2P6I
 
 | | Crystal structure of PH0725 from Pyrococcus horikoshii OT3 | | 分子名称: | S-ADENOSYL-L-HOMOCYSTEINE, diphthine synthase | | 著者 | Yamamoto, H, Matsuura, Y, Morikawa, Y, Shimada, H, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | 登録日 | 2007-03-18 | | 公開日 | 2007-09-18 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | Crystal structure of PH0725 from Pyrococcus horikoshii OT3
To be Published
|
|
2P6K
 
 | | Crystal structure of PH0725 from Pyrococcus horikoshii OT3 | | 分子名称: | S-ADENOSYL-L-HOMOCYSTEINE, SULFATE ION, diphthine synthase | | 著者 | Yamamoto, H, Matsuura, Y, Ono, N, Nakamoto, T, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | 登録日 | 2007-03-18 | | 公開日 | 2007-09-18 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Crystal structure of PH0725 from Pyrococcus horikoshii OT3
To be Published
|
|
2PCK
 
 | | Crystal structure of PH0725 from Pyrococcus horikoshii OT3 | | 分子名称: | S-ADENOSYL-L-HOMOCYSTEINE, diphthine synthase | | 著者 | Yamamoto, H, Morikawa, Y, Matsuura, Y, Shimada, H, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | 登録日 | 2007-03-30 | | 公開日 | 2007-10-02 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Crystal structure of PH0725 from Pyrococcus horikoshii OT3
To be Published
|
|
6U5D
 
 | | RT XFEL structure of CypA solved using LCP injection system | | 分子名称: | Peptidyl-prolyl cis-trans isomerase A | | 著者 | Wolff, A.M, Young, I.D, Sierra, R.G, Brewster, A.S, Koralek, J.D, Boutet, S, Sauter, N.K, Fraser, J.S, Thompson, M.C. | | 登録日 | 2019-08-27 | | 公開日 | 2020-01-29 | | 最終更新日 | 2023-10-11 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | Comparing serial X-ray crystallography and microcrystal electron diffraction (MicroED) as methods for routine structure determination from small macromolecular crystals
Iucrj, 7, 2020
|
|
6U5C
 
 | |
6U5G
 
 | | MicroED structure of a FIB-milled CypA Crystal | | 分子名称: | Peptidyl-prolyl cis-trans isomerase A | | 著者 | Wolff, A.M, Martynowycz, M.W, Zhao, W, Gonen, T, Fraser, J.S, Thompson, M.C. | | 登録日 | 2019-08-27 | | 公開日 | 2020-01-29 | | 最終更新日 | 2023-10-11 | | 実験手法 | ELECTRON CRYSTALLOGRAPHY (2.5 Å) | | 主引用文献 | Comparing serial X-ray crystallography and microcrystal electron diffraction (MicroED) as methods for routine structure determination from small macromolecular crystals
Iucrj, 7, 2020
|
|
8CVU
 
 | | 20ns Temperature-Jump (Light) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
5WS5
 
 | | Native XFEL structure of photosystem II (preflash dark dataset) | | 分子名称: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, ... | | 著者 | Suga, M, Shen, J.R. | | 登録日 | 2016-12-05 | | 公開日 | 2017-03-15 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (2.35 Å) | | 主引用文献 | Light-induced structural changes and the site of O=O bond formation in PSII caught by XFEL.
Nature, 543, 2017
|
|
5WS6
 
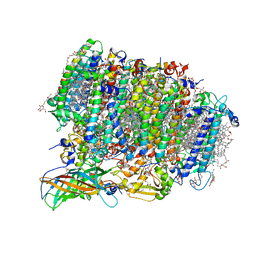 | | Native XFEL structure of Photosystem II (preflash two-flash dataset | | 分子名称: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, ... | | 著者 | Suga, M, Shen, J.R. | | 登録日 | 2016-12-05 | | 公開日 | 2017-03-15 | | 最終更新日 | 2020-07-29 | | 実験手法 | X-RAY DIFFRACTION (2.35 Å) | | 主引用文献 | Light-induced structural changes and the site of O=O bond formation in PSII caught by XFEL.
Nature, 543, 2017
|
|
5JOO
 
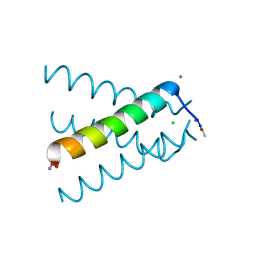 | | XFEL structure of influenza A M2 wild type TM domain at low pH in the lipidic cubic phase at room temperature | | 分子名称: | CALCIUM ION, CHLORIDE ION, Matrix protein 2 | | 著者 | Thomaston, J.L, Woldeyes, R.A, Fraser, J.S, DeGrado, W.F. | | 登録日 | 2016-05-02 | | 公開日 | 2017-08-02 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (1.413 Å) | | 主引用文献 | XFEL structures of the influenza M2 proton channel: Room temperature water networks and insights into proton conduction.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
8CW0
 
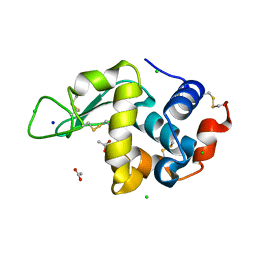 | | 20us Temperature-Jump (Light) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWH
 
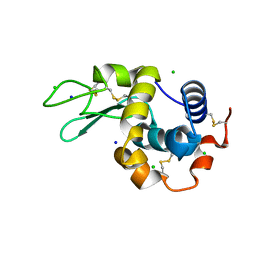 | | 200us Temperature-Jump (Dark2) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW8
 
 | | Laser Off Temperature-Jump XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW5
 
 | | 200us Temperature-Jump (Light) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-30 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWC
 
 | | 20ns Temperature-Jump (Light) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW1
 | | 20us Temperature-Jump (Dark1) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWG
 
 | | 200us Temperature-Jump (Dark1) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW7
 
 | | 200us Temperature-Jump (Dark2) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW6
 
 | | 200us Temperature-Jump (Dark1) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-09 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW3
 
 | | 20us Temperature-Jump (Dark2) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWB
 
 | | Laser Off Temperature-Jump XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.51 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWE
 
 | | 20ns Temperature-Jump (Dark2) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CVV
 
 | | 20ns Temperature-Jump (Dark1) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-30 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWD
 
 | | 20ns Temperature-Jump (Dark1) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CVW
 
 | | 20ns Temperature-Jump (Dark2) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
