3MVW
 
 | | X-ray structure of a "NikA+Iron complex" hybrid, NikA/1 | | Descriptor: | 2-[2-[carboxymethyl(phenylmethyl)amino]ethyl-[(2-hydroxyphenyl)methyl]amino]ethanoic acid, ACETATE ION, CHLORIDE ION, ... | | Authors: | Cavazza, C, Bochot, C, Rousselot-Pailley, P, Carpentier, P, Cherrier, M.V, Martin, L, Marchi-Delapierre, C, Fontecilla-Camps, J.C, Menage, S. | | Deposit date: | 2010-05-05 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Crystallographic snapshots of the reaction of aromatic C-H with O(2) catalysed by a protein-bound iron complex
NAT.CHEM., 2, 2010
|
|
3MVY
 
 | | X-ray structure of the diatomic oxo-intermediate NikA/1-Int', prior hydroxylation | | Descriptor: | 2-[2-[carboxymethyl(phenylmethyl)amino]ethyl-[(2-hydroxyphenyl)methyl]amino]ethanoic acid, ACETATE ION, CHLORIDE ION, ... | | Authors: | Cavazza, C, Bochot, C, Rousselot-Pailley, P, Carpentier, P, Cherrier, M.V, Martin, L, Marchi-Delapierre, C, Fontecilla-Camps, J.C, Menage, S. | | Deposit date: | 2010-05-05 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystallographic snapshots of the reaction of aromatic C-H with O(2) catalysed by a protein-bound iron complex
NAT.CHEM., 2, 2010
|
|
3MVZ
 
 | | X-ray structure of the (hydro)peroxo intermediate NikA/1-Int", after monohydroxylation of the iron complex | | Descriptor: | 2-[2-[carboxymethyl-[(2-hydroxyphenyl)methyl]amino]ethyl-[(2-hydroxyphenyl)methyl]amino]ethanoic acid, ACETATE ION, FE (III) ION, ... | | Authors: | Cavazza, C, Bochot, C, Rousselot-Pailley, P, Carpentier, P, Cherrier, M.V, Martin, L, Marchi-Delapierre, C, Fontecilla-Camps, J.C, Menage, S. | | Deposit date: | 2010-05-05 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystallographic snapshots of the reaction of aromatic C-H with O(2) catalysed by a protein-bound iron complex
NAT.CHEM., 2, 2010
|
|
3MVX
 
 | | X-ray structure of the reduced NikA/1 hybrid, NikA/1-Red | | Descriptor: | (2R,3S)-1,4-DIMERCAPTOBUTANE-2,3-DIOL, (2S,3S)-1,4-DIMERCAPTOBUTANE-2,3-DIOL, 2-[2-[carboxymethyl(phenylmethyl)amino]ethyl-[(2-hydroxyphenyl)methyl]amino]ethanoic acid, ... | | Authors: | Cavazza, C, Bochot, C, Rousselot-Pailley, P, Carpentier, P, Cherrier, M.V, Martin, L, Marchi-Delapierre, C, Fontecilla-Camps, J.C, Menage, S. | | Deposit date: | 2010-05-05 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystallographic snapshots of the reaction of aromatic C-H with O(2) catalysed by a protein-bound iron complex
NAT.CHEM., 2, 2010
|
|
3MZ9
 
 | | X-ray structure of NikA in complex with HBED | | Descriptor: | 2-[2-[carboxymethyl-[(2-hydroxyphenyl)methyl]amino]ethyl-[(2-hydroxyphenyl)methyl]amino]ethanoic acid, ACETATE ION, CHLORIDE ION, ... | | Authors: | Cavazza, C, Bochot, C, Rousselot-Pailley, P, Carpentier, P, Cherrier, M.V, Martin, L, Marchi-Delapierre, C, Fontecilla-Camps, J.C, Menage, S. | | Deposit date: | 2010-05-12 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystallographic snapshots of the reaction of aromatic C-H with O(2) catalysed by a protein-bound iron complex
NAT.CHEM., 2, 2010
|
|
3MW0
 
