8ZEY
 
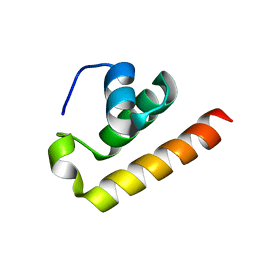 | |
1UM8
 
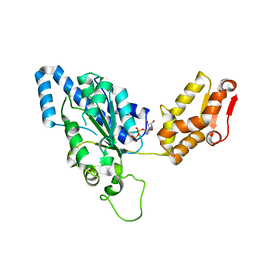 | | Crystal structure of helicobacter pylori ClpX | | 分子名称: | ADENOSINE-5'-DIPHOSPHATE, ATP-dependent Clp protease ATP-binding subunit clpX | | 著者 | Kim, D.Y, Kim, K.K. | | 登録日 | 2003-09-25 | | 公開日 | 2003-12-23 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Crystal Structure of ClpX Molecular Chaperone from Helicobacter pylori
J.Biol.Chem., 278, 2003
|
|
1L1J
 
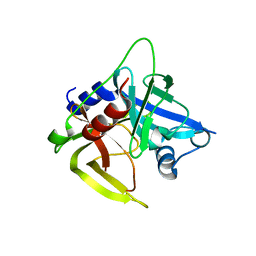 | | Crystal structure of the protease domain of an ATP-independent heat shock protease HtrA | | 分子名称: | heat shock protease HtrA | | 著者 | Kim, D.Y, Kim, D.R, Ha, S.C, Lokanath, N.K, Hwang, H.Y, Kim, K.K. | | 登録日 | 2002-02-18 | | 公開日 | 2003-04-01 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Crystal Structure of the Protease Domain of a Heat-shock Protein HtrA from Thermotoga maritima
J.BIOL.CHEM., 278, 2003
|
|
2P4B
 
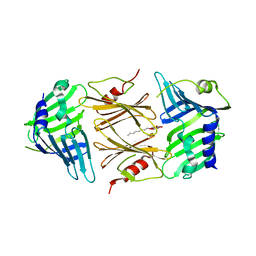 | | Crystal structure of E.coli RseB | | 分子名称: | Sigma-E factor regulatory protein rseB, octyl beta-D-glucopyranoside | | 著者 | Kim, D.Y, Kim, K.K. | | 登録日 | 2007-03-12 | | 公開日 | 2007-05-22 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Crystal structure of RseB and a model of its binding mode to RseA
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2ZL2
 
 | |
2ZL3
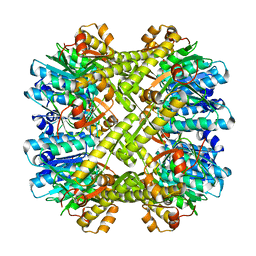 
 | | Crystal structure of H.pylori ClpP S99A | | 分子名称: | ATP-dependent Clp protease proteolytic subunit | | 著者 | Kim, D.Y, Kim, K.K. | | 登録日 | 2008-04-02 | | 公開日 | 2008-04-22 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (2.81 Å) | | 主引用文献 | The structural basis for the activation and peptide recognition of bacterial ClpP
J.Mol.Biol., 379, 2008
|
|
2ZL0
 
 | | Crystal structure of H.pylori ClpP | | 分子名称: | ATP-dependent Clp protease proteolytic subunit | | 著者 | Kim, D.Y, Kim, K.K. | | 登録日 | 2008-04-02 | | 公開日 | 2008-04-22 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | The structural basis for the activation and peptide recognition of bacterial ClpP
J.Mol.Biol., 379, 2008
|
|
2ZL4
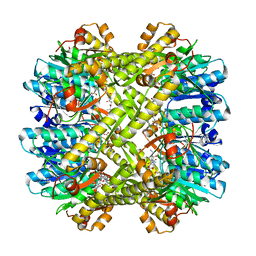 
 | |
3M4W
 
 | | Structural basis for the negative regulation of bacterial stress response by RseB | | 分子名称: | Sigma-E factor negative regulatory protein, Sigma-E factor regulatory protein rseB, ZINC ION | | 著者 | Kim, D.Y, Kwon, E, Choi, J.K, Hwang, H.-Y, Kim, K.K. | | 登録日 | 2010-03-12 | | 公開日 | 2010-05-05 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Structural basis for the negative regulation of bacterial stress response by RseB
Protein Sci., 19, 2010
|
|
8I2E
 
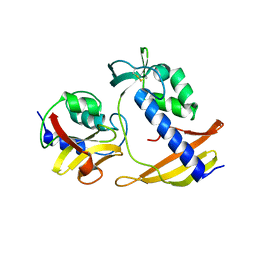 | |
8I2D
 
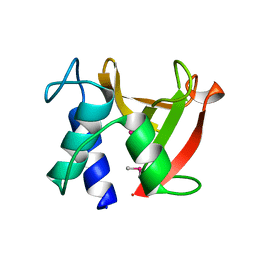 | |
8I2F
 
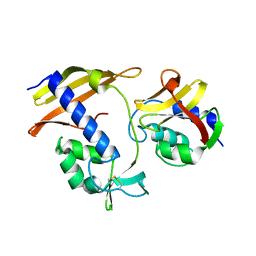 | |
8WTB
 
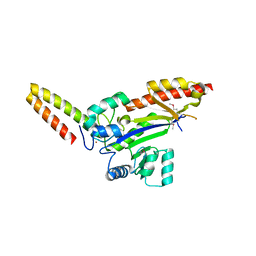 | |
8WTC
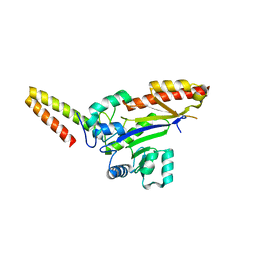 
 | |
2P52
 
