014
|
|
3WCW
 
 | | The structure of a deoxygenated 400 kda hemoglobin provides a more accurate description of the cooperative mechanism of giant hemoglobins: MG bound form | | Descriptor: | A1 globin chain of giant V2 hemoglobin, A2 globin chain of giant V2 hemoglobin, B1 globin chain of giant V2 hemoglobin, ... | | Authors: | Numoto, N, Nakagawa, T, Ohara, R, Hasegawa, T, Kita, A, Yoshida, T, Maruyama, T, Imai, K, Fukumori, Y, Miki, K. | | Deposit date: | 2013-06-01 | | Release date: | 2014-06-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The structure of a deoxygenated 400 kDa haemoglobin reveals ternary- and quaternary-structural changes of giant haemoglobins
Acta Crystallogr.,Sect.D, 70, 2014
|
|
3A5C
 
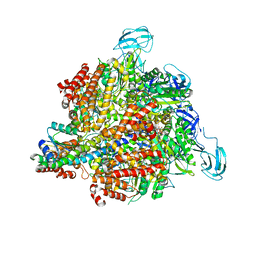 | | Inter-subunit interaction and quaternary rearrangement defined by the central stalk of prokaryotic V1-ATPase | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, V-type ATP synthase alpha chain, V-type ATP synthase beta chain, ... | | Authors: | Numoto, N, Hasegawa, Y, Takeda, K, Miki, K. | | Deposit date: | 2009-08-06 | | Release date: | 2009-10-13 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (4.51 Å) | | Cite: | Inter-subunit interaction and quaternary rearrangement defined by the central stalk of prokaryotic V1-ATPase
Embo Rep., 10, 2009
|
|
2ZZW
 
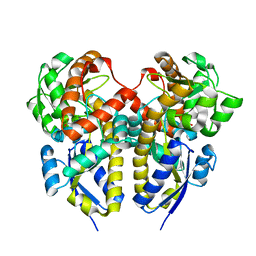 | | Crystal Structure of a Periplasmic Substrate Binding Protein in Complex with Zinc and Lactate | | Descriptor: | ABC transporter, solute-binding protein, LACTIC ACID, ... | | Authors: | Akiyama, N, Takeda, K, Miki, K. | | Deposit date: | 2009-02-27 | | Release date: | 2009-08-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of a periplasmic substrate-binding protein in complex with calcium lactate
J.Mol.Biol., 392, 2009
|
|
3AAC
 
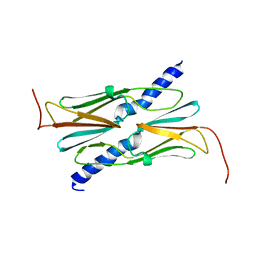 | | Small heat shock protein hsp14.0 with the mutations of I120F and I122F in the form II crystal | | Descriptor: | Putative uncharacterized protein ST1653 | | Authors: | Takeda, K, Hayashi, T, Abe, T, Hirano, Y, Hanazono, Y, Yohda, M, Miki, K. | | Deposit date: | 2009-11-13 | | Release date: | 2010-11-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Dimer structure and conformational variability in the N-terminal region of an archaeal small heat shock protein, StHsp14.0
J.Struct.Biol., 174, 2011
|
|
3WJV
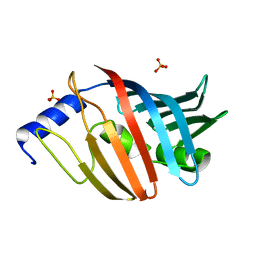 
 | | Crystal structure of the L68E variant of mLolB | | Descriptor: | Outer-membrane lipoprotein LolB, SULFATE ION | | Authors: | Takeda, K, Tokuda, H, Miki, K. | | Deposit date: | 2013-10-16 | | Release date: | 2014-03-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Roles of the Protruding Loop of Factor B Essential for the Localization of Lipoproteins (LolB) in the Anchoring of Bacterial Triacylated Proteins to the Outer Membran
J.Biol.Chem., 289, 2014
|
|
3WCT
 
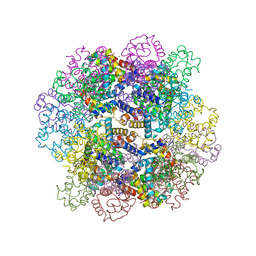 | | The structure of a deoxygenated 400 kda hemoglobin provides a more accurate description of the cooperative mechanism of giant hemoglobins: Oxygenated form | | Descriptor: | A1 globin chain of giant V2 hemoglobin, A2 globin chain of giant V2 hemoglobin, B1 globin chain of giant V2 hemoglobin, ... | | Authors: | Numoto, N, Nakagawa, T, Ohara, R, Hasegawa, T, Kita, A, Yoshida, T, Maruyama, T, Imai, K, Fukumori, Y, Miki, K. | | Deposit date: | 2013-06-01 | | Release date: | 2014-06-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The structure of a deoxygenated 400 kDa haemoglobin reveals ternary- and quaternary-structural changes of giant haemoglobins
Acta Crystallogr.,Sect.D, 70, 2014
|
|
3WOA
 
