2FOR
 
 | | Crystal Structure of the Shigella flexneri Farnesyl Pyrophosphate Synthase Complex with an Isopentenyl Pyrophosphate | | Descriptor: | Geranyltranstransferase, ISOPENTYL PYROPHOSPHATE, PHOSPHATE ION | | Authors: | Minasov, G, Brunzelle, J.S, Shuvalova, L, Collart, F.R, Joachimiak, A, Anderson, W.F, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-01-13 | | Release date: | 2006-01-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of the Shigella flexneri Farnesyl Pyrophosphate Synthase Complex with an Isopentenyl Pyrophosphate
To be Published
|
|
2FZV
 
 | | Crystal Structure of an apo form of a Flavin-binding Protein from Shigella flexneri | | Descriptor: | CALCIUM ION, CHLORIDE ION, putative arsenical resistance protein | | Authors: | Vorontsov, I.I, Minasov, G, Brunzelle, J.S, Shuvalova, L, Collart, F.R, Joachimiak, A, Anderson, W.F, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-02-10 | | Release date: | 2006-02-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of an apo form of Shigella flexneri ArsH protein with an NADPH-dependent FMN reductase activity
Protein Sci., 16, 2007
|
|
2D7F
 
 | | Crystal structure of A lectin from canavalia gladiata seeds complexed with alpha-methyl-mannoside and alpha-aminobutyric acid | | Descriptor: | CALCIUM ION, Concanavalin A, D-ALPHA-AMINOBUTYRIC ACID, ... | | Authors: | Delatorre, P, Rocha, B.A.M, Souza, E.P, Freitas, B.T, Moreno, F.B.B.M, Sampaio, A.H, Azevedo Jr, W.F, Cavada, B.S. | | Deposit date: | 2005-11-19 | | Release date: | 2006-12-19 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Structure of a lectin from Canavalia gladiata seeds: new structural insights for old molecules
Bmc Struct.Biol., 7, 2007
|
|
2G85
 
 | | Crystal structure of chorismate synthase from Mycobacterium tuberculosis at 2.22 angstrons of resolution | | Descriptor: | Chorismate synthase | | Authors: | Dias, M.V.B, dos Santos, B.B, Ely, F, Basso, L.A, Santos, D.S, de Azevedo Jr, W.F. | | Deposit date: | 2006-03-01 | | Release date: | 2007-03-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Crystal structure of chorismate synthase from Mycobacterium tuberculosis at 2.22 angstron of resolution
To be Published
|
|
2CWM
 
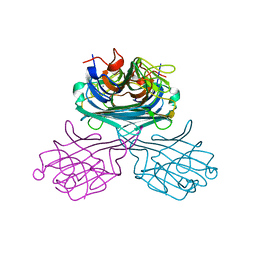 | | Native Crystal Structure of NO releasing inductive lectin from seeds of the Canavalia maritima (ConM) | | Descriptor: | CALCIUM ION, MANGANESE (II) ION, lectin | | Authors: | Cavada, B.S, De Azevedo Jr, W.F, Assreuy, A.M.S, Criddle, D.N, Gadelha, C.A.A, Delatorre, P, Souza, E.P, Rocha, B.A.M, Santi-Gadelha, T, Moreno, F.B.M.B. | | Deposit date: | 2005-06-22 | | Release date: | 2006-01-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Native crystal structure of a nitric oxide-releasing lectin from the seeds of Canavalia maritima
J.Struct.Biol., 152, 2005
|
|
2GJV
 
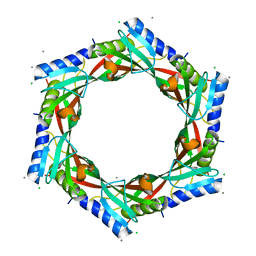 | | Crystal Structure of a Protein of Unknown Function from Salmonella typhimurium | | Descriptor: | CALCIUM ION, CHLORIDE ION, putative cytoplasmic protein | | Authors: | Minasov, G, Shuvalova, L, Brunzelle, J.S, Vorontsov, I.I, Collart, F.R, Joachimiak, A, Anderson, W.F, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-03-31 | | Release date: | 2006-04-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Crystal structure of a Hypothetical Protein from Salmonella typhimurium
To be Published
|
|
2GSJ
 
