6YLA
 
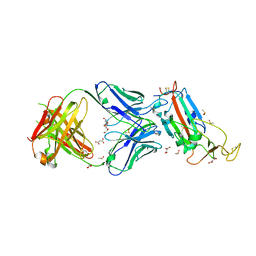 | | Crystal structure of the SARS-CoV-2 receptor binding domain in complex with CR3022 Fab | | Descriptor: | 2-(2-METHOXYETHOXY)ETHANOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, DIMETHYL SULFOXIDE, ... | | Authors: | Huo, J, Zhao, Y, Ren, J, Zhou, D, Ginn, H.M, Fry, E.E, Owens, R, Stuart, D.I. | | Deposit date: | 2020-04-06 | | Release date: | 2020-04-15 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Neutralization of SARS-CoV-2 by Destruction of the Prefusion Spike.
Cell Host Microbe, 28, 2020
|
|
2DBE
 
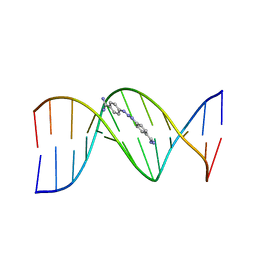 | | CRYSTAL STRUCTURE OF A BERENIL-DODECANUCLEOTIDE COMPLEX: THE ROLE OF WATER IN SEQUENCE-SPECIFIC LIGAND BINDING | | Descriptor: | BERENIL, DNA (5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3') | | Authors: | Brown, D.G, Sanderson, M.R, Skelly, J.V, Jenkins, T.C, Brown, T, Garman, E, Stuart, D.I, Neidle, S. | | Deposit date: | 1990-03-19 | | Release date: | 1991-07-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of a berenil-dodecanucleotide complex: the role of water in sequence-specific ligand binding.
EMBO J., 9, 1990
|
|
9FQW
 
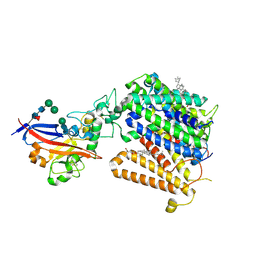 | | Cryo-EM structure of MmCAT1 bound with FrMLV-RBD in the ornithine-bound inward-occluded state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHOLESTEROL HEMISUCCINATE, High affinity cationic amino acid transporter 1,Green fluorescent protein, ... | | Authors: | Ye, M, Zhou, D, Pike, A.C.W, Wang, S, Wang, D, Bakshi, S, Brooke, L, Williams, E, Elkins, J, Stuart, D.I, Sauer, D.B. | | Deposit date: | 2024-06-17 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.01 Å) | | Cite: | Amino acid and viral binding by the high-affinity Cationic Amino acid Transporter 1 (CAT1) from Mus musculus
To Be Published
|
|
9FQU
 
 | | Cryo-EM structure of MmCAT1 bound with FrMLV-RBD in the arginine-bound inward-occluded state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ARGININE, ... | | Authors: | Ye, M, Zhou, D, Pike, A.C.W, Wang, S, Wang, D, Bakshi, S, Brooke, L, Williams, E, Elkins, J, Stuart, D.I, Sauer, D.B. | | Deposit date: | 2024-06-17 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (2.79 Å) | | Cite: | Amino acid and viral binding by the high-affinity Cationic Amino acid Transporter 1 (CAT1) from Mus musculus
To Be Published
|
|
9FQT
 
 | | Cryo-EM structure of MmCAT1 bound with FrMLV-RBD in the apo inward-open state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHOLESTEROL HEMISUCCINATE, ... | | Authors: | Ye, M, Zhou, D, Pike, A.C.W, Wang, S, Wang, D, Bakshi, S, Brooke, L, Williams, E, Elkins, J, Stuart, D.I, Sauer, D.B. | | Deposit date: | 2024-06-17 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Amino acid and viral binding by the high-affinity Cationic Amino acid Transporter 1 (CAT1) from Mus musculus
To Be Published
|
|
9FQV
 
 | | Cryo-EM structure of MmCAT1 bound with FrMLV-RBD in the lysine-bound inward-occluded state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHOLESTEROL HEMISUCCINATE, High affinity cationic amino acid transporter 1,Green fluorescent protein, ... | | Authors: | Ye, M, Zhou, D, Pike, A.C.W, Wang, S, Wang, D, Bakshi, S, Brooke, L, Williams, E, Elkins, J, Stuart, D.I, Sauer, D.B. | | Deposit date: | 2024-06-17 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (2.95 Å) | | Cite: | Amino acid and viral binding by the high-affinity Cationic Amino acid Transporter 1 (CAT1) from Mus musculus
To Be Published
|
|
2CDG
 