 | | X-ray structure of the doubly hydroxylated iron complex-NikA species, NikA1/O2 | | Descriptor: | 2-[2-[carboxymethyl-[(2-hydroxyphenyl)methyl]amino]ethyl-[(2,3-dihydroxyphenyl)methyl]amino]ethanoic acid, DITHIANE DIOL, FE (III) ION, ... | | Authors: | Cavazza, C, Bochot, C, Rousselot-Pailley, P, Carpentier, P, Cherrier, M.V, Martin, L, Marchi-Delapierre, C, Fontecilla-Camps, J.C, Menage, S. | | Deposit date: | 2010-05-05 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystallographic snapshots of the reaction of aromatic C-H with O(2) catalysed by a protein-bound iron complex
NAT.CHEM., 2, 2010
|
|
3MZB
 
 | | X-ray structure of NikA in complex with the doubly hydroxylated iron complex, 1-O2 | | Descriptor: | 2-[2-[carboxymethyl-[(2-hydroxyphenyl)methyl]amino]ethyl-[(2,3-dihydroxyphenyl)methyl]amino]ethanoic acid, ACETATE ION, CHLORIDE ION, ... | | Authors: | Cavazza, C, Bochot, C, Rousselot-Pailley, P, Carpentier, P, Cherrier, M.V, Martin, L, Marchi-Delapierre, C, Fontecilla-Camps, J.C, Menage, S. | | Deposit date: | 2010-05-12 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystallographic snapshots of the reaction of aromatic C-H with O(2) catalysed by a protein-bound iron complex
NAT.CHEM., 2, 2010
|
|
4R9T
 
 | | L-ficolin complexed to sulphates | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ACETATE ION, CALCIUM ION, ... | | Authors: | Laffly, E, Lacroix, M, Martin, L, Vassal-Stermann, E, Thielens, N, Gaboriaud, C. | | Deposit date: | 2014-09-08 | | Release date: | 2014-11-05 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Human ficolin-2 recognition versatility extended: An update on the binding of ficolin-2 to sulfated/phosphated carbohydrates.
Febs Lett., 588, 2014
|
|
4R9J
 | | L-ficolin complexed to glucosamine-6-sulfate | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-amino-2-deoxy-6-O-sulfo-alpha-D-glucopyranose, ACETATE ION, ... | | Authors: | Laffly, E, Lacroix, M, Martin, L, Vassal-Stermann, E, Thielens, N, Gaboriaud, C. | | Deposit date: | 2014-09-05 | | Release date: | 2014-11-05 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Human ficolin-2 recognition versatility extended: An update on the binding of ficolin-2 to sulfated/phosphated carbohydrates.
Febs Lett., 588, 2014
|
|
4UE2
 
 | | Structure of air-treated anaerobically purified D. fructosovorans NiFe-hydrogenase | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, FE3-S4 CLUSTER, GLYCEROL, ... | | Authors: | Volbeda, A, Martin, L, Liebgott, P.-P, Fontecilla-Camps, J.C. | | Deposit date: | 2014-12-15 | | Release date: | 2015-03-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | [Nife]-Hydrogenases Revisited: Nickel-Carboxamido Bond Formation in a Variant with Accrued O2-Tolerance and a Tentative Re-Interpretation of Ni-Si States.
Metallomics, 7, 2015
|
|
4UD2
 
 | | Structure of anaerobically purified D. fructosovorans NiFe- hydrogenase | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, FE3-S4 CLUSTER, GLYCEROL, ... | | Authors: | Volbeda, A, Martin, L, Liebgott, P.-P, Fontecilla-Camps, J.C. | | Deposit date: | 2014-12-05 | | Release date: | 2015-03-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | [Nife]-Hydrogenases Revisited: Nickel-Carboxamido Bond Formation in a Variant with Accrued O2-Tolerance and a Tentative Re-Interpretation of Ni-Si States.
Metallomics, 7, 2015
|
|
4UD6
 
