6NJF
 
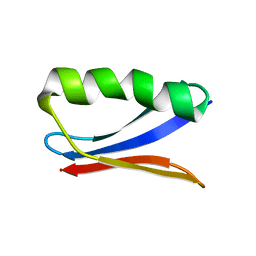 | | Solution NMR Structure of DANCER3-F34A, a rigid and natively folded single mutant of the dynamic protein DANCER-3 | | 分子名称: | Immunoglobulin G-binding protein G | | 著者 | Damry, A.M, Mayer, M.M, Goto, N.K, Chica, R.A. | | 登録日 | 2019-01-03 | | 公開日 | 2019-08-21 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Origin of conformational dynamics in a globular protein.
Commun Biol, 2, 2019
|
|
5UBS
 | | Solution NMR Structure of NERD-S, a natively folded pentamutant of the B1 domain of streptococcal protein G (GB1) with a solvent-exposed Trp43 | | 分子名称: | Immunoglobulin G-binding protein G | | 著者 | Damry, A.M, Davey, J.A, Goto, N.K, Chica, R.A. | | 登録日 | 2016-12-21 | | 公開日 | 2017-08-23 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Rational design of proteins that exchange on functional timescales.
Nat. Chem. Biol., 13, 2017
|
|
5UCE
 | | Solution NMR structure of the major species of DANCER-2, a dynamic and natively folded pentamutant of the B1 domain of streptococcal protein G (GB1) | | 分子名称: | Immunoglobulin G-binding protein G | | 著者 | Damry, A.M, Davey, J.A, Goto, N.K, Chica, R.A. | | 登録日 | 2016-12-22 | | 公開日 | 2017-08-23 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Rational design of proteins that exchange on functional timescales.
Nat. Chem. Biol., 13, 2017
|
|
5UCF
 | | Solution NMR-derived model of the minor species of DANCER-2, a dynamic and natively folded pentamutant of the B1 domain of streptococcal protein G (GB1) | | 分子名称: | Immunoglobulin G-binding protein G | | 著者 | Damry, A.M, Davey, J.A, Goto, N.K, Chica, R.A. | | 登録日 | 2016-12-22 | | 公開日 | 2017-08-23 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Rational design of proteins that exchange on functional timescales.
Nat. Chem. Biol., 13, 2017
|
|
5UB0
 
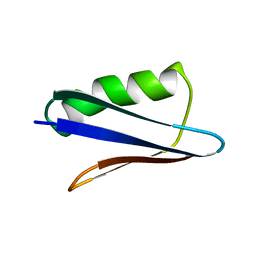 | | Solution NMR Structure of NERD-C, a natively folded tetramutant of the B1 domain of streptococcal protein G (GB1) | | 分子名称: | Immunoglobulin G-binding protein G | | 著者 | Damry, A.M, Davey, J.A, Goto, N.K, Chica, R.A. | | 登録日 | 2016-12-20 | | 公開日 | 2017-08-23 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Rational design of proteins that exchange on functional timescales.
Nat. Chem. Biol., 13, 2017
|
|
8EKG
 
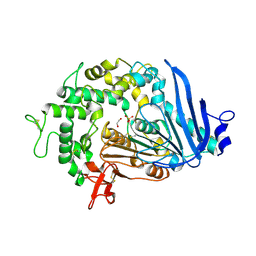 | | MHETase variant Thr159Val, Met192Tyr, Tyr252Phe, Tyr503Trp | | 分子名称: | 1,2-ETHANEDIOL, CALCIUM ION, DI(HYDROXYETHYL)ETHER, ... | | 著者 | Saunders, J.W, Frkic, R.L, Jackson, C.J. | | 登録日 | 2022-09-20 | | 公開日 | 2023-10-18 | | 最終更新日 | 2024-07-10 | | 実験手法 | X-RAY DIFFRACTION (2.65 Å) | | 主引用文献 | Increasing the Soluble Expression and Whole-Cell Activity of the Plastic-Degrading Enzyme MHETase through Consensus Design.
Biochemistry, 63, 2024
|
|
7LOO
 
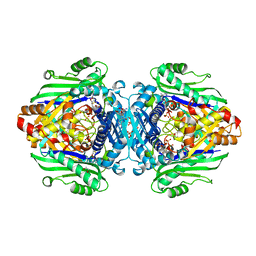 | | S-adenosyl methionine transferase cocrystallized with ATP | | 分子名称: | 1,2-ETHANEDIOL, MAGNESIUM ION, PHOSPHATE ION, ... | | 著者 | Jackson, C.J, Tan, L.L, Laurino, P. | | 登録日 | 2021-02-10 | | 公開日 | 2021-09-15 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | Substrate Dynamics Contribute to Enzymatic Specificity in Human and Bacterial Methionine Adenosyltransferases.
Jacs Au, 1, 2021
|
|
7LL3
 
 | | S-adenosylmethionine synthetase co-crystallized with UppNHp | | 分子名称: | (DIPHOSPHONO)AMINOPHOSPHONIC ACID, 1,2-ETHANEDIOL, 5'-O-[(R)-hydroxy{[(S)-hydroxy(phosphonoamino)phosphoryl]oxy}phosphoryl]uridine, ... | | 著者 | Tan, L.L, Jackson, C.J. | | 登録日 | 2021-02-03 | | 公開日 | 2021-03-31 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.24 Å) | | 主引用文献 | Substrate Dynamics Contribute to Enzymatic Specificity in Human and Bacterial Methionine Adenosyltransferases.
Jacs Au, 1, 2021
|
|
7LOW
 