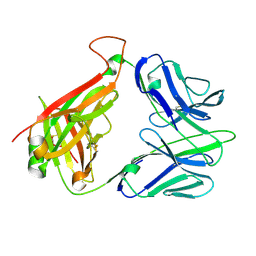 | | Structure and binding kinetics of three different human CD1d-alpha- Galactosylceramide-specific T cell receptors (TCR 5B) | | Descriptor: | TCR 5E | | Authors: | Gadola, S.D, Koch, M, Marles-Wright, J, Lissin, N.M, Sheperd, D, Matulis, G, Harlos, K, Villiger, P.M, Stuart, D.I, Jakobsen, B.K, Cerundolo, V, Jones, E.Y. | | Deposit date: | 2006-01-23 | | Release date: | 2006-03-07 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structrue and Binding Kinetics of Three Different Human Cd1D-Alpha-Galactosylceramide-Specific T Cell Receptors
J.Exp.Med., 203, 2006
|
|
2YIA
 
 | | Structure of the RNA polymerase VP1 from Infectious Pancreatic Necrosis Virus | | Descriptor: | POTASSIUM ION, RNA-DIRECTED RNA POLYMERASE | | Authors: | Graham, S.C, Sarin, L.P, Bahar, M.W, Myers, R.A, Stuart, D.I, Bamford, D.H, Grimes, J.M. | | Deposit date: | 2011-05-11 | | Release date: | 2011-07-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.02 Å) | | Cite: | The N-Terminus of the RNA Polymerase from Infectious Pancreatic Necrosis Virus is the Determinant of Genome Attachment.
Plos Pathog., 7, 2011
|
|
2YI9
 
 | | Structure of the RNA polymerase VP1 from Infectious Pancreatic Necrosis Virus in complex with magnesium | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, POTASSIUM ION, ... | | Authors: | Graham, S.C, Sarin, L.P, Bahar, M.W, Myers, R.A, Stuart, D.I, Bamford, D.H, Grimes, J.M. | | Deposit date: | 2011-05-11 | | Release date: | 2011-07-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The N-Terminus of the RNA Polymerase from Infectious Pancreatic Necrosis Virus is the Determinant of Genome Attachment.
Plos Pathog., 7, 2011
|
|
2YI8
 
 | | Structure of the RNA polymerase VP1 from Infectious Pancreatic Necrosis Virus | | Descriptor: | CHLORIDE ION, POTASSIUM ION, RNA-DIRECTED RNA POLYMERASE | | Authors: | Graham, S.C, Sarin, L.P, Bahar, M.W, Myers, R.A, Stuart, D.I, Bamford, D.H, Grimes, J.M. | | Deposit date: | 2011-05-11 | | Release date: | 2011-07-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The N-Terminus of the RNA Polymerase from Infectious Pancreatic Necrosis Virus is the Determinant of Genome Attachment.
Plos Pathog., 7, 2011
|
|
2CDF
 
 | | Structure and binding kinetics of three different human CD1d-alpha- Galactosylceramide-specific T cell receptors (TCR 5E) | | Descriptor: | TCR 5E | | Authors: | Gadola, S.D, Koch, M, Marles-Wright, J, Lissin, N.M, Sheperd, D, Matulis, G, Harlos, K, Villiger, P.M, Stuart, D.I, Jakobsen, B.K, Cerundolo, V, Jones, E.Y. | | Deposit date: | 2006-01-23 | | Release date: | 2006-03-07 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structrue and Binding Kinetics of Three Different Human Cd1D-Alpha-Galactosylceramide-Specific T Cell Receptors
J.Exp.Med., 203, 2006
|
|
2CME
 
