3FOI
 
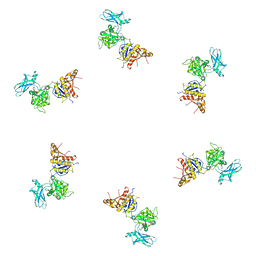 | | Fitting of gp18M crystal structure into 3D cryo-EM reconstruction of bacteriophage T4 contracted tail | | Descriptor: | Tail sheath protein Gp18 | | Authors: | Aksyuk, A.A, Leiman, P.G, Kurochkina, L.P, Shneider, M.M, Kostyuchenko, V.A, Mesyanzhinov, V.V, Rossmann, M.G. | | Deposit date: | 2008-12-30 | | Release date: | 2009-03-10 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (16 Å) | | Cite: | The tail sheath structure of bacteriophage T4: a molecular machine for infecting bacteria.
Embo J., 28, 2009
|
|
3FOA
 
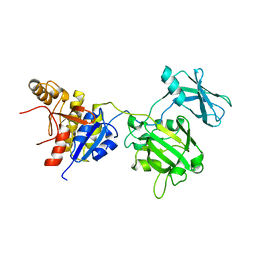 | | Crystal structure of the bacteriophage T4 tail sheath protein, deletion mutant gp18M | | Descriptor: | Tail sheath protein Gp18 | | Authors: | Aksyuk, A.A, Leiman, P.G, Kurochkina, L.P, Shneider, M.M, Kostyuchenko, V.A, Mesyanzhinov, V.V, Rossmann, M.G. | | Deposit date: | 2008-12-29 | | Release date: | 2009-03-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | The tail sheath structure of bacteriophage T4: a molecular machine for infecting bacteria.
Embo J., 28, 2009
|
|
3P0S
 
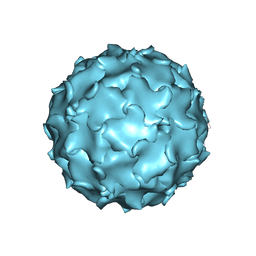 | |
3N9G
 
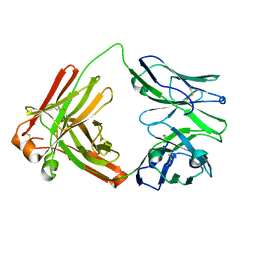 | |
1TJA
 
 | | Fitting of gp8, gp9, and gp11 into the cryo-EM reconstruction of the bacteriophage T4 contracted tail | | Descriptor: | Baseplate structural protein Gp11, Baseplate structural protein Gp8, Baseplate structural protein Gp9 | | Authors: | Leiman, P.G, Chipman, P.R, Kostyuchenko, V.A, Mesyanzhinov, V.V, Rossmann, M.G. | | Deposit date: | 2004-06-03 | | Release date: | 2004-08-31 | | Last modified: | 2024-02-14 | | Method: | ELECTRON MICROSCOPY (16 Å) | | Cite: | Three-dimensional rearrangement of proteins in the tail of bacteriophage t4 on infection of its host
Cell(Cambridge,Mass.), 118, 2004
|
|
3GQK
 
 | |
3GQ9
 
 | |
1QJU
 
 | | HUMAN RHINOVIRUS 16 COAT PROTEIN IN COMPLEX WITH ANTIVIRAL COMPOUND VP61209 | | Descriptor: | 2,6-DIMETHYL-1-(3-[3-METHYL-5-ISOXAZOLYL]-PROPANYL)-4-[2N-METHYL-2H-TETRAZOL-5-YL]-PHENOL, MYRISTIC ACID, PROTEIN VP1, ... | | Authors: | Hadfield, A.T, Minor, I, Diana, G.D, Rossmann, M.G. | | Deposit date: | 1999-07-05 | | Release date: | 1999-07-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Analysis of Three Structurally Related Antiviral Compounds in Complex with Human Rhinovirus 16
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
3FOH
 
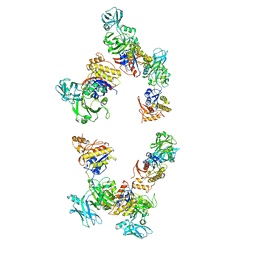 | | Fitting of gp18M crystal structure into 3D cryo-EM reconstruction of bacteriophage T4 extended tail | | Descriptor: | Tail sheath protein Gp18 | | Authors: | Aksyuk, A.A, Leiman, P.G, Kurochkina, L.P, Shneider, M.M, Kostyuchenko, V.A, Mesyanzhinov, V.V, Rossmann, M.G. | | Deposit date: | 2008-12-30 | | Release date: | 2009-03-10 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (15 Å) | | Cite: | The tail sheath structure of bacteriophage T4: a molecular machine for infecting bacteria.
Embo J., 28, 2009
|
|
3GQ8
 