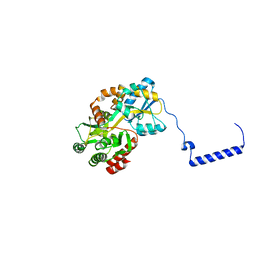 | |
3WMM
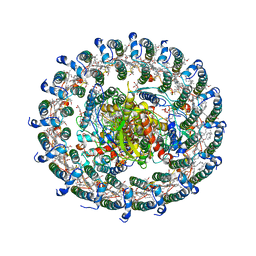 
 | | Crystal structure of the LH1-RC complex from Thermochromatium tepidum in C2 form | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, ... | | Authors: | Niwa, S, Takeda, K, Wang-Otomo, Z.-Y, Miki, K. | | Deposit date: | 2013-11-22 | | Release date: | 2014-04-09 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.008 Å) | | Cite: | Structure of the LH1-RC complex from Thermochromatium tepidum at 3.0 angstrom
Nature, 508, 2014
|
|
2ZC8
 
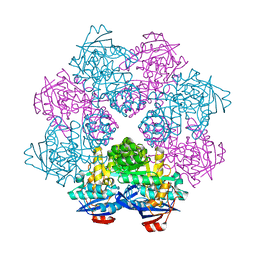 | | Crystal structure of N-Acylamino Acid Racemase from Thermus thermophilus HB8 | | Descriptor: | N-acylamino acid racemase | | Authors: | Hayashida, M, Kim, S.H, Takeda, K, Hisano, T, Miki, K. | | Deposit date: | 2007-11-05 | | Release date: | 2008-02-26 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of N-acylamino acid racemase from Thermus thermophilus HB8
Proteins, 71, 2008
|
|
3W3A
 
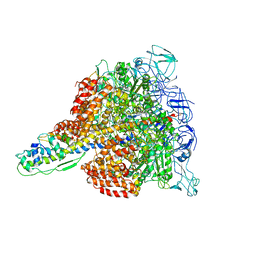 | | Crystal structure of V1-ATPase at 3.9 angstrom resolution | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, V-type ATP synthase alpha chain, V-type ATP synthase beta chain, ... | | Authors: | Nagamatsu, Y, Takeda, K, Kuranaga, T, Numoto, N, Miki, K. | | Deposit date: | 2012-12-14 | | Release date: | 2013-05-15 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.9 Å) | | Cite: | Origin of Asymmetry at the Intersubunit Interfaces of V1-ATPase from Thermusthermophilus
J.Mol.Biol., 425, 2013
|
|
3AAB
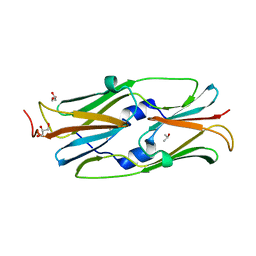 
 | | Small heat shock protein hsp14.0 with the mutations of I120F and I122F in the form I crystal | | Descriptor: | GLYCEROL, ISOPROPYL ALCOHOL, Putative uncharacterized protein ST1653 | | Authors: | Takeda, K, Hayashi, T, Abe, T, Hirano, Y, Hanazono, Y, Yohda, M, Miki, K. | | Deposit date: | 2009-11-13 | | Release date: | 2010-11-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.851 Å) | | Cite: | Dimer structure and conformational variability in the N-terminal region of an archaeal small heat shock protein, StHsp14.0
J.Struct.Biol., 174, 2011
|
|
3WJU
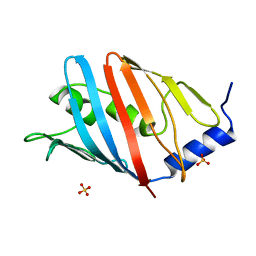 
 | |
2Z4I
 
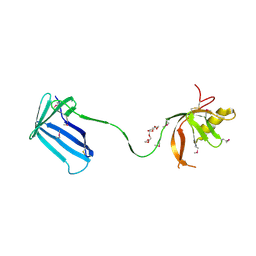 | | Crystal structure of the Cpx pathway activator NlpE from Escherichia coli | | Descriptor: | Copper homeostasis protein cutF, HEXAETHYLENE GLYCOL, SULFATE ION | | Authors: | Hirano, Y, Hossain, M.M, Takeda, K, Tokuda, H, Miki, K. | | Deposit date: | 2007-06-18 | | Release date: | 2007-09-04 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural Studies of the Cpx Pathway Activator NlpE on the Outer Membrane of Escherichia coli
Structure, 15, 2007
|
|
2Z6W
 
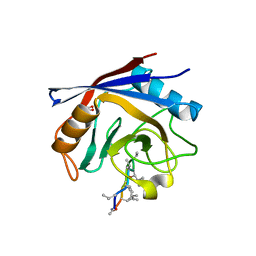 | | Crystal structure of human cyclophilin D in complex with cyclosporin A | | Descriptor: | CITRIC ACID, CYCLOSPORIN A, PEPTIDYL-PROLYL CIS-TRANS ISOMERASE | | Authors: | Kajitani, K, Fujihashi, M, Kobayashi, Y, Shimizu, S, Tsujimoto, Y, Miki, K. | | Deposit date: | 2007-08-09 | | Release date: | 2008-04-29 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (0.96 Å) | | Cite: | Crystal Structure of Human Cyclophilin D in Complex with its Inhibitor, Cyclosporin a at 0.96-A Resolution.
Proteins, 70, 2008
|
|
3WJT
 