 | | Structure of methylviologen-treated anaerobically purified D. fructosovorans NiFe-hydrogenase | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, FE3-S4 CLUSTER, GLYCEROL, ... | | Authors: | Volbeda, A, Martin, L, Liebgott, P.-P, Fontecilla-Camps, J.C. | | Deposit date: | 2014-12-08 | | Release date: | 2015-03-25 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | [NiFe]-hydrogenases revisited: nickel-carboxamido bond formation in a variant with accrued O2-tolerance and a tentative re-interpretation of Ni-SI states.
Metallomics, 7, 2015
|
|
4UEQ
 
 | | Structure of the V74C large subunit mutant of D. fructosovorans NiFe- hydrogenase | | Descriptor: | CALCIUM ION, CARBONATE ION, CARBONMONOXIDE-(DICYANO) IRON, ... | | Authors: | Volbeda, A, Martin, L, Liebgott, P.-P, Fontecilla-Camps, J.C. | | Deposit date: | 2014-12-18 | | Release date: | 2015-03-25 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | [NiFe]-hydrogenases revisited: nickel-carboxamido bond formation in a variant with accrued O2-tolerance and a tentative re-interpretation of Ni-SI states.
Metallomics, 7, 2015
|
|
4UE6
 
 | | Structure of methylene blue-treated anaerobically purified D. fructosovorans NiFe-hydrogenase | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, FE3-S4 CLUSTER, GLYCEROL, ... | | Authors: | Volbeda, A, Martin, L, Liebgott, P.-P, Fontecilla-Camps, J.C. | | Deposit date: | 2014-12-16 | | Release date: | 2015-03-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | [Nife]-Hydrogenases Revisited: Nickel-Carboxamido Bond Formation in a Variant with Accrued O2-Tolerance and a Tentative Re-Interpretation of Ni-Si States.
Metallomics, 7, 2015
|
|
4UEW
 
 | | Structure of H2-treated anaerobically purified D. fructosovorans NiFe- hydrogenase | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, FE3-S4 CLUSTER, GLYCEROL, ... | | Authors: | Volbeda, A, Martin, L, Liebgott, P.-P, Fontecilla-Camps, J.C. | | Deposit date: | 2014-12-20 | | Release date: | 2015-03-25 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | [NiFe]-hydrogenases revisited: nickel-carboxamido bond formation in a variant with accrued O2-tolerance and a tentative re-interpretation of Ni-SI states.
Metallomics, 7, 2015
|
|
4JY9
 
 | | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters | | Descriptor: | CHAPSO, CHLORIDE ION, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Nicolet, Y, Rohac, R, Martin, L, Fontecilla-Camps, J.C. | | Deposit date: | 2013-03-29 | | Release date: | 2013-05-01 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4JYF
 
 | | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters. | | Descriptor: | CARBONATE ION, CHAPSO, CHLORIDE ION, ... | | Authors: | Nicolet, Y, Rohac, R, Martin, L, Fontecilla-Camps, J.C. | | Deposit date: | 2013-03-29 | | Release date: | 2013-05-01 | | Last modified: | 2013-05-15 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4JYE
 
 | | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters. | | Descriptor: | BROMIDE ION, Biotin synthetase, putative, ... | | Authors: | Nicolet, Y, Rohac, R, Martin, L, Fontecilla-Camps, J.C. | | Deposit date: | 2013-03-29 | | Release date: | 2013-05-01 | | Last modified: | 2013-05-15 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4JYD
 
 | | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters. | | Descriptor: | 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, BROMIDE ION, CHAPSO, ... | | Authors: | Nicolet, Y, Rohac, R, Martin, L, Fontecilla-Camps, J.C. | | Deposit date: | 2013-03-29 | | Release date: | 2013-05-01 | | Last modified: | 2013-05-15 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4JY8
 
 | | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters | | Descriptor: | CHLORIDE ION, FEFE-HYDROGENASE MATURASE, HYDROSULFURIC ACID, ... | | Authors: | Nicolet, Y, Rohac, R, Martin, L, Fontecilla-Camps, J.C. | | Deposit date: | 2013-03-29 | | Release date: | 2013-05-01 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.901 Å) | | Cite: | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
6TN6
 