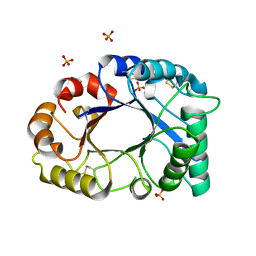 | | cDNA cloning and 1.75A crystal structure determination of PPL2, a novel chimerolectin from Parkia platycephala seeds exhibiting endochitinolytic activity | | Descriptor: | SULFATE ION, protein PPL-2 | | Authors: | Cavada, B.S, Moreno, F.B, da Rocha, B.A, de Azevedo Jr, W.F, Castellon, R.E.R, Goersch, G.V, Nagano, C.S, de Souza, E.P, Nascimento, K.S, Radis-Baptista, G, Delatorre, P, Leroy, Y, Toyama, M.H, Pinto, V.P, Sampaio, A.H, Barettino, D, Debray, H, Calvete, J.J, Sanz, L. | | Deposit date: | 2006-04-26 | | Release date: | 2007-03-13 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | cDNA cloning and 1.75 A crystal structure determination of PPL2, an endochitinase and N-acetylglucosamine-binding hemagglutinin from Parkia platycephala seeds
Febs J., 273, 2006
|
|
1PIW
 
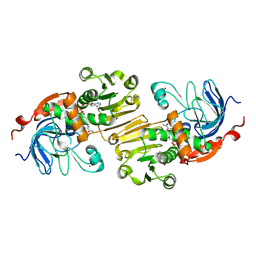 | | APO AND HOLO STRUCTURES OF AN NADP(H)-DEPENDENT CINNAMYL ALCOHOL DEHYDROGENASE FROM SACCHAROMYCES CEREVISIAE | | Descriptor: | Hypothetical zinc-type alcohol dehydrogenase-like protein in PRE5-FET4 intergenic region, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ZINC ION | | Authors: | Valencia, E, Larroy, C, Ochoa, W.F, Pares, X, Fita, I, Biosca, J.A. | | Deposit date: | 2003-05-30 | | Release date: | 2004-08-10 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Apo and Holo Structures of an NADP(H)-dependent Cinnamyl Alcohol Dehydrogenase from Saccharomyces cerevisiae
J.Mol.Biol., 341, 2004
|
|
1PV0
 
 | | Structure of the Sda antikinase | | Descriptor: | Sda | | Authors: | Rowland, S.L, Burkholder, W.F, Maciejewski, M.W, Grossman, A.D, King, G.F. | | Deposit date: | 2003-06-26 | | Release date: | 2004-04-13 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure and mechanism of Sda: an inhibitor of the histidine kinases that regulate initiation of sporulation in Bacillus subtilis
Mol.Cell, 13, 2004
|
|
7OAR
 
 | | Crystal structure of helicase Pif1 from Thermus oshimai in complex with parallel G-quadruplex | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, DNA (28-MER), MAGNESIUM ION, ... | | Authors: | Dai, Y.X, Liu, N.N, Guo, H.L, Chen, W.F, Rety, S, Xi, X.G. | | Deposit date: | 2021-04-20 | | Release date: | 2022-03-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.58 Å) | | Cite: | Structural mechanism underpinning Thermus oshimai Pif1-mediated G-quadruplex unfolding.
Embo Rep., 23, 2022
|
|
1AF9
 
 | | TETANUS NEUROTOXIN C FRAGMENT | | Descriptor: | TETANUS NEUROTOXIN | | Authors: | Umland, T.C, Wingert, L, Swaminathan, S, Furey, W.F, Schmidt, J.J, Sax, M. | | Deposit date: | 1997-03-24 | | Release date: | 1998-04-29 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of the receptor binding fragment HC of tetanus neurotoxin.
Nat.Struct.Biol., 4, 1997
|
|
1NLN
 