 | | The crystal structure of SARS coronavirus ORF-9b protein | | Descriptor: | DECANE, HYPOTHETICAL PROTEIN 5 | | Authors: | Meier, C, Aricescu, A.R, Assenberg, R, Aplin, R.T, Gilbert, R.J.C, Grimes, J.M, Stuart, D.I. | | Deposit date: | 2006-05-06 | | Release date: | 2006-07-19 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The Crystal Structure of Orf-9B, a Lipid Binding Protein from the Sars Coronavirus.
Structure, 14, 2006
|
|
2YGC
 
 | |
9F59
 
 | | Poliovirus type 2 (strain MEF-1) stabilised virus-like particle (PV2 SC6b) from a mammalian expression system. | | Descriptor: | Capsid protein VP0, Capsid protein VP1, Capsid protein VP3, ... | | Authors: | Bahar, M.W, Porta, C, Fry, E.E, Stuart, D.I. | | Deposit date: | 2024-04-28 | | Release date: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (2.3 Å) | | Cite: | Recombinant expression systems for production of stabilised virus-like particles as next-generation polio vaccines.
Nat Commun, 16, 2025
|
|
9F5P
 
 | | Poliovirus type 2 (strain MEF-1) stabilised virus-like particle (PV2 SC6b) from an insect cell expression system. | | Descriptor: | Capsid protein VP0, Capsid protein VP1, Capsid protein VP3, ... | | Authors: | Bahar, M.W, Porta, C, Fry, E.E, Stuart, D.I. | | Deposit date: | 2024-04-29 | | Release date: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Recombinant expression systems for production of stabilised virus-like particles as next-generation polio vaccines.
Nat Commun, 16, 2025
|
|
9EZ0
 
 | | Poliovirus type 1 (strain Mahoney) expanded conformation stabilised virus-like particle (PV1 SC6b) from a yeast expression system. | | Descriptor: | Capsid protein VP0, Capsid protein VP1, Capsid protein VP3 | | Authors: | Bahar, M.W, Sherry, L, Stonehouse, N.J, Rowlands, D.J, Fry, E.E, Stuart, D.I. | | Deposit date: | 2024-04-10 | | Release date: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Recombinant expression systems for production of stabilised virus-like particles as next-generation polio vaccines.
Nat Commun, 16, 2025
|
|
9EYY
 
 | | Poliovirus type 1 (strain Mahoney) native conformation stabilised virus-like particle (PV1 SC6b) from a yeast expression system. | | Descriptor: | Capsid protein VP0, Capsid protein VP1, Capsid protein VP3, ... | | Authors: | Bahar, M.W, Sherry, L, Stonehouse, N.J, Rowlands, D.J, Fry, E.E, Stuart, D.I. | | Deposit date: | 2024-04-09 | | Release date: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Recombinant expression systems for production of stabilised virus-like particles as next-generation polio vaccines.
Nat Commun, 16, 2025
|
|
9F0K
 
 | | Poliovirus type 1 (strain Mahoney) expanded conformation stabilised virus-like particle (PV1 SC6b) from a mammalian expression system | | Descriptor: | Capsid protein VP0, Capsid protein VP1, Capsid protein VP3 | | Authors: | Bahar, M.W, Porta, C, Fry, E.E, Stuart, D.I. | | Deposit date: | 2024-04-16 | | Release date: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Recombinant expression systems for production of stabilised virus-like particles as next-generation polio vaccines.
Nat Commun, 16, 2025
|
|
9G6V
 
 | | Dissociated FMDV SAT2 Pentamer in complex with ultralong Fab117 | | Descriptor: | Fab117 Heavy Chain, Genome polyprotein | | Authors: | Clarke, J.D, Duyvesteyn, H.M.E, Ren, J, Fry, E.E, Owens, R.J, Stuart, D.I, Hammond, J.A. | | Deposit date: | 2024-07-19 | | Release date: | 2024-10-30 | | Last modified: | 2024-11-20 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | A broadly reactive ultralong bovine antibody that can determine the integrity of foot-and-mouth disease virus capsids.
J.Gen.Virol., 105, 2024
|
|
9H94
 