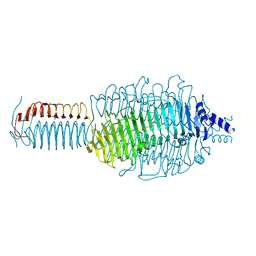 | |
3GQA
 
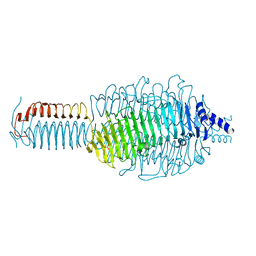 | |
3H3W
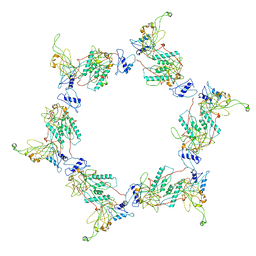 
 | | Fitting of the gp6 crystal structure into 3D cryo-EM reconstruction of bacteriophage T4 dome-shaped baseplate | | Descriptor: | Baseplate structural protein Gp6 | | Authors: | Aksyuk, A.A, Leiman, P.G, Shneider, M.M, Mesyanzhinov, V.V, Rossmann, M.G. | | Deposit date: | 2009-04-17 | | Release date: | 2009-05-19 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (12 Å) | | Cite: | The structure of gene product 6 of bacteriophage T4, the hinge-pin of the baseplate.
Structure, 17, 2009
|
|
3H3Y
 
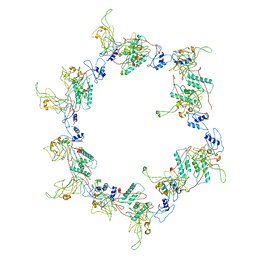 | | Fitting of the gp6 crystal structure into 3D cryo-EM reconstruction of bacteriophage T4 star-shaped baseplate | | Descriptor: | Baseplate structural protein Gp6 | | Authors: | Aksyuk, A.A, Leiman, P.G, Shneider, M.M, Mesyanzhinov, V.V, Rossmann, M.G. | | Deposit date: | 2009-04-17 | | Release date: | 2009-05-19 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (16 Å) | | Cite: | The structure of gene product 6 of bacteriophage T4, the hinge-pin of the baseplate.
Structure, 17, 2009
|
|
3N7X
 
 | |
3MUU
 
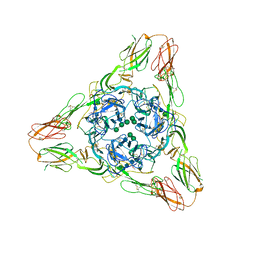 | | Crystal structure of the Sindbis virus E2-E1 heterodimer at low pH | | Descriptor: | Structural polyprotein, alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Li, L, Jose, J, Xiang, Y, Kuhn, R.J, Rossmann, M.G. | | Deposit date: | 2010-05-03 | | Release date: | 2010-11-24 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.29 Å) | | Cite: | Structural changes of envelope proteins during alphavirus fusion.
Nature, 468, 2010
|
|
3MUW
 
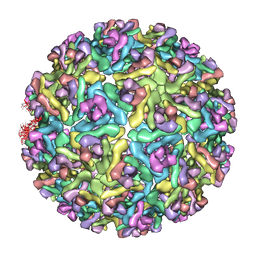 | | Pseudo-atomic structure of the E2-E1 protein shell in Sindbis virus | | Descriptor: | Structural polyprotein | | Authors: | Li, L, Jose, J, Xiang, Y, Kuhn, R.J, Rossmann, M.G. | | Deposit date: | 2010-05-03 | | Release date: | 2010-11-24 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (9 Å) | | Cite: | Structural changes of envelope proteins during alphavirus fusion.
Nature, 468, 2010
|
|
1TGE
 