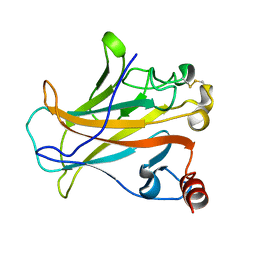 | |
1YGZ
 
 | | Crystal Structure of Inorganic Pyrophosphatase from Helicobacter pylori | | 分子名称: | Inorganic pyrophosphatase | | 著者 | Wu, C.A, Lokanath, N.K, Kim, D.Y, Park, H.J, Hwang, H.Y, Kim, S.T, Suh, S.W, Kim, K.K. | | 登録日 | 2005-01-06 | | 公開日 | 2005-11-01 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Structure of inorganic pyrophosphatase from Helicobacter pylori.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
3BG4
 
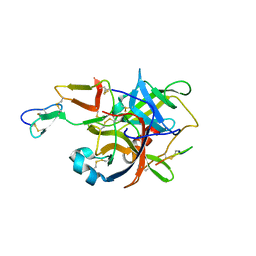 | | The crystal structure of guamerin in complex with chymotrypsin and the development of an elastase-specific inhibitor | | 分子名称: | Chymotrypsin A chain A, Chymotrypsin A chain B, Chymotrypsin A chain C, ... | | 著者 | Kim, H, Chu, T.T.T, Kim, D.Y, Kim, D.R, Nguyen, C.M.T, Choi, J, Lee, J.R, Hahn, M.J, Kim, K.K. | | 登録日 | 2007-11-26 | | 公開日 | 2008-07-29 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | The crystal structure of guamerin in complex with chymotrypsin and the development of an elastase-specific inhibitor.
J.Mol.Biol., 376, 2008
|
|
3OEO
 
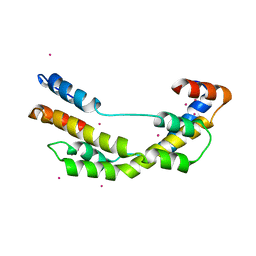 | | The crystal structure E. coli Spy | | 分子名称: | CADMIUM ION, Spheroplast protein Y | | 著者 | Kwon, E, Kim, D.Y, Gross, C.A, Gross, J.D, Kim, K.K. | | 登録日 | 2010-08-13 | | 公開日 | 2010-09-22 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | The crystal structure Escherichia coli Spy.
Protein Sci., 19, 2010
|
|
3V67
 
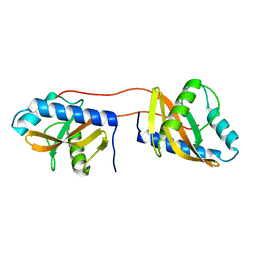 | | Periplasmic domain of Vibrio parahaemolyticus CpxA | | 分子名称: | Sensor protein CpxA | | 著者 | Kwon, E, Kim, D.Y, Ngo, T.D, Gross, J.D, Kim, K.K. | | 登録日 | 2011-12-19 | | 公開日 | 2012-09-26 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | The crystal structure of the periplasmic domain of Vibrio parahaemolyticus CpxA
Protein Sci., 21, 2012
|
|
7WA4
 
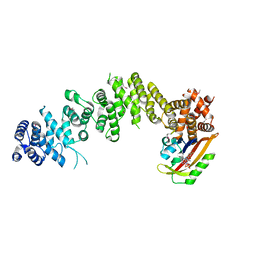 | | Crystal structure of GIGANTEA in complex with LKP2 | | 分子名称: | Adagio protein 2, FLAVIN MONONUCLEOTIDE, Protein GIGANTEA | | 著者 | Pathak, D, Dahal, P, Kwon, E, Kim, D.Y. | | 登録日 | 2021-12-12 | | 公開日 | 2022-04-27 | | 最終更新日 | 2022-05-04 | | 実験手法 | X-RAY DIFFRACTION (3.5 Å) | | 主引用文献 | Structural analysis of the regulation of blue-light receptors by GIGANTEA.
Cell Rep, 39, 2022
|
|
6JHK
 
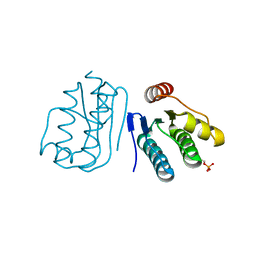 | |
6M37
 
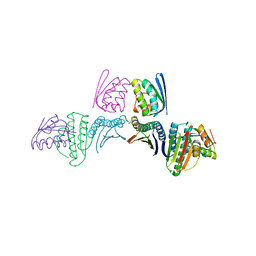 | |
6M36
 
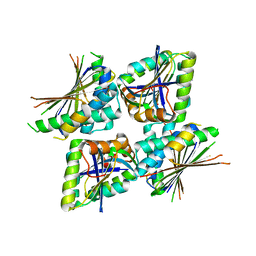 | |
6C3R
 
 | |
7W42
 
 | | Crystal structure of Bacillus subtilis YjoB | | 分子名称: | Uncharacterized ATPase YjoB | | 著者 | Dahal, P, Kwon, E, Kim, D.Y. | | 登録日 | 2021-11-26 | | 公開日 | 2022-10-19 | | 最終更新日 | 2024-05-29 | | 実験手法 | X-RAY DIFFRACTION (2.619 Å) | | 主引用文献 | Crystal structure and biochemical analysis suggest that YjoB ATPase is a putative substrate-specific molecular chaperone.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
