6A19
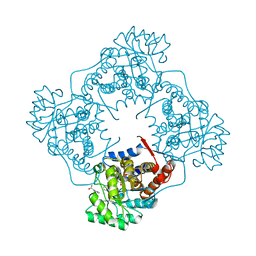 
 | | Mandelate oxidase mutant-Y128F with Benzoylformic acid | | Descriptor: | 4-hydroxymandelate oxidase, BENZOYL-FORMIC ACID, FLAVIN MONONUCLEOTIDE | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-06 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
5HGX
 
 | |
5HJE
 
 | |
7EUZ
 
 | | Structural and mechanistic studies of a novel non-heme iron epimerase/lyase and its utilization in chemoselective synthesis. | | Descriptor: | (2S,3R)-2-azanyl-3-(1H-indol-3-yl)butanoic acid, Cupin domain-containing protein, FE (III) ION, ... | | Authors: | Li, T.L, Li, Y.S, Chen, M.H. | | Deposit date: | 2021-05-19 | | Release date: | 2022-03-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.91028166 Å) | | Cite: | Structural and Mechanistic Bases for StnK3 and Its Mutant-Mediated Lewis-Acid-Dependent Epimerization and Retro-Aldol Reactions.
Acs Catalysis, 12, 2022
|
|
7EUE
 
 | |
7F6X
 
 | | Structural and mechanistic studies of a novel non-heme iron epimerase/lyase and its utilization in chemoselective synthesis. | | Descriptor: | 1H-INDOLE-3-CARBALDEHYDE, Cupin domain-containing protein, FE (III) ION, ... | | Authors: | Li, T.L, Li, Y.S, Chen, M.H. | | Deposit date: | 2021-06-26 | | Release date: | 2022-03-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Structural and Mechanistic Bases for StnK3 and Its Mutant-MediatedLewis-Acid-Dependent Epimerization and Retro-Aldol Reactions.
Acs Catalysis, 12, 2022
|
|
7EUP
 
 | | Structural and mechanistic studies of a novel non-heme iron epimerase/lyase and its utilization in chemoselective synthesis. | | Descriptor: | (2S,3R)-2-azanyl-3-phenyl-butanoic acid, Cupin domain-containing protein, FE (III) ION | | Authors: | Li, T.L, Li, Y.S, Chen, M.H. | | Deposit date: | 2021-05-18 | | Release date: | 2022-03-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.11043429 Å) | | Cite: | Structural and Mechanistic Bases for StnK3 and Its Mutant-Mediated Lewis-Acid-Dependent Epimerization and Retro-Aldol Reactions.
Acs Catalysis, 12, 2022
|
|
7EQK
 
 | | Structural and mechanistic studies of a novel non-heme iron epimerase/lyase and its utilization in chemoselective synthesis. | | Descriptor: | (E)-3-(1H-indol-3-yl)-2-oxidanyl-but-2-enoic acid, 1-(1~{H}-indol-3-yl)ethanone, Cupin domain-containing protein, ... | | Authors: | Li, T.L, Li, Y.S, Chen, M.H. | | Deposit date: | 2021-05-03 | | Release date: | 2022-03-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.04001021 Å) | | Cite: | Structural and Mechanistic Bases for StnK3 and Its Mutant-Mediated Lewis-Acid-Dependent Epimerization and Retro-Aldol Reactions.
Acs Catalysis, 12, 2022
|
|
7EU6
 
 | |
6A01
 
 | | The crystal structure of Mandelate oxidase Y128F with 3,3-difluoro-2,2-dihydroxy-3-phenylpropionic acid | | Descriptor: | 3,3-difluoro-2,2-dihydroxy-3-phenylpropanoic acid, 4-hydroxymandelate oxidase, FLAVIN MONONUCLEOTIDE | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-05 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.868 Å) | | Cite: | Structural and chemical trapping of flavin-oxide intermediates reveals substrate-directed reaction multiplicity.
Protein Sci., 29, 2020
|
|
5WQE
 