 | | X-ray structure of the endo-beta-1,4-mannanase from Thermotoga petrophila | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ... | | Authors: | da Silva, V.M, Squina, F.M, Sperenca, M, Martin, L, Muniz, J.R.C, Garcia, W, Nicolet, Y. | | Deposit date: | 2019-12-06 | | Release date: | 2020-04-29 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | High-resolution structure of a modular hyperthermostable endo-beta-1,4-mannanase from Thermotoga petrophila: The ancillary immunoglobulin-like module is a thermostabilizing domain.
Biochim Biophys Acta Proteins Proteom, 1868, 2020
|
|
4JXC
 
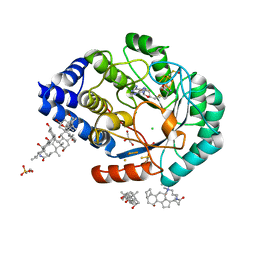 | | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters | | Descriptor: | CHAPSO, CHLORIDE ION, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Nicolet, Y, Rohac, R, Martin, L, Fontecilla-Camps, J.C. | | Deposit date: | 2013-03-28 | | Release date: | 2013-05-01 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | X-ray snapshots of possible intermediates in the time course of synthesis and degradation of protein-bound Fe4S4 clusters.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
1RHH
 
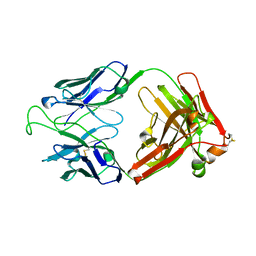 | | Crystal Structure of the Broadly HIV-1 Neutralizing Fab X5 at 1.90 Angstrom Resolution | | Descriptor: | Fab X5, heavy chain, light chain | | Authors: | Darbha, R, Phogat, S, Labrijn, A.F, Shu, Y, Gu, Y, Andrykovitch, M, Zhang, M.Y, Pantophlet, R, Martin, L, Vita, C, Burton, D.R, Dimitrov, D.S, Ji, X. | | Deposit date: | 2003-11-14 | | Release date: | 2004-02-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of the Broadly Cross-Reactive HIV-1-Neutralizing Fab X5 and Fine Mapping of Its Epitope
Biochemistry, 43, 2004
|
|
4NYT
 
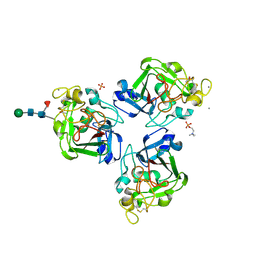 | | L-Ficolin Complexed to Phosphocholine | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETATE ION, CALCIUM ION, ... | | Authors: | Laffly, E, Gaboriaud, C, Martin, L, Thielens, N. | | Deposit date: | 2013-12-11 | | Release date: | 2014-10-29 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Human L-ficolin recognizes phosphocholine moieties of pneumococcal teichoic Acid
J.Immunol., 193, 2014
|
|
7KQP
 
 | | Crystal structure of SARS-CoV-2 NSP3 macrodomain in complex with ADP-ribose (P43 crystal form) | | Descriptor: | Non-structural protein 3, [(2R,3S,4R,5R)-5-(6-AMINOPURIN-9-YL)-3,4-DIHYDROXY-OXOLAN-2-YL]METHYL [HYDROXY-[[(2R,3S,4R,5S)-3,4,5-TRIHYDROXYOXOLAN-2-YL]METHOXY]PHOSPHORYL] HYDROGEN PHOSPHATE | | Authors: | Correy, G.J, Young, I.D, Thompson, M.C, Fraser, J.S. | | Deposit date: | 2020-11-17 | | Release date: | 2020-12-09 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (0.88 Å) | | Cite: | Fragment binding to the Nsp3 macrodomain of SARS-CoV-2 identified through crystallographic screening and computational docking.
Sci Adv, 7, 2021
|
|