 | | Poliovirus type 2 (strain MEF-1) stabilised virus-like particle (PV2 SC5a) from a yeast expression system. | | Descriptor: | Capsid protein VP0, Capsid protein VP1, Capsid protein VP3, ... | | Authors: | Bahar, M.W, Sherry, L, Stonehouse, N.J, Rowlands, D.J, Fry, E.E, Stuart, D.I. | | Deposit date: | 2024-10-29 | | Release date: | 2025-04-02 | | Last modified: | 2025-04-09 | | Method: | ELECTRON MICROSCOPY (2.1 Å) | | Cite: | Production of an immunogenic trivalent poliovirus virus-like particle vaccine candidate in yeast using controlled fermentation.
Npj Vaccines, 10, 2025
|
|
9H93
 
 | | Poliovirus type 2 (strain MEF-1) stabilised virus-like particle (PV2 SC6b) from a yeast expression system. | | Descriptor: | Capsid protein VP1, Capsid protein VP3, Capsid protein, ... | | Authors: | Bahar, M.W, Sherry, L, Stonehouse, N.J, Rowlands, D.J, Fry, E.E, Stuart, D.I. | | Deposit date: | 2024-10-29 | | Release date: | 2025-04-02 | | Last modified: | 2025-04-09 | | Method: | ELECTRON MICROSCOPY (2.4 Å) | | Cite: | Production of an immunogenic trivalent poliovirus virus-like particle vaccine candidate in yeast using controlled fermentation.
Npj Vaccines, 10, 2025
|
|
9F3Q
 
 | | Poliovirus type 1 (strain Mahoney) stabilised virus-like particle (PV1 SC6b) in complex with GPP3 and GSH. | | Descriptor: | 1-[(3S)-5-[4-[(E)-ETHOXYIMINOMETHYL]PHENOXY]-3-METHYL-PENTYL]-3-PYRIDIN-4-YL-IMIDAZOLIDIN-2-ONE, Capsid protein VP0, Capsid protein VP1, ... | | Authors: | Bahar, M.W, Nasta, V, Sherry, L, Stonehouse, N.J, Rowlands, D.J, Fry, E.E, Stuart, D.I. | | Deposit date: | 2024-04-25 | | Release date: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (2.75 Å) | | Cite: | Recombinant expression systems for production of stabilised virus-like particles as next-generation polio vaccines.
Nat Commun, 16, 2025
|
|
8BBO
 
 | | SARS-CoV-2 Delta-RBD complexed with BA.2-36 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, IGH@ protein, Immunoglobulin kappa light chain, ... | | Authors: | Zhou, D, Ren, J, Stuart, D.I. | | Deposit date: | 2022-10-14 | | Release date: | 2023-03-22 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Rapid escape of new SARS-CoV-2 Omicron variants from BA.2-directed antibody responses.
Cell Rep, 42, 2023
|
|
8BBN
 
 | | SARS-CoV-2 Delta-RBD complexed with BA.2-10 and EY6A Fabs | | Descriptor: | BA.2-10 heavy chain, BA.2-10 light chain, EY6A Heavy chain, ... | | Authors: | Zhou, D, Ren, J, Stuart, D.I. | | Deposit date: | 2022-10-14 | | Release date: | 2023-03-22 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (3.58 Å) | | Cite: | Rapid escape of new SARS-CoV-2 Omicron variants from BA.2-directed antibody responses.
Cell Rep, 42, 2023
|
|
8BCZ
 
 | | SARS-CoV-2 Delta-RBD complexed with Fabs BA.2-36, BA.2-23, EY6A and COVOX-45 | | Descriptor: | BA.2-23 heavy chain, BA.2-23 light chain, BA.2-36 heavy chain, ... | | Authors: | Duyvesteyn, H.M.E, Ren, J, Stuart, D.I. | | Deposit date: | 2022-10-17 | | Release date: | 2023-03-22 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Rapid escape of new SARS-CoV-2 Omicron variants from BA.2-directed antibody responses.
Cell Rep, 42, 2023
|
|