 | | S-adenosylmethionine synthetase | | 分子名称: | 1,2-ETHANEDIOL, MAGNESIUM ION, PHOSPHATE ION, ... | | 著者 | Tan, L.L, Jackson, C.J. | | 登録日 | 2021-02-10 | | 公開日 | 2021-03-31 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.39 Å) | | 主引用文献 | Substrate Dynamics Contribute to Enzymatic Specificity in Human and Bacterial Methionine Adenosyltransferases.
Jacs Au, 1, 2021
|
|
7LOZ
 
 | | S-adenosylmethionine synthetase | | 分子名称: | 1,2-ETHANEDIOL, MAGNESIUM ION, PHOSPHATE ION, ... | | 著者 | Tan, L.L, Jackson, C.J. | | 登録日 | 2021-02-11 | | 公開日 | 2021-03-31 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.25 Å) | | 主引用文献 | Substrate Dynamics Contribute to Enzymatic Specificity in Human and Bacterial Methionine Adenosyltransferases.
Jacs Au, 1, 2021
|
|
7LNN
 
 | |
7LO2
 
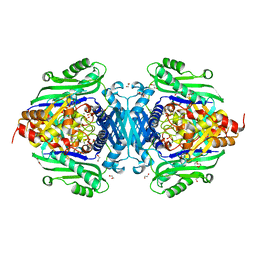 | |
7LNH
 
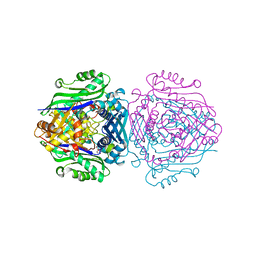 | | S-adenosylmethionine synthetase co-crystallized with UppNHp | | 分子名称: | (2~{S})-2-azanyl-4-[[(2~{S},3~{S},4~{R},5~{R})-5-[2,4-bis(oxidanylidene)pyrimidin-1-yl]-3,4-bis(oxidanyl)oxolan-2-yl]methyl-methyl-$l^{3}-sulfanyl]butanoic acid, 1,2-ETHANEDIOL, S-adenosylmethionine synthase isoform type-2 | | 著者 | Tan, L.L, Jackson, C.J. | | 登録日 | 2021-02-07 | | 公開日 | 2022-03-02 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Substrate Dynamics Contribute to Enzymatic Specificity in Human and Bacterial Methionine Adenosyltransferases.
Jacs Au, 1, 2021
|
|
8E9N
 
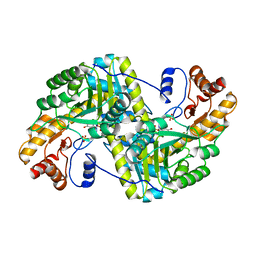 | | Crystal structure of E. coli aspartate aminotransferase mutant VFIY in the ligand-free form at 278 K | | 分子名称: | Aspartate aminotransferase, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.88 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9L
 | | Crystal structure of E. coli aspartate aminotransferase mutant VFIT in the ligand-free form at 278 K | | 分子名称: | Aspartate aminotransferase, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.31 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9M
 
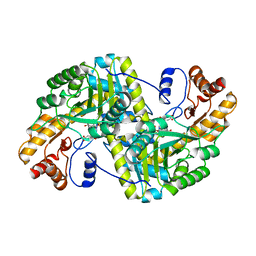 | | Crystal structure of E. coli aspartate aminotransferase mutant VFIT bound to maleic acid at 278 K | | 分子名称: | Aspartate aminotransferase, MALEIC ACID, PYRIDOXAL-5'-PHOSPHATE | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.76 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9R
 
 | | Crystal structure of E. coli aspartate aminotransferase mutant VFCS in the ligand-free form at 278 K | | 分子名称: | Aspartate aminotransferase, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9K
 
 | | Crystal structure of wild-type E. coli aspartate aminotransferase bound to maleate at 278 K | | 分子名称: | Aspartate aminotransferase, MALEIC ACID, PYRIDOXAL-5'-PHOSPHATE | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.83 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9Q
 
 | | Crystal structure of E. coli aspartate aminotransferase mutant HEX bound to maleic acid at 278 K | | 分子名称: | Aspartate aminotransferase, MALEIC ACID, PYRIDOXAL-5'-PHOSPHATE | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9O
 
 | | Crystal structure of E. coli aspartate aminotransferase mutant VFIY bound to maleic acid at 278 K | | 分子名称: | Aspartate aminotransferase, MALEIC ACID, PYRIDOXAL-5'-PHOSPHATE | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.96 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9T
 
 | | Crystal structure of wild-type E. coli aspartate aminotransferase in the ligand-free form at 303 K | | 分子名称: | Aspartate aminotransferase, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.13 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9S
 
 | | Crystal structure of E. coli aspartate aminotransferase mutant VFCS bound to maleic acid at 278 K | | 分子名称: | Aspartate aminotransferase, MALEIC ACID, PYRIDOXAL-5'-PHOSPHATE | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9V
 
 | | Crystal structure of E. coli aspartate aminotransferase mutant VFIT in the ligand-free form at 303 K | | 分子名称: | Aspartate aminotransferase, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-10-05 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.01 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9J
 
 | | Crystal structure of E. coli aspartate aminotransferase mutant HEX in the ligand-free form at 278 K | | 分子名称: | Aspartate aminotransferase, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-11-02 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.09 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
8E9P
 
 | | Crystal structure of wild-type E. coli aspartate aminotransferase in the ligand-free form at 278 K | | 分子名称: | Aspartate aminotransferase, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | 著者 | Chica, R.A, St-Jacques, A.D, Rodriguez, J.M, Thompson, M.C. | | 登録日 | 2022-08-26 | | 公開日 | 2022-11-02 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.08 Å) | | 主引用文献 | Computational remodeling of an enzyme conformational landscape for altered substrate selectivity.
Nat Commun, 14, 2023
|
|