 | | CRYSTAL STRUCTURE OF HUMAN ADENOVIRUS 2 PROTEINASE WITH ITS 11 AMINO ACID COFACTOR AT 1.6 ANGSTROM RESOLUTION | | Descriptor: | ACETIC ACID, Adenain, PVIC | | Authors: | McGrath, W.J, Ding, J, Sweet, R.M, Mangel, W.F. | | Deposit date: | 2003-01-07 | | Release date: | 2003-08-26 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystallographic structure at 1.6-A resolution of the human adenovirus proteinase in a covalent complex with its 11-amino-acid peptide cofactor: insights on a new fold
Biochim.Biophys.Acta, 1648, 2003
|
|
1P0C
 
 | | Crystal Structure of the NADP(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8) | | Descriptor: | GLYCEROL, NADP-dependent ALCOHOL DEHYDROGENASE, PHOSPHATE ION, ... | | Authors: | Rosell, A, Valencia, E, Pares, X, Fita, I, Farres, J, Ochoa, W.F. | | Deposit date: | 2003-04-10 | | Release date: | 2003-04-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of the Vertebrate NADP(H)-dependent Alcohol Dehydrogenase (ADH8)
J.Mol.Biol., 330, 2003
|
|
1QTX
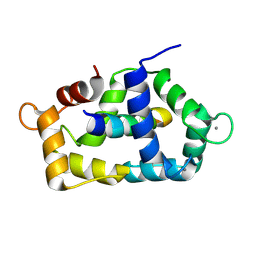 
 | |
1P0F
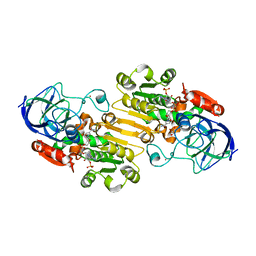 
 | | Crystal Structure of the Binary Complex: NADP(H)-Dependent Vertebrate Alcohol Dehydrogenase (ADH8) with the cofactor NADP | | Descriptor: | GLYCEROL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NADP-dependent ALCOHOL DEHYDROGENASE, ... | | Authors: | Rosell, A, Valencia, E, Pares, X, Fita, I, Farres, J, Ochoa, W.F. | | Deposit date: | 2003-04-10 | | Release date: | 2003-04-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of the vertebrate NADP(H)-dependent alcohol dehydrogenase (ADH8)
J.Mol.Biol., 330, 2003
|
|
1QS7
 
 | |
1QP6
 
 | | SOLUTION STRUCTURE OF ALPHA2D | | Descriptor: | PROTEIN (ALPHA2D) | | Authors: | Hill, R.B, DeGrado, W.F. | | Deposit date: | 1999-06-01 | | Release date: | 1999-06-09 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of Alpha2D, A Nativelike De Novo Designed Protein
J.Am.Chem.Soc., 120, 1998
|
|
1Q1N
 | | APO AND HOLO STRUCTURES OF AN NADP(H)-DEPENDENT CINNAMYL ALCOHOL DEHYDROGENASE FROM SACCHAROMYCES CEREVISIAE | | Descriptor: | Hypothetical zinc-type alcohol dehydrogenase-like protein in PRE5-FET4 intergenic region, ZINC ION | | Authors: | Valencia, E, Larroy, C, Ochoa, W.F, Pares, X, Fita, I, Biosca, J.A. | | Deposit date: | 2003-07-22 | | Release date: | 2004-08-03 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Apo and Holo Structures of an NADP(H)-dependent Cinnamyl Alcohol Dehydrogenase from Saccharomyces cerevisiae
J.Mol.Biol., 341, 2004
|
|
1PS0
 | | Crystal Structure of the NADP(H)-Dependent Cinnamyl Alcohol Dehydrogenase from Saccharomyces cerevisiae | | Descriptor: | Hypothetical zinc-type alcohol dehydrogenase-like protein in PRE5-FET4 intergenic region, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ZINC ION | | Authors: | Valencia, E, Larroy, C, Ochoa, W.F, Pares, X, Fita, I, Biosca, J.A. | | Deposit date: | 2003-06-20 | | Release date: | 2004-06-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | Apo and Holo Structures of an NADP(H)-dependent Cinnamyl Alcohol Dehydrogenase from Saccharomyces cerevisiae
J.Mol.Biol., 341, 2004
|
|
1AL1
 
