8K3B
 
 | |
6K7Z
 
 | | Crystal structure of a GH18 chitinase from Pseudoalteromonas aurantia | | 分子名称: | GH18 chiitnase | | 著者 | Wang, Y.J, Li, P.Y, Cao, H.Y, Chen, X.L, Zhang, Y.Z. | | 登録日 | 2019-06-10 | | 公開日 | 2020-06-10 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.799 Å) | | 主引用文献 | Structural Insight Into Chitin Degradation and Thermostability of a Novel Endochitinase From the Glycoside Hydrolase Family 18.
Front Microbiol, 10, 2019
|
|
8J0I
 
 | | Aldo-keto reductase KmAKR | | 分子名称: | NADPH-dependent alpha-keto amide reductase, SODIUM ION | | 著者 | Xu, S.Y, Zhou, L, Xu, Y, Wang, Y.J, Zheng, Y.G. | | 登録日 | 2023-04-11 | | 公開日 | 2024-04-17 | | 実験手法 | X-RAY DIFFRACTION (1.91 Å) | | 主引用文献 | Aldo-keto reductase KmAKR from Kluyveromyces marxianus
To Be Published
|
|
8WJW
 
 | |
8WK9
 
 | |
8WK5
 
 | |
8WK7
 
 | |
8WKA
 
 | |
8IKU
 
 | |
7V9U
 
 | | Cryo-EM structure of E.coli retron-Ec86 (RT-msDNA-RNA) at 3.2 angstrom | | 分子名称: | DNA (105-MER), RNA (5'-R(P*CP*GP*UP*AP*AP*GP*GP*G)-3'), RNA (81-MER), ... | | 著者 | Wang, Y.J, Guan, Z.Y, Zou, T.T. | | 登録日 | 2021-08-26 | | 公開日 | 2022-08-31 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.12 Å) | | 主引用文献 | Cryo-EM structures of Escherichia coli Ec86 retron complexes reveal architecture and defence mechanism.
Nat Microbiol, 7, 2022
|
|
7XJG
 
 | | Cryo-EM structure of E.coli retron-Ec86 in complex with its effector at 2.5 angstrom | | 分子名称: | DNA (105-MER), MAGNESIUM ION, RNA (14-MER), ... | | 著者 | Wang, Y.J, Guan, Z.Y, Zou, T.T. | | 登録日 | 2022-04-17 | | 公開日 | 2022-09-14 | | 最終更新日 | 2024-07-03 | | 実験手法 | ELECTRON MICROSCOPY (2.51 Å) | | 主引用文献 | Cryo-EM structures of Escherichia coli Ec86 retron complexes reveal architecture and defence mechanism.
Nat Microbiol, 7, 2022
|
|
7V9X
 
 | |
8FBO
 
 | |
8FBJ
 
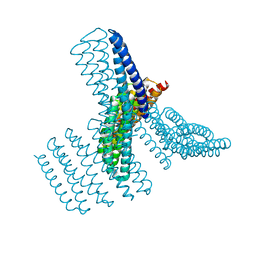 | |
8FBI
 
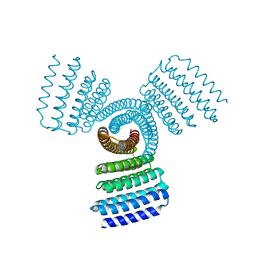 | |
8FBK
 
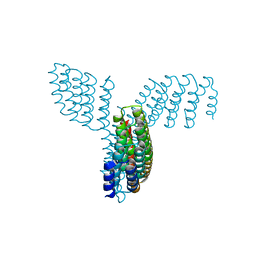 | |
8FBN
 
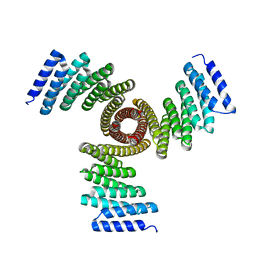 | |
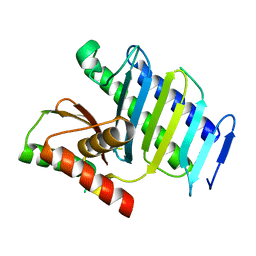
8GUO
 