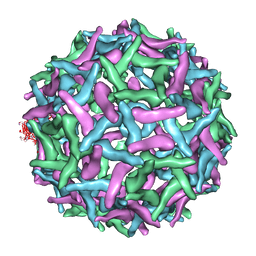 | | The structure of immature Dengue virus at 12.5 angstrom | | Descriptor: | envelope glycoprotein | | Authors: | Zhang, Y, Zhang, W, Ogata, S, Clements, D, Strauss, J.H, Baker, T.S, Kuhn, R.J, Rossmann, M.G. | | Deposit date: | 2004-05-28 | | Release date: | 2004-09-28 | | Last modified: | 2024-02-14 | | Method: | ELECTRON MICROSCOPY (12.5 Å) | | Cite: | Conformational changes of the flavivirus e glycoprotein.
Structure, 12, 2004
|
|
3GQH
 
 | |
1TG8
 
 | | The structure of Dengue virus E glycoprotein | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose, envelope glycoprotein | | Authors: | Zhang, Y, Zhang, W, Ogata, S, Clements, D, Strauss, J.H, Baker, T.S, Rossmann, M.G. | | Deposit date: | 2004-05-28 | | Release date: | 2004-09-28 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Conformational changes of the flavivirus e glycoprotein
Structure, 12, 2004
|
|
3EPD
 
 | | CryoEM structure of poliovirus receptor bound to poliovirus type 3 | | Descriptor: | MYRISTIC ACID, Poliovirus Type3 peptide, Poliovirus receptor, ... | | Authors: | Zhang, P, Mueller, S, Morais, M.C, Bator, C.M, Bowman, V.D, Hafenstein, S, Wimmer, E, Rossmann, M.G. | | Deposit date: | 2008-09-29 | | Release date: | 2008-11-11 | | Last modified: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (9 Å) | | Cite: | Crystal structure of CD155 and electron microscopic studies of its complexes with polioviruses.
Proc.Natl.Acad.Sci.USA, 105, 2008
|
|
3EPC
 
 | | CryoEM structure of poliovirus receptor bound to poliovirus type 1 | | Descriptor: | MYRISTIC ACID, Poliovirus receptor, Protein VP1, ... | | Authors: | Zhang, P, Mueller, S, Morais, M.C, Bator, C.M, Bowman, V.D, Hafenstein, S, Wimmer, E, Rossmann, M.G. | | Deposit date: | 2008-09-29 | | Release date: | 2008-11-11 | | Last modified: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (8 Å) | | Cite: | Crystal structure of CD155 and electron microscopic studies of its complexes with polioviruses.
Proc.Natl.Acad.Sci.USA, 105, 2008
|
|
3EPF
 
 | | CryoEM structure of poliovirus receptor bound to poliovirus type 2 | | Descriptor: | 1[2-CHLORO-4-METHOXY-PHENYL-OXYMETHYL]-4-[2,6-DICHLORO-PHENYL-OXYMETHYL]-BENZENE, MYRISTIC ACID, Poliovirus receptor, ... | | Authors: | Zhang, P, Mueller, S, Morais, M.C, Bator, C.M, Bowman, V.D, Hafenstein, S, Wimmer, E, Rossmann, M.G. | | Deposit date: | 2008-09-29 | | Release date: | 2008-11-11 | | Last modified: | 2024-04-17 | | Method: | ELECTRON MICROSCOPY (9 Å) | | Cite: | Crystal structure of CD155 and electron microscopic studies of its complexes with polioviruses.
Proc.Natl.Acad.Sci.USA, 105, 2008
|
|
3H2T
 
 | | Crystal structure of gene product 6, baseplate protein of bacteriophage T4 | | Descriptor: | Baseplate structural protein Gp6 | | Authors: | Aksyuk, A.A, Leiman, P.G, Shneider, M.M, Mesyanzhinov, V.V, Rossmann, M.G. | | Deposit date: | 2009-04-14 | | Release date: | 2009-05-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | The structure of gene product 6 of bacteriophage t4, the hinge-pin of the baseplate.
Structure, 17, 2009
|
|
3GQ7
 
 | |
1WTH
 
 | | Crystal structure of gp5-S351L mutant and gp27 complex | | Descriptor: | Baseplate structural protein Gp27, PHOSPHATE ION, POTASSIUM ION, ... | | Authors: | Kanamaru, S, Ishiwata, Y, Suzuki, T, Rossmann, M.G, Arisaka, F. | | Deposit date: | 2004-11-23 | | Release date: | 2005-03-08 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Control of bacteriophage t4 tail lysozyme activity during the infection process
J.Mol.Biol., 346, 2005
|
|