 | |
6M2O
 
 | |
6M2U
 
 | |
7DWC
 
 | | Bacteroides thetaiotaomicron VPI5482 BTAxe1 | | Descriptor: | Xylanase | | Authors: | Wang, L.Y, Wang, Y.L, Xin, F.J, Sun, L.C. | | Deposit date: | 2021-01-17 | | Release date: | 2022-01-19 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.804 Å) | | Cite: | Rational Design for Broadened Substrate Specificity and Enhanced Activity of a Novel Acetyl Xylan Esterase from Bacteroides thetaiotaomicron.
J.Agric.Food Chem., 69, 2021
|
|
6A23
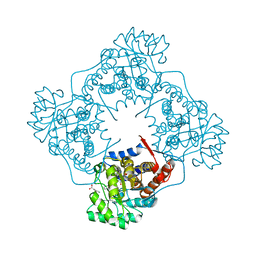 
 | | Mandelate oxidase mutant-Y128F with the N5-benzyl-FMN adduct | | Descriptor: | 1-[5-(benzenecarbonyl)-7,8-dimethyl-2,4-dioxo-1,3,4,5-tetrahydrobenzo[g]pteridin-10(2H)-yl]-1-deoxy-5-O-phosphono-D-ribitol, 4-hydroxymandelate oxidase, BENZOYL-FORMIC ACID, ... | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-08 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A1A
 
 | | Mandelate oxidase mutant-Y128F with 4-hydroxymandelic acid | | Descriptor: | (2S)-hydroxy(4-hydroxyphenyl)ethanoic acid, 4-hydroxymandelate oxidase, FLAVIN MONONUCLEOTIDE | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-07 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Biochemical and structural explorations of alpha-hydroxyacid oxidases reveal a four-electron oxidative decarboxylation reaction.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
5ZZZ
 
 | | The crystal structure of Mandelate oxidase Y128C with benzoyl-formic acid | | Descriptor: | 4-hydroxymandelate oxidase, BENZOYL-FORMIC ACID, FLAVIN MONONUCLEOTIDE | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-05 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.446 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A21
 
 | | Mandelate oxidase mutant-Y128F with the N5-malonyl-FMN adduct | | Descriptor: | 3-[7,8-dimethyl-2,4-bis(oxidanylidene)-10-[(2S,3S,4R)-2,3,4-tris(oxidanyl)-5-phosphonooxy-pentyl]-1H-benzo[g]pteridin-5-yl]-3-oxidanylidene-propanoic acid, 4-hydroxymandelate oxidase | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-08 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.501 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
5ZZQ
 
 | | The crystal structure of Mandelate oxidase with (S)-4-Hydroxymandelic acid | | Descriptor: | (2S)-hydroxy(4-hydroxyphenyl)ethanoic acid, 4-hydroxymandelate oxidase, FLAVIN MONONUCLEOTIDE | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-04 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Biochemical and structural explorations of alpha-hydroxyacid oxidases reveal a four-electron oxidative decarboxylation reaction.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A11
 
 | | Mandelate oxidase mutant-Y128F with phenylpyruvic acid | | Descriptor: | 3-PHENYLPYRUVIC ACID, 4-hydroxymandelate oxidase, FLAVIN MONONUCLEOTIDE | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-06 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Biochemical and structural explorations of alpha-hydroxyacid oxidases reveal a four-electron oxidative decarboxylation reaction.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A1M
 
 | | Mandelate oxidase mutant-Y128F with 5-deazariboflavin mononucleotide and benzoylformic acid | | Descriptor: | 1-deoxy-1-(7,8-dimethyl-2,4-dioxo-3,4-dihydropyrimido[4,5-b]quinolin-10(2H)-yl)-5-O-phosphono-D-ribitol, 4-hydroxymandelate oxidase, BENZOYL-FORMIC ACID, ... | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-07 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
5ZZX
 
 | |
6A00
 
 | | The crystal structure of Mandelate oxidase with (S)-2-phenylpropionate | | Descriptor: | (2~{S})-2-phenylpropanoic acid, 4-hydroxymandelate oxidase, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-05 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Biochemical and structural explorations of alpha-hydroxyacid oxidases reveal a four-electron oxidative decarboxylation reaction.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A1P
 
 | | Mandelate oxidase mutant-Y128F with 5-deazariboflavin mononucleotide and phenylpyruvic acid | | Descriptor: | 1-deoxy-1-(7,8-dimethyl-2,4-dioxo-3,4-dihydropyrimido[4,5-b]quinolin-10(2H)-yl)-5-O-phosphono-D-ribitol, 3-PHENYLPYRUVIC ACID, 4-hydroxymandelate oxidase, ... | | Authors: | Li, T.L, Lin, K.H. | | Deposit date: | 2018-06-07 | | Release date: | 2019-06-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.51 Å) | | Cite: | The flavin mononucleotide cofactor in alpha-hydroxyacid oxidases exerts its electrophilic/nucleophilic duality in control of the substrate-oxidation level.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
6A0D
 
 | |