 | |
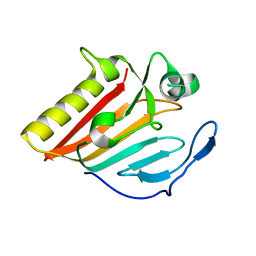
8GUN
 
 | |
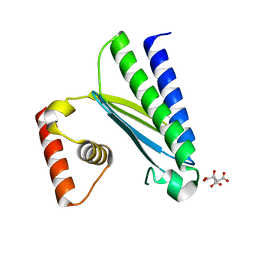
8GUP
 
 | |
7XY9
 
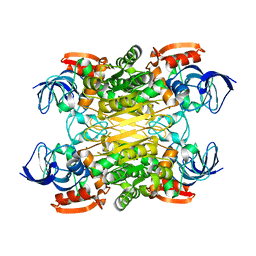 | | Cryo-EM structure of secondary alcohol dehydrogenases TbSADH after carrier-free immobilization based on weak intermolecular interactions | | 分子名称: | MAGNESIUM ION, NADP-dependent isopropanol dehydrogenase, ZINC ION | | 著者 | Chen, Q, Li, X, Yang, F, Qu, G, Sun, Z.T, Wang, Y.J. | | 登録日 | 2022-06-01 | | 公開日 | 2023-06-07 | | 最終更新日 | 2024-05-08 | | 実験手法 | ELECTRON MICROSCOPY (2.12 Å) | | 主引用文献 | Active and stable alcohol dehydrogenase-assembled hydrogels via synergistic bridging of triazoles and metal ions.
Nat Commun, 14, 2023
|
|
6J76
 
 | | Structure of 3,6-anhydro-L-galactose Dehydrogenase in Complex with NAP | | 分子名称: | Aldehyde dehydrogenase A, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | 著者 | Li, P.Y, Wang, Y, Chen, X.L, Zhang, Y.Z. | | 登録日 | 2019-01-17 | | 公開日 | 2020-01-22 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.368 Å) | | 主引用文献 | 3,6-Anhydro-L-Galactose Dehydrogenase VvAHGD is a Member of a New Aldehyde Dehydrogenase Family and Catalyzes by a Novel Mechanism with Conformational Switch of Two Catalytic Residues Cysteine 282 and Glutamate 248.
J.Mol.Biol., 432, 2020
|
|
6J75
 
 | | Structure of 3,6-anhydro-L-galactose Dehydrogenase | | 分子名称: | Aldehyde dehydrogenase A | | 著者 | Li, P.Y, Wang, Y, Chen, X.L, Zhang, Y.Z. | | 登録日 | 2019-01-17 | | 公開日 | 2020-01-22 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.695 Å) | | 主引用文献 | 3,6-Anhydro-L-Galactose Dehydrogenase VvAHGD is a Member of a New Aldehyde Dehydrogenase Family and Catalyzes by a Novel Mechanism with Conformational Switch of Two Catalytic Residues Cysteine 282 and Glutamate 248.
J.Mol.Biol., 432, 2020
|
|
6KZQ
 
 | | structure of PTP-MEG2 and NSF-pY83 peptide complex | | 分子名称: | NSF-pY83 peptide, Tyrosine-protein phosphatase non-receptor type 9 | | 著者 | Xu, Y.F, Chen, X, Yu, X, Sun, J.P. | | 登録日 | 2019-09-25 | | 公開日 | 2020-09-30 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | PTP-MEG2 regulates quantal size and fusion pore opening through two distinct structural bases and substrates.
Embo Rep., 22, 2021
|
|
6L03
 
 | | structure of PTP-MEG2 and MUNC18-1-pY145 peptide complex | | 分子名称: | Tyrosine-protein phosphatase non-receptor type 9, stxbp1-pY145 peptide | | 著者 | Xu, Y.F, Chen, X, Yu, X, Sun, J.P. | | 登録日 | 2019-09-25 | | 公開日 | 2020-09-30 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (2.084 Å) | | 主引用文献 | PTP-MEG2 regulates quantal size and fusion pore opening through two distinct structural bases and substrates.
Embo Rep., 22, 2021
|
|