 | | CRYSTAL STRUCTURE OF ALPHA1: IMPLICATIONS FOR PROTEIN DESIGN | | Descriptor: | ALPHA HELIX PEPTIDE: ELLKKLLEELKG, SULFATE ION | | Authors: | Hill, C.P, Anderson, D.H, Wesson, L, Degrado, W.F, Eisenberg, D. | | Deposit date: | 1990-07-02 | | Release date: | 1991-10-15 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of alpha 1: implications for protein design.
Science, 249, 1990
|
|
1PF7
 | | CRYSTAL STRUCTURE OF HUMAN PNP COMPLEXED WITH IMMUCILLIN H | | Descriptor: | 1,4-DIDEOXY-4-AZA-1-(S)-(9-DEAZAHYPOXANTHIN-9-YL)-D-RIBITOL, PURINE NUCLEOSIDE PHOSPHORYLASE, SULFATE ION | | Authors: | De Azevedo Jr, W.F, Canduri, F, Dos Santos, D.M, Pereira, J.H, Dias, M.V.B, Silva, R.G, Mendes, M.A, Palma, M.S, Basso, L.A, Santos, D.S. | | Deposit date: | 2003-05-24 | | Release date: | 2004-06-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural basis for inhibition of human PNP by immucillin-H
Biochem.Biophys.Res.Commun., 309, 2003
|
|
1CE4
 
 | |
1COI
 | | DESIGNED TRIMERIC COILED COIL-VALD | | Descriptor: | COIL-VALD, SULFATE ION | | Authors: | Ogihara, N.L, Weiss, M.S, Degrado, W.F, Eisenberg, D. | | Deposit date: | 1996-08-10 | | Release date: | 1997-02-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The crystal structure of the designed trimeric coiled coil coil-VaLd: implications for engineering crystals and supramolecular assemblies.
Protein Sci., 6, 1997
|
|
1DKZ
 
 | | THE SUBSTRATE BINDING DOMAIN OF DNAK IN COMPLEX WITH A SUBSTRATE PEPTIDE, DETERMINED FROM TYPE 1 NATIVE CRYSTALS | | Descriptor: | SUBSTRATE BINDING DOMAIN OF DNAK, SUBSTRATE PEPTIDE (7 RESIDUES) | | Authors: | Zhu, X, Zhao, X, Burkholder, W.F, Gragerov, A, Ogata, C.M, Gottesman, M.E, Hendrickson, W.A. | | Deposit date: | 1996-06-03 | | Release date: | 1996-12-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural analysis of substrate binding by the molecular chaperone DnaK.
Science, 272, 1996
|
|
1DUG
 
 | | STRUCTURE OF THE FIBRINOGEN G CHAIN INTEGRIN BINDING AND FACTOR XIIIA CROSSLINKING SITES OBTAINED THROUGH CARRIER PROTEIN DRIVEN CRYSTALLIZATION | | Descriptor: | GLUTATHIONE, chimera of GLUTATHIONE S-TRANSFERASE-synthetic LINKEr-C-TERMINAL FIBRINOGEN GAMMA CHAIN | | Authors: | Ware, S, Donahue, J.P, Hawiger, J, Anderson, W.F. | | Deposit date: | 2000-01-17 | | Release date: | 2000-02-02 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of the fibrinogen gamma-chain integrin binding and factor XIIIa cross-linking sites obtained through carrier protein driven crystallization.
Protein Sci., 8, 1999
|
|