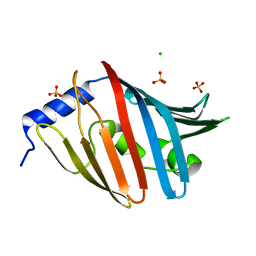 | | Crystal structure of the L68D variant of mLolB | | Descriptor: | CHLORIDE ION, Outer-membrane lipoprotein LolB, SULFATE ION | | Authors: | Takeda, K, Tokuda, H, Miki, K. | | Deposit date: | 2013-10-16 | | Release date: | 2014-03-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Roles of the Protruding Loop of Factor B Essential for the Localization of Lipoproteins (LolB) in the Anchoring of Bacterial Triacylated Proteins to the Outer Membran
J.Biol.Chem., 289, 2014
|
|
2ZPD
 
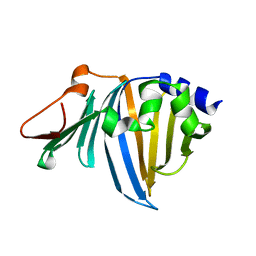 | | Crystal structure of the R43L mutant of LolA in the open form | | Descriptor: | Outer-membrane lipoprotein carrier protein | | Authors: | Takeda, K, Yokota, N, Oguchi, Y, Tokuda, H, Miki, K. | | Deposit date: | 2008-07-10 | | Release date: | 2008-08-05 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Opening and closing of the hydrophobic cavity of LolA coupled to lipoprotein binding and release.
J.Biol.Chem., 283, 2008
|
|
2Z4H
 
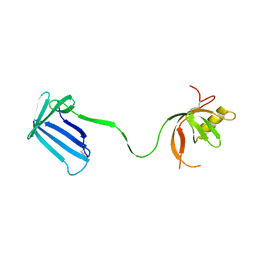 | | Crystal structure of the Cpx pathway activator NlpE from Escherichia coli | | Descriptor: | Copper homeostasis protein cutF, SULFATE ION | | Authors: | Hirano, Y, Hossain, M.M, Takeda, K, Tokuda, H, Miki, K. | | Deposit date: | 2007-06-18 | | Release date: | 2007-09-04 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural Studies of the Cpx Pathway Activator NlpE on the Outer Membrane of Escherichia coli
Structure, 15, 2007
|
|
3A5D
 
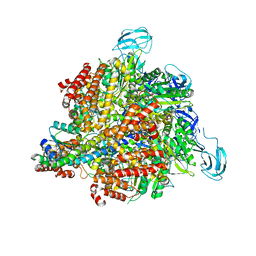 | | Inter-subunit interaction and quaternary rearrangement defined by the central stalk of prokaryotic V1-ATPase | | Descriptor: | V-type ATP synthase alpha chain, V-type ATP synthase beta chain, V-type ATP synthase subunit D, ... | | Authors: | Numoto, N, Hasegawa, Y, Takeda, K, Miki, K. | | Deposit date: | 2009-08-06 | | Release date: | 2009-10-13 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (4.8 Å) | | Cite: | Inter-subunit interaction and quaternary rearrangement defined by the central stalk of prokaryotic V1-ATPase
Embo Rep., 10, 2009
|
|
3AZC
 
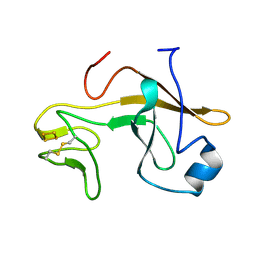 | | Crystal structure of the soluble part of cytochrome b6f complex iron-sulfur subunit from Thermosynechococcus elongatus BP-1 | | Descriptor: | Cytochrome b6-f complex iron-sulfur subunit, FE2/S2 (INORGANIC) CLUSTER | | Authors: | Veit, S, Takeda, K, Tsunoyama, Y, Roegner, M, Miki, K. | | Deposit date: | 2011-05-23 | | Release date: | 2012-05-23 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of a thermophilic cyanobacterial b(6)f-type Rieske protein
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3W07
 
 | |
3VX3
 
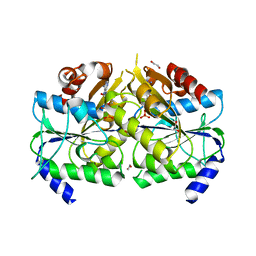 | | Crystal structure of [NiFe] hydrogenase maturation protein HypB from Thermococcus kodakarensis KOD1 | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-DIPHOSPHATE, ATPase involved in chromosome partitioning, ... | | Authors: | Sasaki, D, Watanabe, S, Miki, K. | | Deposit date: | 2012-09-09 | | Release date: | 2013-02-27 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Identification and Structure of a Novel Archaeal HypB for [NiFe] Hydrogenase Maturation
J.Mol.Biol., 425, 2013
|
|
